Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
Southern African Journal of HIV Medicine
On-line version ISSN 2078-6751
Print version ISSN 1608-9693
South. Afr. j. HIV med. (Online) vol.22 n.1 Johannesburg 2021
http://dx.doi.org/10.4102/sajhivmed.v22i1.1185
ORIGINAL RESEARCH
HIV-1/2 differentiation in a South African public laboratory
Rendani T. MafuyekaI; Lynne M. WebberI, II; Piet BeckerIII; Simnikiwe H. MayaphiI
IDepartment of Medical Virology, Faculty of Health Sciences, University of Pretoria, Pretoria, South Africa
IIDepartment of Virology, Tshwane Academic division, National Health Laboratory, Pretoria, South Africa
IIIResearch Office, Faculty of Health Sciences, University of Pretoria, Pretoria, South Africa
ABSTRACT
BACKGROUND: The human immunodeficiency virus type-2 (HIV-2) prevalence in South Africa (SA) is unknown, however, sporadic cases have been reported. Human immunodeficiency virus -1 and 2 differentiation is not part of most South African public laboratories' testing algorithm. Human immunodeficiency virus -2 diagnosis using serology assays may be complicated by HIV-1 and HIV-2 antibody cross-reactivity.
OBJECTIVES: To determine the proportion of HIV-2 infections in specimens that tested HIV-1/2 positive at a public laboratory in Tshwane.
METHOD: A total of 480 specimens that were previously tested with fourth generation ELISA platforms (Modular E170 [Roche, Switzerland] and Architect i2000 [Abbott, Germany]) were randomly selected. Human immunodeficiency virus -1 and 2 antibody differentiation testing was carried out using the Multispot HIV-1/2 rapid assay (Bio-Rad Laboratories, USA). An in-house nested HIV-2 PCR assay targeting the 5′-long terminal repeats (5′-LTR) region was evaluated and used as a confirmatory test
RESULTS: The study tested 480 HIV-1/2 seropositive patients and their mean age was 36.7 years (range 3-82 years). Of the 480 patients, 292 (60.8%) were female, 182 (37.9%) were male and 6 (1.3%) were not specified. Human immunodeficiency virus differentiation results were as follows: 466 (97.1%) were positive for only HIV-1 antibodies, 11 (2.3%) [95%CI: (0.98%; 3.74%)] were positive for both HIV-1 and HIV-2 antibodies, 3 (0.6%) were negative for both antibodies and none were positive for only HIV-2 antibodies. Of the 11 specimens with both HIV-1 and HIV-2 antibodies, seven had sufficient volume for confirmatory testing and were all negative on the in-house HIV-2 PCR assay.
CONCLUSION: The multispot HIV-1/2 rapid assay demonstrated cross-reactivity between HIV-1 and HIV-2 antibodies. Human immunodeficiency virus -2 infections were not detected.
Keywords: HIV-1/2 differentiation; HIV-2 testing; HIV1/2 antibody cross reaction; HIV-2 in South Africa; HIV-2 PCR.
Introduction
Human immunodeficiency virus type 2 (HIV-2) belongs to the Retroviridae family, Lentivirus genus.1 It is most prevalent in West Africa.2 Most countries with historical ties with West Africa have also detected a significant proportion of HIV-2 infections.3 Although the HIV pandemic is mainly caused by HIV-1, HIV-2 is also an important cause of acquired immunodeficiency syndrome (AIDS).4 Both HIV-1 and HIV-2 have similar clinical manifestations, however, HIV-2 progresses much slower to AIDS.5
South African data on HIV-2 infections is limited. Sporadic cases have been reported in the past, which is an indication that HIV-2 may be circulating in South Africa (SA).6,7 In addition, SA has a large number of tourists and immigrants from the West African region and other parts of the world where HIV-2 infections have been reported,8,9 which may contribute to the spread of the virus in the country.
Routine diagnostic serology assays that are widely used in SA national public sector health laboratories do not include HIV-1 and HIV-2 differentiation.10 This may have implications for HIV management,11 as the treatment of HIV-2 differs from that of HIV-1. Human immunodeficiency virus-2 is intrinsically resistant to non-nucleoside reverse transcriptase inhibitors (NNRTIs) and fusion inhibitors. It is also only weakly suppressed by some protease inhibitors. Human immunodeficiency virus-2 has a lower genetic barrier compared with HIV-1 to nucleoside reverse transcriptase inhibitors (NRTIs) resistance.
Dolutegravir is effective against HIV-2, but strains with raltegravir mutations may have low levels of resistance.12
The virology diagnostic laboratory is currently using a fourth generation enzyme-linked immunosorbent assay (4th gen ELISA) for HIV-1/2 testing. The fourth generation ELISA can detect both HIV-1 and HIV-2 antibodies and p24 antigen, but it does not differentiate between the two types. The World Health Organization (WHO) HIV testing guidelines recommend HIV-2 differentiation with serological and/or molecular assays in settings where HIV-2 infections have been detected.13 Human immunodeficiency virus -2 molecular testing is an important confirmatory assay, owing to the high HIV-1 and HIV-2 antibody cross-reactivity with serological assays.14
Aims
This study aimed to determine the proportion of HIV-2 infections in specimens that tested HIV-1/2 positive in a public laboratory in Tshwane.
Objectives
•Human immunodeficiency virus -1 and -2 differentiation using the Multispot HIV-1/2 rapid test.
•Evaluation of the in-house HIV-2 nested polymerase chain reaction (PCR) assay.
•Confirmatory testing using the in-house HIV-2 PCR assay.
Methods and materials
Study design
This was a retrospective, cross-sectional study conducted between February 2013 and July 2016.
Study setting
This study was conducted in the National Health Laboratory Services (NHLS), Tshwane Academic Division (TAD) Virology Laboratory at the University of Pretoria, Faculty of Health Sciences. The specimens used came from the public sector hospitals and clinics around Tshwane Metropolitan Municipality, submitted for routine HIV screening.
Specimen selection
A total of 480 plasma and serum specimens that had previously tested positive for HIV-1/2 antibody, were randomly selected. The inclusion criteria were age of more than 24 months and HIV-1/2 antibody positive results on two different fourth generation ELISA platforms, the Modular E170 (Roche, Switzerland) and the Architect i2000 (Abbott, Germany).
Statistics
The HIV-2 prevalence is unknown in SA. Based on the HIV-2 prevalence data from other parts of the world outside the West African region,15,16,17 a prevalence of 1% was assumed for SA. It was determined that a sample size of 459 tests would have a 0.99 probability to detect at least one HIV-2 positive specimen. The sample size calculation was done using software nQuery version 7.0.18
HIV-1 and 2 antibody differentiation
The Multispot HIV-1/2 rapid assay (Bio-Rad Laboratories, USA) was used for HIV-1 and HIV-2 differentiation.
The procedure was performed according to the manufacturer's instructions. Four known HIV-2 antibody positive specimens, from a laboratory in Portugal, that were previously tested using only the Western Blot assay, and 52 HIV-1 and HIV-2 antibody negative specimens that were previously tested using HIV-1/2 fourth generation ELISA, Modular E170 (Roche, Switzerland) were included to evaluate performance of the Multispot HIV-1/2 rapid assay.
In-house HIV-2 nested polymerase chain reaction assay
Two sets of primers, targeting the conserved 5′ long terminal repeat (5′-LTR) region of HIV-2, were designed and numbered according to the HIV-2 ROD reference strain, accession number M15390.1 (Table 1). The in-house HIV-2 nested PCR assay was evaluated using the four known HIV-2 antibody positive specimens. These specimens were also processed at a private laboratory using a commercial real-time HIV-2 PCR assay (Genesig Standard kit) that targets the integrase (pol) gene region. The Genesig standard kit detects HIV-2 subtypes A and B only. The internal control was also included in the commercial real-time assay. Two known HIV-1 specimens with high viral load (132 000 copies/mL and 30 200 copies/mL) were included to determine the specificity of the in-house PCR primers. The in-house HIV-2 nested PCR assay was used to confirm specimens that were positive for both HIV-1 and HIV-2 antibodies on the Multispot HIV-1/2 rapid assay.
Nucleic acid extraction
Except for the initial centrifugation step to get a purified pellet, the extraction of total nucleic acids was performed manually using the QIAamp UltraSens Virus kit (Qiagen, Germany) according to the manufacturer's instructions.
Centrifugation was optimised at 800 relative centrifugal force (RCF) for 3 min instead of the recommended 1200 RCF. Optimisation to a lower RCF produced an easily dissolvable pellet. Sample input volume for extraction was 500 µL, and lower input volume of 250 µL was used for samples with insufficient volume. The final eluate volume was 60 µL.
Amplification, detection and analysis
A nested PCR was carried out using the Superscript III One-Step RT-PCR kit with platinum Taq deoxyribonucleic acid polymerase (5 U/µL) (Life technologies, USA) according to the manufacturer's instructions. The total reaction volumes for both first and second round PCR were 50 µL. First round PCR contained 25 µL of 2x reaction mix (a buffer containing 0.4 mM of each dNTP, 2.4 mM MgSO4); 1 µL of each primer (outer forward and reverse) (10 µM); 1 µL of Taq enzyme (5 U/µL), 5 µL of template and 17 µL of nuclease-free H2O. The second round PCR reaction contained 25 µL of 2x reaction mix (a buffer containing 0.4 mM of each dNTP, 2.4 mM MgSO4); 1 µL of each primer (inner forward and reverse)(10 µM); 0.2 µL of Taq enzyme (5 U/µL), 0.5 µL of template and 22.3 µL of nuclease-free H2O.
The first round PCR conditions were as follows: cDNA synthesis for 1 cycle at 50 °C for 30 min; denaturation for 1 cycle at 94 °C for 2 min; amplification 40 cycles (denature at 94 °C for 30 s, annealing at 50 °C for 30 s, extension at 68 °C for 1 min); final extension for 1 cycle at 68 °C for 7 min and hold at 4 °C. The second round PCR conditions were as follows: denaturation for 1 cycle at 94 °C for 2 min; amplification 40 cycles (denature at 94 °C for 30 s, annealing at 50 °C for 30 s, extension at 68 °C for 30 s); final extension for 1 cycle at 68 °C for 7 min and hold at 4 °C. Polymerase chain reaction amplicons were visualised on a 1.5% agarose gel by ultraviolet (UV) illumination.
HIV-2 sequencing
Polymerase chain reaction amplicons were sequenced to evaluate the in-house HIV-2 nested PCR, at Inqaba Biotechnical Industries (Pty) Ltd using the second round PCR primers (MMFw-2 and MMRv-2) (Table 1). The original PCR products were sent to Inqaba where they performed a clean-up procedure and Sanger sequencing. Sequence analysis comprised of sequence editing and alignment performed using BioEdit Sequence Alignment Editor Version 7.2.5 programme.19 Molecular Evolutionary Genetics Analysis (MEGA) version 6 programme,20 was used for constructing the phylogenetic tree using Neighbor-Joining statistical method (with 1000 bootstrap replicates).
Ethical consideration
This study was approved by the research ethics committee, Faculty Health Sciences, University of Pretoria (ethics reference number: 11/2013). Permission to use the blood specimens and to retrieve demographics information from the laboratory information system (LIS) was obtained from Tshwane Academic Division of the NHLS. The information of each specimen was managed with strict confidentiality and each specimen was given a unique study number to ensure patient anonymity.
Results
HIV-1 and 2 antibodies differentiation results
The study tested 480 HIV-1 and HIV-2 seropositive patients, and their mean age was 36.7 years (range: 3-82 years). Of the 480, 292 (60.8%) were female, 182 (37.9%) were male and 6 (1.3%) were not specified. All four known HIV-2 antibody positive specimens used to evaluate the Multispot HIV-1/2 rapid assay were positive for HIV-2 antibodies only. All 52 known HIV-1 and HIV-2 ELISA negative specimens were negative. Differentiation results of the 480 specimens were as follows: 466 (97.1%) were positive for only HIV-1 antibodies, 11 (2.3%) 95%CI: 0.98%; 3.74% were positive for both HIV-1 and HIV-2 antibodies, 3 (0.6%) were negative for both antibodies and none were positive for only HIV-2 antibodies (Figure 1).
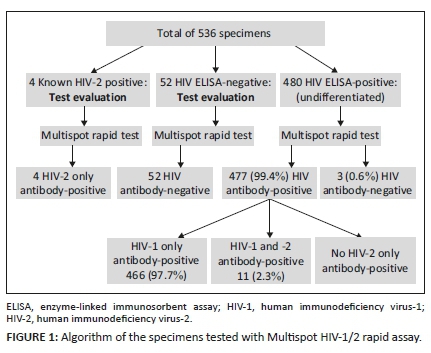
In-house HIV-2 nested polymerase chain reaction assay results
Evaluation results of the in-house nested HIV-2 PCR, on four known HIV-2 antibody positive specimens, were comparable to a commercial real-time PCR, which was carried out in a private laboratory (Table 2). Two HIV-1 specimens, with high viral load, were also not detected by the in-house HIV-2 PCR (Figure 2). Of the 11 specimens, which were positive for both HIV-1 and HIV-2 antibodies, only seven had sufficient volume for further testing. All tested HIV-2 PCR negative on the in-house assay.

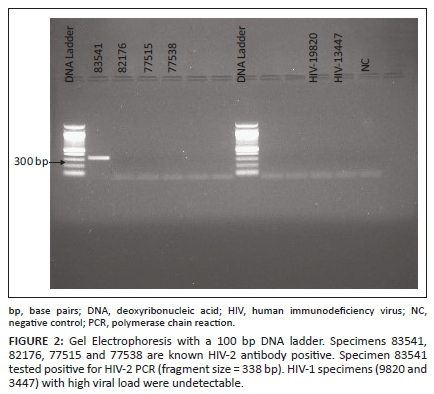
HIV-2 sequencing results
The PCR products of the confirmed HIV-2 subtype A/B specimen (83541) were sent for sequencing. Phylogenetic analysis revealed that this sequence clustered with subtype A of HIV-2 (Figure 3). Sequencing was not indicated for those specimens that were HIV-2 negative on the in-house PCR assay.
Discussion
This study aimed to determine the proportion of HIV-2 infections in specimens that tested HIV-1/2 antibody positive at a virology diagnostic laboratory using the Multispot HIV-1/2 rapid assay for differentiation. The Multispot rapid test has been reported as satisfactory with regard to the detection of HIV-1 and HIV-2.21,22,23,24,25,26
Differentiation results showed HIV-1 and HIV-2 dual seropositivity in 11 (2.3%) of the specimens. Of the 11, seven specimens with sufficient volume for the in-house HIV-2 PCR tested negative. This may indicate HIV-1 and HIV-2 antibody cross-reactivity. These results were not surprising, as HIV-1 and HIV-2 antibody cross-reactivity have been observed in other studies.24,27 A study conducted in SA demonstrated high levels of false HIV-2 seropositivity, at 15%, 71.2% and 10.7%, using the SD Bioline HIV1/2 3.0 rapid test, the NewLAV Western blots, and Pepti-LAV1-2, respectively.27 This South African study demonstrated a high rate of antibody cross reactivity with negative HIV-2 molecular testing. Another study carried out in the United States of America, demonstrated antibody cross-reactivity rates of 46.5% and 11.4% using the Centres for Disease Control (CDC) HIV-1 Western blot (WB) interpretive criteria and the alternative more stringent HIV-1 WB interpretive criteria, respectively.24 The HIV-1 and HIV-2 antibody cross-reactivity rate was 2.3% in this study. Other previous studies that also used the Multispot assay demonstrated a cross-reactivity rate of 0.5%26 and 1.3%.24 These previous studies are further demonstrating that HIV-2 testing using serological assays may be complicated owing to high levels of antibody cross-reactivity and should be interpreted with caution. Highly sensitive and specific molecular confirmatory testing is always important for those specimens with dual HIV seropositivity14 and for research purposes.
The first two reported HIV-2 cases in SA were documented in 19886 and 1993.7 These cases were both diagnosed and confirmed using antibody detection assays. Confirmation of the 1988 case was done using an immunofluorescence assay and that of the 1993 case was done using a Western blot assay. Possibility of HIV-1 and HIV-2 antibody cross reactivity in these two cases cannot be ruled out as no molecular testing was performed.
The in-house nested HIV-2 PCR had comparable results, on the four known HIV-2 antibody positive specimens, with the commercial assay (Genesig Standard kit) processed in a private laboratory (Table 2). Sequencing of the PCR products of the known HIV-2 positive specimen (83541) indicated HIV-2 subtype A (Figure 3). The other three HIV-2 seropositive specimens that failed to amplify using HIV-2-specific PCRs (the in-house and the commercial assays) may have reflected naturally low levels of HIV-2 virus or the effect of antiretroviral therapy exposure. There was no relevant clinical information provided on these known HIV-2 antibody positive specimens from the laboratory in Portugal, which could have explained these results. In addition, the in-house nested HIV-2 PCR assay did not detect HIV-1 in two specimens with high viral loads highlighting a lower likelihood of false positive results in individuals with HIV-1 infection. To further evaluate the sensitivity and specificity of the primers used for in-house nested PCR, more HIV-1 and HIV-2 samples need to be tested.
The negative HIV-2 PCR on specimens with dual HIV seropositivity may not completely rule out dual HIV infections, as there are many factors that may contribute to false-negative HIV-2 PCR. Such factors include naturally low viral load in HIV-2 infections because of the low replication rate.28 In a study by Drylewicz J et al., plasma HIV-2 viral load was undetectable in up to 85% of individuals positive for HIV-2 antibodies.29 Another possible explanation could be that HIV-1 suppresses HIV-2 replication in individuals with dual infection; hence, the HIV-2 antibodies and viral load remain very low or undetectable. The quality of specimens used in this study may have been compromised as well. Specimens were initially stored at 4 °C for more than 7 days prior to testing, which may have led to RNA degradation. Serum and plasma specimens used in this study may not be the best for HIV-2 detection as evidence suggests that HIV-2 replication is restricted in vivo.30 Peripheral blood mononuclear cells (PBMCs) may yield a better outcome with regard to HIV-2 detection.30,31
The limitations of this study include small sample size, quality and type of the specimens used as described here.
Low specimen volumes (250 µL - 500 µL) were used for extraction because of specimen insufficiency. Nucleic acid quantification and quality assessment were also not done. This may have led to false negative results. Polymerase chain reaction failure of the in-house PCR could not be excluded because of the lack of internal controls; however, the commercial HIV-2 PCR with comparable results and with internal controls, amplified. The developed in-house nested HIV-2 PCR needs further evaluation with more HIV-2 positive specimens, including other subtypes and addition of more HIV-1 only specimens. The commercial assay can only detect HIV-2 subtypes A and B.32 Detection of other HIV-2 subtypes (C-G) could not be evaluated.
Conclusion
The Multispot HIV-1/2 rapid assay demonstrated some cross-reactivity between HIV-1 and HIV-2 antibodies. Human immunodeficiency virus-2 infections were not detected. Dual infections could not be excluded in this study. The in-house HIV-2 PCR assay needs further evaluation to determine its sensitivity and specificity. These results cannot be generalised because the sample size was too small. Further studies are needed to explore HIV-2 infections and treatment response in SA.
Postscript
The Multispot HIV-1 and HIV-2 rapid assay was later discontinued in 2016 and was replaced by the Geenius HIV-1 and HIV-2 Supplemental Assay.33
Acknowledgements
(1) Tshwane Academic division of the NHLS and the University of Pretoria Virology Department for support and supply of all patients' specimens, (2) Dr Osman for arranging the known HIV-2 specimens from a Portugal laboratory and (3) Dr Allison Glass (private laboratory) for testing HIV-2 PCR on the four known HIV-2 specimens.
This research was conducted for the fulfilment of the MMed in Pathology (Clinical Virology) and supervised by Simnikiwe Horatious Mayaphi and Lynne Margaret Webber.
Competing interests
The authors declare that they have no financial or personal relationships that may have inappropriately influenced them in writing this article.
Authors' contributions
S.H.M. and L.M.W. developed the study concept, design, manuscript editing and they were also supervisors. P.B. contributed in data analysis, statistics and manuscript editing. R.T.M. conducted laboratory tests, data collection, data analysis and writing manuscript.
Funding information
This work was supported by the NHLS Research Trust (GRANT004 94395), Discovery Foundation (reference number 034211) and the University of Pretoria Research grant.
Data availability
The data that support the findings of this study are available from the corresponding author, R.T.M., upon reasonable request.
Disclaimer
The views and opinions expressed in this article are those of the authors and do not necessarily reflect the official policy or position of any affiliated agency of the authors.
References
1.Richman DD, Whitley RJ, Hayden FG. Clinical virology. 3rd ed. Washington, DC: ASM Press; 2009, p. 737. [ Links ]
2.Armstrong-James D, Stebbing J, Scourfield A, et al. Clinical outcome in resistant HIV-2 infection treated with Raltegravir and Maraviroc. Antiviral Res. 2010;86(2):224-226. https://doi.org/10.1016/j.antiviral.2010.02.324 [ Links ]
3.Soriano V, Gomes P, Heneine W, et al. Human immunodeficiency virus type 2 (HIV-2) in Portugal: Clinical spectrum, circulating subtypes, virus isolation, and plasma viral load. J Med Virol. 2000;61(1):111-116. https://doi.org/10.1002/(SICI)1096-9071(200005)61:1%3C111::AID-JMV18%3E3.0.CO;2-W [ Links ]
4.Kashyap B, Gautam H, Bhalla P. Epidemiology and seroprevalence of human immunodeficiency virus type 2. Intervirology. 2011;54(3):151-155. https://doi.org/10.1159/000319929 [ Links ]
5.Nyamweya S, Hegedus A, Jaye A, Rowland-Jones S, Flanagan KL, Macallan DC. Comparing HIV-1 and HIV-2 infection: Lessons for viral immunopathogenesis. Rev Med Virol. 2013;23(4):221-240. https://doi.org/10.1002/rmv.1739 [ Links ]
6.Lyons SF, Smith AN, McGillivray GM, Schoub BD. HIV-2 infection in South Africa. T Roy Soc Trop Med H. 1988;82(5):757. https://doi.org/10.1016/0035-9203(88)90227-1 [ Links ]
7.Steele AD, Bezuidenhout S, Du Toit D, Lecatsas G. HIV-2 at Ga-Rankuwa hospital. SAMJ. 1993;83(7):530-531. [ Links ]
8.Statistics South Africa. Documented immigrants in South Africa [homepage on the Internet]. 2013 [cited 2016 Jul 05]. Available from: http://www.statssa.gov.za/ [ Links ]
9.Statistics South Africa. Tourism report [homepage on the Internet]. 2014 [cited 2016 Jul 05]. Available from: http://www.statssa.gov.za/ [ Links ]
10.SADoH. HIV counselling and testing (HCT) policy guidelines. Pretoria: Department of Health; 2010. [ Links ]
11.Chan PA, Wakeman SE, Flanigan T, Cu-Uvin S, Kojic E, Kantor R. HIV-2 diagnosis and quantification in high-risk patients. AIDS Res Ther. 2008;5(1):1-5. https://doi.org/10.1186/1742-6405-5-18 [ Links ]
12.Menéndez-Arias L, Álvarez M. Antiretroviral therapy and drug resistance in human immunodeficiency virus type 2 infection. Antiviral Res. 2014;102:70-86. https://doi.org/10.1016/j.antiviral.2013.12.001 [ Links ]
13.WHO. HIV testing guidelines. Geneva: World Health Organization; 2015. [ Links ]
14.Kannangai R, Ramalingam S, Prakash KJ, et al. Molecular confirmation of human immunodeficiency virus (HIV) type 2 in HIV-seropositive subjects in South India. Clin Diagn Lab Immunol. 2000;7(6):987-989. https://doi.org/10.1128/CDLI.7.6.987-989.2000 [ Links ]
15.Burgard M, Jasseron C, Matheron S, et al. Mother-to-child transmission of HIV-2 infection from 1986 to 2007 in the ANRS French perinatal cohort EPF-CO1. Clin Infect Dis. 2010;51(7):833-843. https://doi.org/10.1086/656284 [ Links ]
16.Maueia C, Costa D, Meggi B, et al. Frequency of human immunodeficiency virus type-2 in HIV infected patients in Maputo City, Mozambique. Virol J. 2011;8:408. https://doi.org/10.1186/1743-422X-8-408 [ Links ]
17.Pipero P, Qyra SH, Basho M, et al. Human immunodeficiency virus type-2 infection in Albania. Anglisticum. 2014;3(6):61. [ Links ]
18.nQuery & nTerim4.0 by Statistical Solutions Ltd [homepage on the Internet]. [cited 2017 Aug 08]. Available from: https://www.statsols.com/ [ Links ]
19.Bioedit Sequence Alignment Editor Programme version 7.2.5 [homepage on the Internet]. [cited 2016 Sep 20]. Available from: http://www.mbio.ncsu.edu/bioedit/bioedit.html/ [ Links ]
20.Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. Molecular Evolutionary Genetics Analysis (MEGA) version 6 programme. Mol Biol Evol. 2013;30(12):2725-2729. https://doi.org/10.1093/molbev/mst197 [ Links ]
21.Delaney KP, Branson BM, Uniyal A, et al. Evaluation of the performance characteristics of 6 rapid HIV antibody tests. Clin Infect Dis. 2011;52(2):257-263. https://doi.org/10.1093/cid/ciq068 [ Links ]
22.Holguı́n A. Evaluation of three rapid tests for detection of antibodies to HIV-1 non-B subtypes. J Virol Methods. 2004;115(1):105-107. https://doi.org/10.1016/j.jviromet.2003.09.028 [ Links ]
23.Mallocha L, Kadivara K, Putzb J, et al. Comparative evaluation of the Bio-Rad Geenius HIV-1/2 confirmatory assay and the Bio-Rad Multispot HIV-1/2 rapid test as an alternative differentiation assay for CLSI M53 algorithm-I. J Clin Virol. 2013;58S:85-91. https://doi.org/10.1016/j.jcv.2013.08.008 [ Links ]
24.Nasrullah M, Ethridge SF, Delaney KP, et al. Comparison of alternative interpretive criteria for the HIV-1 Western Blot and results of the Multispot HIV-1/HIV-2 rapid test for classifying HIV-1 and HIV-2 infections. J Clin Virol. 2011;52(Suppl 1):S23-S27. https://doi.org/10.1016/j.jcv.2011.09.020 [ Links ]
25.O'Connell RJ, Agan BK, Anderson SA, Malia JA, Michael NL. Sensitivity of the Multispot HIV-1/HIV-2 rapid test using samples from human immunodeficiency virus type-1 positive individuals with various levels of exposure to highly active antiretroviral therapy. J Clin Microbiol. 2006;44(5):1831-1833. https://doi.org/10.1128/JCM.44.5.1831-1833.2006 [ Links ]
26.Ramos EM, Harb S, Dragavon J, Coombs RW. Clinical performance of the Multispot HIV-1/HIV-2 rapid test to correctly differentiate HIV-2 from HIV-1 infection in screening algorithms using third and fourth generation assays and to identify cross reactivity with the HIV-1 Western Blot. J Clin Virol. 2013;58(Supp 1):S104-S107. https://doi.org/10.1016/j.jcv.2013.08.018 [ Links ]
27.Singh L, Parboosing R, Manasa J, Moodley P, De Oliveira T. High level of HIV-2 false positivity in KwaZulu-Natal province: A region of South Africa with a very high HIV-1 subtype C prevalence. J Med Virol. 2013;85(12):2065-2071. https://doi.org/10.1002/jmv.23716 [ Links ]
28.Delarue S, Didier E, Damond F, et al. Highly sensitive plasma RNA quantification by real-time PCR in HIV-2 group A and group B infection. J Clin Virol. 2013;58(2):461-467. https://doi.org/10.1016/j.jcv.2013.08.003 [ Links ]
29.Drylewicz J, Matheron S, Lazaro E, et al. Comparison of viro-immunological marker changes between HIV-1 and HIV-2-infected patients in France. AIDS. 2008;22(4):457-468. https://doi.org/10.1097/QAD.0b013e3282f4ddfc [ Links ]
30.MacNeil A, Sarr AD, Sankalé JL, Meloni ST, Mboup S, Kanki P. Direct evidence of lower viral replication rates in vivo in human immunodeficiency virus type 2 (HIV-2) infection than in HIV-1 infection. J Virol. 2007;81(10):5325-5330. https://doi.org/10.1128/JVI.02625-06 [ Links ]
31.Kageyama S, Maniar JK, Iwasaki M, et al. Seronegative HIV-2 carriers in India. Int J STD AIDS. 2000;11(1):31-37. https://doi.org/10.1258/0956462001914878 [ Links ]
32.HIV-2 Integrase (Pol) gene region. Genesig standard kit handbook HB10.02.08. US: Primerdesign Ltd. [ Links ]
33.Penner MC, National Alliance of State & Territorial AIDS Directors (NASTAD) [homepage on the Internet]. 2016 [2020 June 22]. Available from: https://www.nastad.org/sites/default/files/resources/docs/February-16Discontinuation-Multispot-HIV-Rapid-Test.pdf [ Links ]
 Correspondence:
Correspondence:
Rendani Mafuyeka
rendani.mafuyeka@up.ac.za
Received: 26 Oct. 2020
Accepted: 30 Jan. 2021
Published: 12 Mar. 2021














