Servicios Personalizados
Articulo
Indicadores
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares en Google
Similares en Google
Compartir
Water SA
versión On-line ISSN 1816-7950
versión impresa ISSN 0378-4738
Water SA vol.35 no.3 Pretoria abr. 2009
Low prevalence of antibiotic-resistant gram-negative bacteria isolated from rural south-western Ugandan groundwater
Olusegun O SogeI; Michael A GiardinoI; Iana C IvanovaI; Amber L PearsonII; John S MeschkeI ; Marilyn C RobertsI, *
IDepartment of Environmental & Occupational Health Sciences, Box 357234, School of Public Health, University of Washington, Seattle, WA 98195-7234
IIDepartments of Geography & Global Health, University of Washington, Seattle, WA 98195
ABSTRACT
The objective of this study was to determine antibiotic resistance patterns and specific resistance genes in Gram-negative enteric bacteria recovered from 42 different drinking water sources servicing 2 rural villages in south-western Uganda. These water sites were prone to contamination by both human and cattle activity. Of the 52 isolates examined, 26 carried antibiotic resistance genes with 25 being ampicillin resistant, 21 carrying the blaTEM gene, and no isolate carrying genes coding for extended-spectrum β−lactamases. Twelve isolates were tetracycline resistant and these bacteria carried between 1 and 3 different tet genes, with the tet(A) gene the most common. Six isolates carried the macrolide resistance mef(A) and/or the macrolide-lincosamide-streptogramin B resistance erm(B) genes. Four isolates carried the sul1 gene, and 4 isolates carried the sul1 and int1 genes indicating the presence of Class 1 integrons. The Ugandan isolates in this study had lower than expected carriage rates of antibiotic and multi-drug resistance genes, carriage of Class 1 integrons and lacked genes coding for extended-spectrum β−lactamases as compared to antibiotic resistance carriage in clinical African isolates.
Keywords: drinking water, antibiotic resistance, resistance genes, Gram-negative, conjugal transfer
Introduction
Pearson et al. (2008) reported on the isolation and biochemical characterisation of water-borne Gram-negative bacteria isolated from boreholes, ponds and valley water tanks used by 2 rural villages, from Nyabushozi County in the Mbarara District of south-western Uganda. Both humans and animals used the water sources and the E. coli counts confirmed that 38 of the 42 water sites did not meet international drinking water standards. The inhabitants of these villages were settled and semi-nomadic Bahima pastoralists, Bairu agriculturalists and a few internal migrants. The villages had undergone dramatic water resource changes as a result of land privatisation and the creation of a national park in 1989. The villagers were 10 to 15 km from the nearest local private medical clinic, which charged money for its services and 40 to 50 km from the nearest hospital. These communities lacked the infrastructure required for basic services, such as health care and a municipal drinking water supply. As a result, these people had little interaction with western medicine and relied on traditional herbal treatments for human and livestock diseases (authors' unpublished observations).
The currently published literature suggests that levels of antibiotic-resistant bacteria are high and continue to rise in Africa (Okeke et al., 2007). There are no data on antimicrobial susceptibilities of water bacteria from Uganda. Therefore it was of interest to characterise the level of antibiotic resistance and corresponding resistance genes in the Ugandan bacteria previously isolated from 42 of the 47 Ugandan water sites tested which serviced the 2 villages. In this study, the antibiotic resistance phenotypes and genotypes of 52 randomly selected Gram-negative enteric bacteria from Nyabushozi County in the Mbarara District of south-western Uganda were characterised.
Materials and methods
Bacterial isolates
Previously, water-borne Gram-negative bacteria were recovered from 1 mℓ water samples taken from 47 water sites within 2 villages in Nyabushozi County in the Mbarara District of south-western Uganda. The water samples were plated onto EC 3MTM PetrifilmTM according to the manufacturer's instructions and incubated at 37ºC for 24 h. Forty-two of the 47 water sites were positive for enteric bacteria. Random isolates were taken from the Petrifilms and biochemically identified and then representative isolates were verified by sequencing the variable region of their 16S rRNA gene as previously reported (Pearson et al., 2008). Many of the 16S rRNA gene sequences obtained allowed identification to the genus but not species level. From the initial study, 52 genetically distinct archived bacteria were available for further study. These included: 2 Citrobacter spp., 20 E. coli, 7 Enterobacter spp., 5 Klebsiella spp., 3 Morganella morganii, 2 Proteus spp., 3 Providencia rettgeri, 4 Pseudomonas spp., 4 Salmonella spp., and 2 Serratia odorifera. Because E. coli 0157:H7 had previously been reported in the area (Majalija et al., 2008), we cultured the 20 E. coli isolates on Sorbitol-MacConkey medium-SMAC (Remel, Inc., Lenexa, KS, USA; March and Ratnam, 1986). No growth was observed indicating that they were not E. coli 0157:H7. Two laboratory strains, E. coli HB101 and E. coli DH5α were used as recipients in conjugation experiments.
Antibiotic susceptibility
Antimicrobial susceptibilities were performed using disc diffusion on Mueller–Hinton agar (Remel, Inc., Lenexa, KS, USA) according to the CLSI guidelines (CLSI, 2003a). E. coli ATCC 25922 was used as a control. The antibiotic disks included ampicillin (10 µg), cefotaxime (30 µg), ceftazidime (30 µg), chloramphenicol (30 µg), kanamycin (30 µg), tetracycline (30 µg), trimethoprim/sulfamethoxazole (25 µg) (Remel, Inc., Lenexa, KS, USA). In addition, minimum inhibitory concentrations (MICs) were performed using an agar dilution method according to the CLSI guidelines (CLSI, 2003b) for ceftazidime, cefotaxime, aztreonam, piperacillin, cefepime, erythromycin, and imipenem for 26 ampicillin resistant (Apr) isolates.
Detection of antibiotic resistance genes
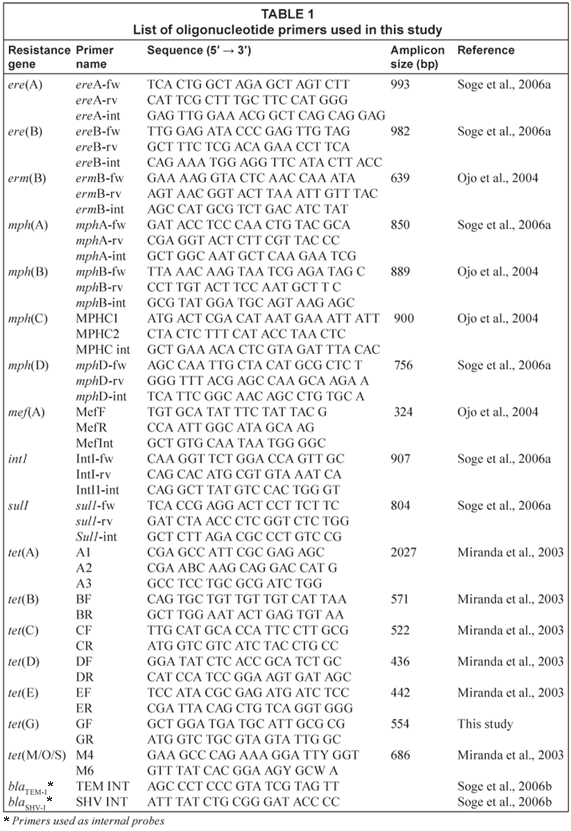
Polymerase chain reaction (PCR) assays were used for the detection of tetracycline resistance genes [tet(A), tet(B), tet(C), tet(D), tet(E), tet(G) and tet(M)]; macrolide or combinations of macrolide-lincosamide-streptogramin [MLS] genes [erm(B), mef(A), ere(A), ere(B), mph(A), mph(B), mph(C), and mph(D)]; the intI1 integrase for Class 1 integron and sul1 gene coding for sulphonamide resistance were performed as previously described (Miranda et al., 2003; Soge et al., 2006a). Positive and negative controls were used for each PCR assay. The PCR products were verified by DNA-DNA hybridisation with internal probes as previously described (Soge et al., 2006a). The ampicillin resistance genes blaTEM and blaSHV were identified as previously described (Soge et al., 2006b). The PCR primers and probes are listed in Table 1.
Plasmids isolation
Plasmid DNA was extracted using a modified alkaline lysis procedure and electrophoresed through 0.7% agarose gel with the E. coli V517 (58 kb); R1 (100 kb) and R478 (274.5 kb) used as markers for plasmid size estimation as previously described (Soge et al., 2006a).
Conjugal gene transfer
Mating experiments were carried out using E. coli HB101 and/or E. coli DH5α as recipients. Both isolates have previously been selected for chromosomal resistance to streptomycin (500 mg/ℓ), fusidic acid (25 mg/ℓ), nalidixic acid (25 mg/ℓ), and rifampicin (25 mg/ℓ) [Strepr, Fusr, Rifr, and Nalr] as recipients (Soge et al., 2006a). E. coli 302, E. coli 304 (Apr, Tcr, SXTr), E. coli 387 (Apr, Tcr, Ermr), Enterobacter cloacae 325, Morganella morganii 390, Serratia odorifera 394, Citrobacter sp. 283 and Proteus sp. 222 were used as donors. Transconjugants were selected on one of the following; Luria-Bertani agar (Difco Laboratories, Kansas, MO, USA) supplemented with rifampicin (25 mg/ℓ) plus ampicillin (50 mg/ℓ), streptomycin (250 mg/ℓ) plus ampicillin (50 mg/ℓ), rifampicin (25 mg/ℓ) plus tetracycline (20 mg/ℓ), rifampicin (25 mg/ℓ) plus erythromycin (128 mg/ℓ)] as described previously (Soge et al., 2006a). Mating experiments were done with a ratio of 1:1 of the donor to recipient with E. coli as the parental strains and ratios of 1:10, 1:50, 1:100, and 1:200 donor to recipient with Enterobacter cloacae, Morganella morganii, Serratia odorifera, Citrobacter sp. and Proteus sp. parental strains. Transconjugants were confirmed biochemically and their antibiotic resistance genes verified by PCR assays followed by DNA-DNA hybridisation of the PCR products using radio-labelled internal probes as previously described (Soge et al., 2006a).
Results
Antibiotic susceptibility and antibiotic resistance genes
Of the 52 Gram-negative isolates examined, 26 were susceptible to all antibiotics and 26 isolates were resistant to > 1 of the antibiotics examined, with 14 isolates exhibiting multidrug resistance. Apr was found in 25 of the 26 antibiotic resistant isolates including 6 E. coli, 5 Enterobacter spp., 1 Salmonella paratyphi, 2 Salmonella spp., 2 Klebsiella spp., 2 Citrobacter spp., 2 Proteus spp., 3 M. morganii, one each of P. rettgeri and S. odorifera. All Apr isolates were susceptible to ceftazidime, cefotaxime, aztreonam, cefepime with MIC < 4 mg/ℓ for all 4 drugs tested suggesting that they did not carry genes coding for extended-β-lactamases. Nineteen of Apr isolates carried a blaTEM gene, 1 E. coli, Klebsiella sp. and M. morganii carried both blaTEM and blaSHV genes, and 1 Klebsiella sp. carried a blaSHV gene (Table 2). Two isolates did not carry either the blaTEM, and blaSHV or blaCTX-M genes.
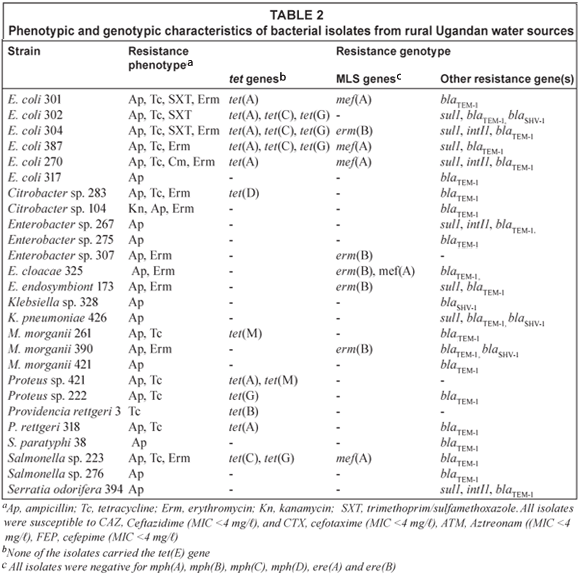
Twelve isolates were tetracycline resistant (Tcr) with 7 isolates carrying a single tet gene. Three isolates carried the tet(A) gene and 1 isolate each with the tet(B), tet(D), tet(G) or the tet(M) gene. The remaining 5 isolates carried multiple tet genes and included 3 isolates with the tet(A), tet(C), and tet(G) genes, 1 isolate with both the tet(A) and tet(M) genes, and 1 isolate with both the tet(C) and tet(G) genes. Eleven isolates had erythromycin MIC >128 mg/ℓ and 5 of these isolates carried macrolide [mef(A)] and/or macrolide-lincosamide-streptogramin B [erm(B)] resistance genes, while all 11 isolates were negative for mph(A), mph(B), mph(C), mph(D), ere(A) and ere(B). Eight isolates carried the sul1 gene and 4 of these isolates also carried the int1 gene suggestive of an integron. One isolate was chloramphenicol resistant though the resistance gene was not determined.
Plasmids
Thirteen isolates, including 5 E. coli, 3 Enterobacter spp., 1 Klebsiella sp., 1 Proteus sp., 1 Providencia rettgeri sp., and 2 Salmonella spp., carried 1 to 3 plasmids with molecular weights of 3 to 150 kb (data not shown) while the remaining 13 antibiotic resistant isolates, had no detectable plasmid DNA using the single plasmid extraction method.
Conjugation transfer experiments
Three E. coli; E. coli 302, E. coli 304 , and E. coli 387, along with E. cloacae 325, M. morganii 390, S. odorifera 394, and Citrobacter sp. 283 and Proteus sp. 222 were used as donors with recipient E. coli HB101 and/or E. coli DH5α. All 3 E. coli donors transferred Apr and Tcr at a frequency ranging from 10-5 to 10-6 /recipient. Enterobacter cloacae 325 carrying 2 large plasmids (>58 kb) and other donors M. morganii 390, S. odorifera 394, and Citrobacter sp. 283 and Proteus sp. 222, which had no detectable plasmid gave no transconjugants (transfer frequency <10−9) using 4 different ratio combinations of donor to recipient (1:10, 1:50, 1:100, 1:1200) with both HB101 and DH5α as recipients in repeated experiments. All the transconjugants from the E. coli to E. coli mating were Apr Tcr and carried a single large plasmid of molecular weights 58 kb, 65 kb, 75 kb for E. coli 304, E. coli 302 and E. coli 387 respectively (Table 3).
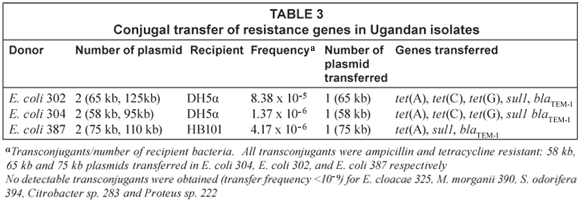
All transconjugants carried the blaTEM conferring resistance to Apr, the sul1 gene but differed in the tet genes that were transferred to the transconjugants. E. coli 302, E. coli 304 donors transferred tet(A), tet(C) and tet(G) genes, while E. coli 387 donor transferred only the tet(A) gene.
Discussion
In the current study, 26 out of 52 water-borne Ugandan Gram-negative bacteria were antibiotic resistant and 14 were multidrug resistant. None of the isolates carried extended-spectrum β-lactamases, and 4 isolates carried genes consistent with a Class 1 integron. Apr was the most common resistance phenotype among the Ugandan isolates. In the E. coli isolates, Apr was associated with conjugative plasmids [58, 65, 75 kb], while the other Gram-negative isolates appeared to have chromosomally mediated Apr genes, which we were unable to conjugally transfer in the study. None of the antibiotic-resistant isolates were resistant to cefotaxime and ceftazidime, while clinical human African enteric bacteria are usually resistant to cephalosporins and often carry multiple β-lactamases encoded by blaCTX-M, blaCMY, and blaVIM type genes in addition to blaSHV and blaTEM-1 (Frank et al., 2006; Gray et al., 2006; Ktari et al., 2006; Soge et al., 2006b). Two isolates from the current Ugandan study, Proteus 421 and Enterobacter 307, were Apr but did not carry either blaSHV , blaTEM-1, blaCTX-M genes. Similarly, 2 Citrobacter spp. 283 and 104 had erythromycin MICs > 128 mg/ℓ but did not carry any of the 8 common MLS genes examined. One SXTr E. coli did not carry the sul1 gene while all 12 Tcr isolates carried previously characterised tet genes.
Three of the 8 isolates used as donors were able to conjugally transfer antibiotic resistance to the recipients. The resulting transconjugants carried a single plasmid, which carried 1-3 tet genes, sul1 and blaTEM-1genes. The blaSHV-1 was not associated with these conjugative plasmids and was not transferred in the experiments. In contrast, the remaining 5 donor isolates did not generate transconjugants.
In one Nigerian study, small differences in antibiotic susceptibility between the clinical bacteria and bacteria isolated from soil, industrial effluent, food and drinking water were found (Lateef et al., 2005), suggesting the possibility that clinical and water-borne bacteria may have the same level of antibiotic resistance. Little work on the level of antibiotic carriage in Ugandan Gram-negative bacteria is available; however, in a 1998 paper, the authors reported that 92% of the endemic Shigella isolated in Mbarara, Uganda were resistant to cotrimoxazole and 58% were resistant to ampicillin (Legros et al., 1998). Previous studies on water enteric bacteria from other African countries also showed higher rates of antibiotic resistance than those found in this study. For instance, two separate Nigerian studies have found high levels of antibiotic-resistant (93 to 94%) water-borne enteric bacteria, isolated from communal well water, and from the lower Niger Delta River (Ibiebele et al., 1989; Ogan et al., 1993). Lin et al. (2004) characterised 113 enteric bacteria, including E. coli, Klebsiella sp., C. freundii, Enterobacter spp., S. marcesens, isolated from the Mhlathuze River in South Africa and found that 94.7 % of these bacteria were resistant to > 1 antibiotic and 75.2% of the isolates were multidrug resistant. The low level of Class 1 integrons found in the Ugandan isolates differs from recent studies of Class 1 integrons where 12% of the E. coli isolated from a remote community of Guarani Indians in Bolivia, and 40% of 100 multi-drug resistant Gram-negative bacteria, from the River Torsa, India, carried Class 1 integrons (Pallecchi et al., 2007; Mukherjee and Chakraborty, 2006).
Unregulated use of antibiotics in agriculture, animal husbandry, and medical therapy has been a major influence in Africa and is often cited as a major reason why there is high prevalence of multidrug resistant bacteria (Okeke et al., 2007). Why the Ugandan water bacteria in this study differ from other studies from remote areas in Africa is intriguing. However, this study does illustrate that the level of antibiotic resistance found in water-borne Gram-negative bacteria can vary among Gram-negative bacteria isolated from various remote parts of the world where exposure to western medicine and antibiotics is minimal.
Acknowledgements
This study was presented in part at the 108th American Society for Microbiology General Meeting, June 1-5, 2008, Boston Massachusetts (Abstract A-115).
References
CLINICAL AND LABORATORY STANDARDS INSTITUTE (CLSI) (2003a) Performance Standards for Antimicrobial Disk Susceptibility Tests (8thedn.): Approved Standard M2-A8. CLSI, Wayne, PA, USA. [ Links ]
CLINICAL AND LABORATORY STANDARDS INSTITUTE (CLSI) (2003b) Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically (6th edn.): Approved Standard M7-A6. CLSI, Wayne, PA, USA. [ Links ]
FRANK T, ALERT G, GAUTIER V, TALARMIN A and BERCION R (2006) Extended-spectrum beta-lactamase-producing Enterobacteriaceae, Central African Republic. Emerg. Infect. Dis. 12 863-865. [ Links ]
GRAY KJ, WILSON LK, PHIRI A, CORKILL JE, FRENCH N and HART CA (2006) Identification and characterization of ceftriaxone resistance and extended-spectrum beta-lactamases in Malawian bacteraemic Enterobacteriaceae. J. Antimicrob. Chemother. 57 661-665. [ Links ]
IBIEBELE DD and SOKARI TG (1989) Occurrence of drug-resistant bacteria in communal well water around Port Harcourt, Nigeria. Epidemiol. Infect. 103 193-202. [ Links ]
KTARI S, ALERT G, MNIF B, GAUTIER V, MAHJOUBI F, BEN JM, BOUAZIZ M and HAMMAMI A (2006) Emergence of multidrug-resistant Klebsiella pneumoniae isolates producing VIM-4 metallo-beta-lactamase, CTX-M-15 extended-spectrum beta-lactamase, and CMY-4 AmpC beta-lactamase in a Tunisian university hospital. Antimicrob. Agents Chemother. 12 4198-4201. [ Links ]
LATEEF A, OLOKE JK, and GUEGUIMKANA EB (2005) The prevalence of bacterial resistance in clinical, food, water and some environmental samples in Southwest Nigeria. Environ. Monit. Assess. 100 59-69. [ Links ]
LEGROS D, OCHOLA D, LWANGA N, and GUMA G (1998) Antibiotic sensitivity of endemic Shigella in Mbarara, Uganda. East Afr. Med. J. 75 160-161. [ Links ]
LIN J, BIYELA, PT and PUCKREE T (2004) Antibiotic resistance profiles of environmental isolates from Mhlathuze River, KwaZulu-Natal (RSA). Water SA 30 (1) 23-28 http://www.wrc.org.za/archives/watersa%20archive/2004/Jan-04/4.pdf. [ Links ]
MAJALIJA S, SEGAL H, EJOBI F and ELISH GB (2008) Shiga toxin gene-containing Escherichia coli from cattle and diarrheic children in the pastoral systems of southwestern Uganda. J. Clin. Microbiol. 46 352-354. [ Links ]
MARCH SB and RATNAM S (1986) Sorbitol-MacConkey medium for detection of Escherichia coli O157:H7 associated with hemorrhagic colitis. J. Clin. Microbiol. 23 869-872. [ Links ]
MIRANDA CD, KEHRENBERG C, ULEP C, SCHWARZ S and ROBERTS MC (2003) Diversity of tetracycline resistance genes in bacteria from Chilean salmon farms. Antimicrob. Agents Chemother. 47 883-888. [ Links ]
MUKHERJEE S and CHAKRABORTY R (2006) Incidence of class 1 integrons in multiple antibiotic-resistant Gram-negative copiotrophic bacteria from the River Torsa in India. Res. Microbiol. 157 220-226. [ Links ]
OGAN MT and NWIIKA DE (1993) Studies on the ecology of aquatic bacteria of the lower Niger Delta: multiple antibiotic resistance among the standard plate count organisms. J. Appl. Bacteriol. 74 595-602. [ Links ]
OJO KK, ULEP C, VAN KIRK N, LUIS H, BERNARDO M, LEITAO J and ROBERTS MC (2004) The mef(A) Gene predominates among seven macrolide resistance genes identified in Gram-negative strains representing 13 genera, isolated from healthy Portuguese children. Antimicrob. Agents Chemother. 48 3451-3456. [ Links ]
OKEKE IN, ABODERIN OA, BYARUGABA DK, OJO KK and OPINTAN JA (2007) Growing problem of multidrug-resistant enteric pathogens in Africa. Emerg. Infect. Dis. 13 1640-1646. [ Links ]
PALLECCHI L, LUCCHETTI C, BARTOLONI A, BARTALESI F, MANTELLA A, GAMBOA H, CARATTOLI A, PARADISI F and ROSSOLINI GM (2007) Population structure and resistance genes in antibiotic-resistant bacteria from a remote community with minimal antibiotic exposure. Antimicrob. Agents Chemother. 51 1179-1184. [ Links ]
PEARSON AL, ROBERTS MC, SOGE OO, IVANOVA I, MAYER JD and MESCHKE JS (2008) Utility of EC 3MTM petrifilmsTM and sanitary surveys for source water assessment in south-western Uganda. Water SA 34 (2) 279-283. http://www.wrc.org.za/downloads/watersa/2008/April/2227.pdf [ Links ]
SOGE OO, ADENIYI BA and ROBERTS MC (2006a) New antibiotic resistance genes associated with CTX-M plasmids from uropathogenic Nigerian Klebsiella pneumoniae. J. Antimicrob. Chemother. 58 1048-1053. [ Links ]
SOGE OO, QUEENAN AM, OJO KK, ADENIYI BA and ROBERTS MC (2006b) CTX-M-15 extended-spectrum beta-lactamase from Nigerian Klebsiella pneumoniae. J. Antimicrob. Chemother. 57 24-30. [ Links ]
Received 13 January 2009; accepted in revised form 20 February 2009.
* To whom all correspondence should be addressed.
206-543-8001; fax: 206-543-3873;
e-mail: marilynr@u.washington.edu














