Servicios Personalizados
Articulo
Indicadores
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares en Google
Similares en Google
Compartir
Journal of the South African Veterinary Association
versión On-line ISSN 2224-9435
versión impresa ISSN 1019-9128
J. S. Afr. Vet. Assoc. vol.82 no.2 Pretoria ene. 2011
ARTICLE ARTIKEL
Detection and characterisation of papillomavirus in skin lesions of giraffe and sable antelope in South Africa
E van DykI,*; A-M BosmanII; E van WilpeIII; J H WilliamsIV; R G BengisV; J van HeerdenVI ; E H VenterII
IDepartment of Companion Animal Clinical Studies, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, Onderstepoort, 0110 South Africa
IIDepartment of Veterinary Tropical Diseases, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, Onderstepoort, 0110 South Africa
IIISection of Electron Microscopy, Department of Anatomy and Physiology, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, Onderstepoort, 0110 South Africa
IVSection of Pathology, Department of Paraclinical Sciences, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, Onderstepoort, 0110 South Africa
VNational Department of Agriculture, PO Box 12, Skukuza, Kruger National Park, 1350 South Africa
VIKimberley Veterinary Clinic, 16 Dalham Road, Kimberley, 8301 South Africa
ABSTRACT
Papillomavirus was detected electron microscopically in cutaneous fibropapillomas of a giraffe (Giraffa camelopardalis) and a sable antelope (Hippotragus niger). The virus particles measured 45 nm in diameter. Histopathologically, the lesions showed histopathological features similar to those of equine sarcoid as well as positive immunoperoxidase-staining of tissue sections for papillomavirus antigen. Polymerase chain reaction (PCR) detected bovine papillomavirus (BPV) DNA. Bovine papillomavirus-1 was characterised by real-time PCR in the sable and giraffe, and cloning and sequencing of the PCR product revealed a similarity to BPV-1. As in the 1st giraffe, the lesions from a 2nd giraffe revealed locally malignant pleomorphism, possibly indicating the lesional end-point of papilloma infection. Neither virus particles nor positively staining papillomavirus antigen could be demonstrated in the 2nd giraffe but papillomavirus DNA was detected by real-time PCR which corresponded with BPV-1 and BPV-2.
Keywords: cutaneous fibropapillomas, Giraffa camelopardalis, giraffe, Hippotragus niger, malignant pleomorphism, papillomavirus, sable antelope.
INTRODUCTION
Papillomaviruses are classified in the family Papillomaviridae6. This large family of animal and human viruses generally infects epithelial cells causing hyperproliferative lesions. Papillomavirus also has oncogenic potential18, infecting cutaneous and mucous epithelia in a variety of hosts through cuts or abrasions and induces the formation of papillomas or warts by targeting the keratinocyte, the viral growth accompanying the steady maturation of the cell to the surface31. These tumours are generally benign and self-limiting and spontaneously regress with the animal recovering completely, but occasionally benign tumours may persist and become malignant by progressing to squamous cell carcinomas10.
Bovine papillomavirus induces exophytic papillomas of cutaneous or mucosal epithelia in cattle. Papillomas are benign tumours which generally regress uneventfully; however, they do occasionally persist and provide the focus for malignant transformation to squamous cell carcinomas, particularly in the presence of environmental co-factors. This has been experimentally demonstrated in cancer of the urinary bladder and upper alimentary tract in cattle feeding on bracken fern (Pteridium aquilinum)39. In equines BPV causes a locally invasive, fibroblastic skin tumour of horses, donkeys, mules23 and zebras37. These tumours appear as different clinical entities and can be classified into 6 clinical types16. This cross-species infection of Equidae by BPV 1 and 2 is the only record until recently of a papillomavirus crossing the species barrier, the resulting tumour being known as a sarcoid22. Bovine papillomavirus has been demonstrated in lesions in the water buffalo (Bubalis bubalis)28 and bison (Bison bonasus)21.
Earlier it was thought that apart from BPV 1 and 2, papillomaviruses are strictly species-specific and only the natural host is infected. Even under experimental conditions papillomaviruses did not appear to infect any host other than the natural one. Lesions usually attributed to papillomavirus infection have been most extensively studied in cattle2,9,14 and horses3,7,11,24,25, but at least 50 mammalian species have been confirmed as being infected by species-specific papillomaviruses33. Papillomaviruses appear to be widespread and have been found in a large number of vertebrate species and are assumed to have co-evolved with their hosts4,5. Virtually all mammalian species are hosts for one or more papillomaviruses33.
CASE HISTORIES
Giraffes 1 and 2
An adult female giraffe (Giraffa camelopardalis) (Giraffe 1) with extensive lesions in the skin of the dorsal neck and back was observed near Shingwedzi rest camp in the northern part of the Kruger National Park (KNP), South Africa (Fig. 1). She was in a group of 6 animals, 1 of which was a bull. The other animals in the group showed no lesions. This area is a natural habitat for giraffe as they occur in a variety of dry savannas ranging from scrub to woodland29. Approximately 1 year later a 2nd affected giraffe (Giraffe 2), was observed in the vicinity of Skukuza rest camp and the Kruger Gate in the southern part of the KNP (Fig. 2). This was also an adult female and part of a group of 5 of mixed sex and age. She was the only animal in the group exhibiting wart-like lesions that were especially prominent in the skin of her head and neck, but had also spread to the rest of her body.


Sable
A group of 30 sable antelope (Hippotragus niger) was kept as a breeding herd on a game farm in the Kimberley district, Northern Cape Province, South Africa. This arid area of South Africa is not the natural habitat of sable as they are a savanna woodland species, but such game species are sometimes translocated to private land and reserves outside their normal range. One cow was lame in the right hind leg and a wart-like lesion of 60 × 60 mm was present in the skin of the lateral aspect of the distal part of the 2nd phalanx proximal to the right hind hoof. After surgical removal, growth recurred at the original site and other well-defined wart-like lesions were also noticed in the skin of the right shoulder and lip.
The purpose of this study was to determine electron microscopically if BPV was present in the lesions of these animals and, if so, to detect and distinguish between BPV-1 and BPV-2 DNA using real-time PCR37.
MATERIALS AND METHODS
Sample collection
Giraffe 1 was shot in May 2007 in the vicinity of the Shingwedzi rest camp (23º97'42"S, 31º42'58"E). Samples of the lesions were collected for PCR by excising pieces of tissue 10×3×3mmin size. These were stored in sterile tubes at 4 ºC until analysis. Lesion samples for histopathological examination were stored in duplicate in 10 % neutral buffered formalin. The 2nd giraffe (Giraffe 2) was shot near Skukuza rest camp (24º59'16"S, 31º34'32"E) in October 2008 and samples were collected as described above. The sable antelope was anaesthetised and the lesion on the pastern surgically removed in an effort to relieve the lameness. Samples of the lesion were taken as described above.
Electron microscopy
Formalin-fixed skin samples from both giraffes were prepared for transmission electron microscopy (TEM) according to standard procedures. Ultra-thin sections were stained with uranyl acetate and lead citrate, and examined in a Philips CM 10 transmission electron microscope operated at 80 kV.
An unfixed skin sample of the sable cow was prepared for TEM by grinding the tissue in a mortar with a pestle in a small volume of sterile water and the mixture was centrifuged at 13 000 rpm for 45 min. The resultant pellet was re-suspended in water, stained with 3 % phosphotungstic acid and a drop of the suspension placed onto a formvar-and carbon-coated grid for examination. A relevant area was also retrieved from the histological wax block (see below) of the skin sample from the pastern and was treated with1%OsO4 in xylene and embedded in an epoxy resin.
Histopathology
The formalin-fixed samples from both giraffes and the sable were dehydrated and paraffin wax-embedded for routine histological processing. All tumour sections were stained with haematoxylin and eosin for light microscopy, as well as immunohistochemically for evaluation of papillomavirus, using a polyclonal rabbit anti-papillomavirus antibody against chemically disrupted BPV-1, by the avidin-biotin technique13. A section of a bovine fibropapilloma was used as a positive control.
DNA extraction
DNA was extracted from 25 mg of lesion tissues from each animal using the QIAamp® DNA extraction kit (Southern Cross Biotechnologies) according to the manufacturer 's instructions. Extracted DNA was eluted in 100ℓ elution buffer and stored at 4 ºC until further analysis.
Real-time PCR
The hybridisation probe real-time PCR assay, as described by van Dyk et al., (2009)37 was used for the detection and differentiation of BPV-1 and BPV-2 DNA in the giraffe (1 & 2) and sable lesions. The primers (forward primer: 5'-CAA AGG CAA GAC TTT CTG AAA CAT-3', and reverse primer: 5'-AGA CCT GTA CAG GAG CAC TCA A-3') amplified a 244 and 247 bp region of the E5 ORF of BPV-1 and BPV-2 respectively. For the specific detection of BPV-1 amplicons, hybridisation probes BPV-1 anchor (5'-ACT GG TGT ACT ATG CCA AAT CTA TGG TTT CTA TTG-Fluor-3') and BPV-1 sensor (5'-LC-640-CTT GGG ACT AGT TGC TGC AAT GCA ACT-Pho-3') were used, and for the detection of BPV-2 amplicons, hybridisation probes BPV-2 anchor (5'-TTT AAT CAC TGC CAT TTG TTT TTT TCA TAT CTC GT-Fluor-3') and BPV-2 sensor (5'-LC705-AGG CAT ACT ATG CCG AAT CTA TGG TTT CTA TTG TT-Pho-3') were used.
All primers and fluorescent-labelled hybridisation probes were manufactured by Metabion International AG (Martinsried, Germany).
The PCR amplification mixture and reaction conditions were as previously described37. Plasmids obtained from Giraffe 1 and the sable were used as template DNA (2 µl) and genomic DNA from Giraffe 2 was used. Fluorescence was measured at 640 and 705 nm and the results were analysed with the Roche LightCycler® Software v4.0. Specimens, from positive zebra sarcoid (BPV-1 and BPV-2)37 were included as positive controls.
Cloning and sequence analysis
Conventional PCR was used for the amplification of a 637 bp DNA fragment of a region of the E5 ORF of both BPV-1 and BPV-2. The primers (F_3610: 5'-GCT AACCAG GTAAAG TGCTAT C-3'; R_4247: 5'-TGC TTG CAT GTC CTG TAC AGG T-3'), PCR amplification mixture and reaction conditions were as previously described37. However, a nested PCR step was included using 1 µℓ of the 1st round amplicons as PCR template in Giraffe 1 and the sable. The amplicons obtained were purified, cloned into the pGEM®-T easy vector (Promega pGEM-T Easy Vector System, Promega, Madison, USA), and transformed into competent JM 109 E. coli cells (Promega, Madison, USA). Recombinant plasmid DNA was isolated, directly sequenced using the ABI BigDyeTM Terminator Cycle Sequencing Ready Reaction kit (PE Applied Biosystems) and analysed on an ABI 3100 sequencer at Inqaba Biotec (Pretoria, South Africa). Sequencing data were assembled and edited with the GAP4 programme of the Staden package (version 1.6.0 for Windows)30 and aligned with published sequences of related genera using ClustalX (version 1.81 for Windows). A BLAST search was performed using the Blastn algorithm. Phylogenetic trees were constructed using the maximum parsimony method (Mega 3.0 software package) 17 in combination with the bootstrap method12 (1000 replicates/tree for distance methods and 100 replicates/tree for parsimony methods). Human papillomavirus type 16 (K02718) was used as an outgroup.
RESULTS
Macroscopic lesions
There were extensive skin lesions on the head, neck and back of the 2 adult female giraffes (Giraffe 1 and 2) in the Kruger National Park (Figs 1, 2).
Electron microscopy
Negatively-stained spherical papillomavirus particles with a diameter of 52 nm and distinct capsomeres were present in the unfixed skin sample of the sable cow (Fig. 3). The nuclei of the stratum granulosum of both the sable cow and Giraffe 1 contained numerous randomly scattered papillomavirus particles measuring 45 nm in diameter (Figs 4, 5). The nuclei of infected cells displayed irregular clumps of condensed chromatin. No virus was demonstrated in samples obtained from Giraffe 2.



Histopathology
The skin lesions of Giraffe 1 were fibropapillomatous in nature and those of the sable resembled equine verrucose sarcoid. Small nuclei in the stratum granulosum of both animals stained immunohistochemically positive for papillomavirus antigen. The lesions of Giraffe 2 were more sarcoid-like, expansile and infiltrative in the dermis. Lesions of both giraffes showed evidence of malignancy. Scattered, large fibroblasts with bizarre large nuclei, some of which appeared multinucleated, occurred throughout the dermal tumour tissue with those of Giraffe 2 being more prevalent. Immunohistochemical staining of sections from Giraffe 2 did not demonstrate papillomavirus antigen. The detailed histopathology is described and illustrated elsewhere38.
Real-time PCR
The primers used in the real-time PCR assay37 amplified either a 244 bp or 247 bp region of the E5 ORF of BPV-1 and BPV-2, respectively. Two separate hybridisation probes sets were used in a multiplex format for the specific detection of BPV-1 and BPV-2 DNA. For the detection of BPV-1 DNA, an increase in fluorescence is expected at 640 nm as well as 2 BPV-1-specific melting peaks at 62.90 ± 1.24 ºC and 68.17 ± 0.71 ºC. Similarly, for the detection of BPV-2 DNA, an increase in fluorescence at 705 nm is expected as well as 2 BPV-2-specific peaks at 58.86 ± 0.60 ºC and 64.06 ± 0.59 ºC.
Bovine PV-1-positive material from sarcoids of Cape mountain zebras induced a graph with melting peaks showing a similar configuration to those of material from Giraffe 1 and the sable. Giraffe 2 demonstrated fluorescence at 58 ºC as well as 68 ºC at 640 nm (Fig. 6) indicating both BPV-1 and BPV-2.
Cloning and sequencing
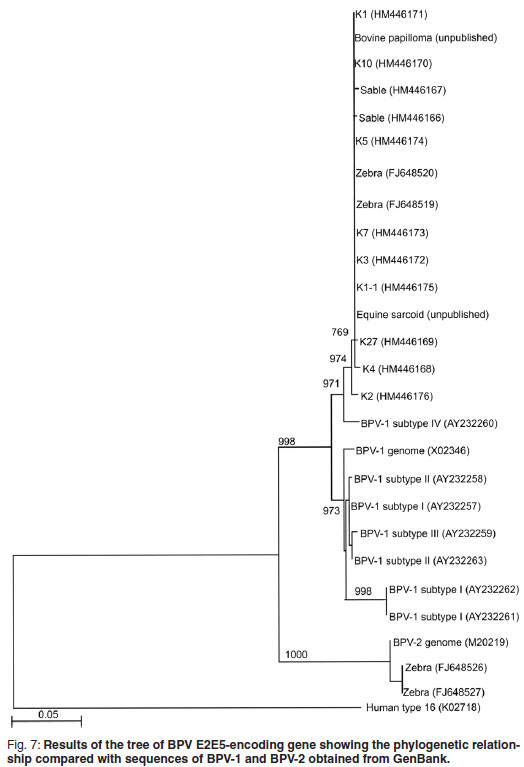
Fragments, 636 bp in size, of the E5 ORF of the sable and Giraffe 1 samples were amplified, cloned and sequenced to confirm that the amplicons obtained were indeed related to BPV. Sequences were edited and truncated to a length of 478 bp. Six [K10 (HM446170); K1 (HM446171); K3 (HM446172); K7 (HM446173); K5 (HM446174); K1-1 (HM446174)] of the 9 Giraffe 1 clones had identical sequences over the 478 bp region and a BLAST search revealed that these sequences were identical to those of the BPV -1 E2 and E5 protein encoding genes previously reported37 to occur in Cape mountain zebras (FJ648519 to FJ648528). The other 3 clones [K4 (HM446168); K27 (HM446169); K2 (HM446176)] were not identical to each other, but the BLAST search showed the highest similarity (~99 %) with the BPV type 1 E2 and E5 protein-encoding genes found in Cape mountain zebras (FJ648519 to FJ648528). Only 2 sequences were obtained from sable clones, Sable 4 (HM 446166) and Sable 2 (HM 44167) and they showed 2 nucleotide differences within the 478 bp fragment. Again, the BLAST search showed the highest similarity (~99 %) with the BPV-E2 and E5 protein encoding genes found in Cape mountain zebras. All sequences obtained from Giraffe 1 and sable samples were closely related (~97 %) to the BPV-1 subtype IV E5 protein-encoding gene (accession number AY232260) detected in equine sarcoid11. A phylogenetic tree was constructed and the results were concurrent with the BLAST results obtained (Fig. 7). In the case of Giraffe 2, no good-quality sequence data could be obtained and the DNA could only be detected by real-time PCR.
DISCUSSION
Numerous descriptions of papillomatous epithelial growths in the skin of many species of mammals have been reported 15,21,26-28,32,33,36. Epithelial growths were reported in giraffes in the Kruger National Park in 197835. They are characterised variably by a conspicuous amount of fibrous connective tissue underlying hyperplastic epithelium and are commonly associated with infection with a papillomavirus. This study documents the presence of BPV and/or DNA in such growths in the skin of the 2 giraffes and a sable antelope.
Papillomatosis is a naturally-occurring, generally species-specific infectious disease. The lesions are regarded as hyperplastic or a form of benign neoplasia as they do not metastasise internally20. Although bovine-associated papillomavirus DNA is consistently found in the sarcoid lesions of the horse1,19 and zebra37, papillomavirus particles have not been demonstrated and the disease is therefore considered to be a non-productive infection in which the viral DNA exists episomally in these species. Most fibropapillomas (sarcoids) in horses contain identifiable BPV DNA of either type 1 or 23,7,11,24,25,34,40. The E1 gene of the papillomavirus identified in sarcoids of donkeys is very similar to BPV-1 and it has been proposed that it is a subtype of BPV-125.In the present study the papillomavirus particles that were demonstrated in the wartlike lesions of the sable antelope and Giraffe 1 are considered to be closely related to BPV-I.
Ultrastructurally, virus particles have also been demonstrated in skin papillomas in an impala and a giraffe in Kenya15 but sequencing was not done. In the water buffalo28 and the bison21 virus has also been demonstrated electron microscopically, and following its sequencing it was reported to be homologous to the LCR of BPV-1 (prototype sequence, accession number X02346) in the water buffalo28 while the DNA sequence of 413 bp amplicon derived from the European bison compared with the consensus sequence of BPV-2 (GenBank Accession number AY300818)21. The virus was also demonstrated in both species of animals by electron microscopy21,28.
The real-time PCR method37 demonstrated the presence of BPV-1 in the sable and Giraffe 1 and was confirmed by cloning and sequencing. In the 2nd giraffe, no good-quality sequences could be obtained, although, real-time assay demonstrated the presence of both BPV-1 and BPV-2. This can be attributed to the high sensitivity of the real-time PCR assay used, which is capable of demonstrating the virus at less than 1.5 gene copies37. Another possible reason why the virus could not be demonstrated was that the papilloma had undergone a degree of cellular malignant transformation, with more pleomorphic fibroblasts being present in the lesions than in Giraffe 1. The structural integrity of the virus appeared to be lost and the virus or viral antigen was no longer present20.
Although the geographical areas are widely separated, the Giraffe 1 and sable clones clustered together with high similarity to BPV-1 sequences, and with some of the zebra sequences reported previously37. None of the sable or Giraffe 1 sequences reported here were of the BPV-2 group, and only some of the previously reported zebra samples37 clustered in BPV-2. The human papillomavirus type 16 outgroup clustered completely differently (Fig. 7).
Clinical manifestation of latent infection in the 3 animals described here may have been provoked by stress. In the case of the sable antelope, not being in its typical habitat may have induced immunosuppression. In the case of the giraffes it can only be speculated that drought conditions and the resultant high tannin content of the Acacia trees as a result of over-browsing acted as a co-factor in BPV-associated carcinogenesis as seen in cattle following bracken fern ingestion8.
ACKNOWLEDGEMENTS
This work was supported by grants from the South African Veterinary Foundation, the Research and Development Fund of the University of Pretoria, and the Department of Veterinary Tropical Diseases, Faculty of Veterinary Science, University of Pretoria. The authors thank Prof. Roy Tustin for his critical comments on the manuscript.
REFERENCES
1. Amtmann E, Muller H, Sauer G 1980 Equine connective tissue tumours contain unintegrated bovine papillomavirus DNA. Journal of Virology 35: 962-964 [ Links ]
2. Anderson R A, Scobie L, O'Neil B W, Grindlay G J, Campo M S 1997 Viral proteins of bovine papillomavirus type 4 during the development of alimentary canal tumours. Veterinary Journal 154: 69-78 [ Links ]
3. Angelos J A, Marti E, Lazary S, Carmichael L E 1991 Characterization of BPV-like DNA in equine sarcoids. Archives of Virology 119: 95-109 [ Links ]
4. Antonsson A, McMillan A J 2006 Papillomavirus in healthy skin of Australian animals. Journal of General Virology 87: 3195-3200 [ Links ]
5. Bernard H-U 1994 Coevolution of papillomaviruses with human populations. Trends in Microbiology 2: 140-143 [ Links ]
6. Bernard, H-U 2006 Phylogeny and taxonomy of papillomaviruses. In Campo M S (ed.) Papillomavirus research: natural history to vaccines and beyond. Caister Academic Press, Norfolk: 11-17 [ Links ]
7. Carr E A, Théon A P, Madewell B R, Griffey S M, HitchcockME2001Bovinepapillomavirus DNA in neoplastic and nonneoplastic tissues obtained from horses with and without sarcoids in the western United States. American Journal of Veterinary Research 62: 741-744 [ Links ]
8. Campo M S 1997 Bovine papillomavirus and cancer. Veterinary Journal 154: 175-188 [ Links ]
9. Campo M S 2002 Animal models of papillomavirus pathogenesis. Virus Research 89: 249-261 [ Links ]
10. Campo M S 2006 In Campo M S (ed.) Papillomavirus research: natural history to vaccines and beyond. Caister Academic Press, Norfolk: 1-2 [ Links ]
11. Chambers G, Ellsmore V A, O'Brien P M, Reid S W J, Love S, Campo M S, Nasir L 2003 Sequence variants of bovine papillomavirus E5 detected in equine sarcoids. Virus Research 96: 141-145 [ Links ]
12. Felsenstein J 1985 Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783-791 [ Links ]
13. Haines D M, Chelack B J 1991 Technical considerations for developing enzyme immunohistochemical staining procedures on formalin-fixed paraffin-embedded tissues for diagnostic pathology. Journal of Veterinary Diagnostic Investigation 3: 101-112 [ Links ]
14. Jarret W F H, McNeal P E, Grimshaw T R, Selman I E, McIntyre W I M 1978 High incidence area of cattle cancer with a possible interaction between an environmental carcininogen and a papillomavirus. Nature 274: 215-217 [ Links ]
15. Karstad L, Kaminjolo J S 1978 Skin papillomas in an impala (Aepyceros melampus) and a giraffe (Giraffa camelopardalis). Journal of Wildlife Diseases 14: 309-313 [ Links ]
16. Knottenbelt D C 2005 A suggested clinical classification for the equine sarcoid. Clinical Techniques in Equine Practice 4: 278-295 [ Links ]
17. Kumar S, Tamura K, Nei M 2004 MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinformation 5: 150-163 [ Links ]
18. Lambert P F, Balsitis S J, Shai A, Simonson S J S, Williams S M G 2006 Transgenic mouse models for the in vivo analysis of papillomavirus oncogene function. In Campo M S (ed.) Papillomavirus research: from natural history to vaccines and beyond. Caister Academic Press, London: 213-228 [ Links ]
19. Lancaster W D 1981 Apparent lack of integration of bovine papilloma virus DNA in virus-induced equine and bovine tumor cells and virus-transformed mouse cells. Virology 108: 251-255 [ Links ]
20. Lancaster W D, Olson C 1982 Animal papillomaviruses. Microbiological Reviews 46: 191-207 [ Links ]
21. Literák I, Tomito Y, Ogawa T, Shirasawa H, Šmid B, Novotný L, Adamec M 2006 Papilloatosis in a European bison. Journal of Wildlife Diseases 42: 149-153 [ Links ]
22. Nasir L, Gault E, Morgan I M, Chambers G, Ellsmore V, Campo M S 2007 Identification and functional analysis of sequence variants in the long control region and the E2 open reading frame of bovine papillomavirus type 1 isolated from equine sarcoids. Virology 364: 355-361 [ Links ]
23. Nasir L, Reid S W J 2006 Bovine papillomaviruses and equine sarcoids. In Campo M S (ed.) Papillomavirus research: natural history to vaccines and beyond. Caister Academic Press, Norfolk: 389-397 [ Links ]
24. Otten N, von Tscharner C, Lazary S, Antczak D F, Gerber H 1993 DNA of bovine papillomavirus type 1 and 2 in equine sarcoids: PCR detection and direct sequencing. Archives of Virology 132: 121-131 [ Links ]
25. Reid S W J, Smith K T, JarrettWFH 1994 Detection, cloning and characterization of papillomaviral DNA present in sarcoid tumours of Equus asinus. Veterinary Record 135: 430-432 [ Links ]
26. Schulman F Y, Krafft A E, Janczewski T 2001 Feline cutaneous fibropapillomas: clinicopathologic findings and association with papillomavirus infection. Veterinary Pathology 38: 291-296 [ Links ]
27. Schulman F Y, Krafft A E, Janczewski T, Reupert R, Jackson K., Garner M M 2003 Camelid mucoutaneous fibropapillomas: clinicopathologic findings and association with papillomavirus. Veterinary Pathology 40: 103-107 [ Links ]
28. Silvestre O, Borzacchiello G, Nava D, Iovane G, Russo V, Vecchio D, D'Ausilio F, Gault E A, Campo M S, Paciello O 2009 Bovine Papillomavirus Type 1 DNA and E5 oncoprotein expression in water buffalo fibropapillomas. Veterinary Pathology 46: 636 [ Links ]
29. Smithers R H N 1983 Order Artiodactyla. In SmithersRHN (ed.) The mammals of the southern African subregion. University of Pretoria, Pretoria: 578-653 [ Links ]
30. Staden R, Beal K F, Bonfield J K 2000 The Staden package, 1998. Methods in Molecular Biology 132: 115-130 [ Links ]
31. Stanley M A 2006 Immunobiology of papillomaviruses. In Campo M S (ed.), Papillomavirus research: natural history to vaccines and beyond. Caister Academic Press, Norfolk: 311-319 [ Links ]
32. Sundberg J P, van Ranst M, Montali R, Homer B L, Miller W H, Rowland P H, Scott D W, England J J, Dunstan R W, Mikaelian I, Jenson A B 2000 Feline papillomas and papillomaviruses. Veterinary Pathology 37: 1-10 [ Links ]
33. Sundberg J P, van Ranst M, Jenson A B 2001 Papillomavirus infections. In Williams E S, Barker I K (eds) Infectious diseases of wild mammals. Iowa State University Press, Ames: 223-231 [ Links ]
34. Trenfield K, Spradbrow P B, Vanselow B 1985 Sequences of papillomavirus DNA in equine sarcoids. Equine Veterinary Journal 17: 449-452 [ Links ]
35. Tustin R C 1978 Cancer in wild animals. Fauna and flora (Transvaal Nature Conservation Division) 33: 21-22 [ Links ]
36. Uzal F A, Latorraca A, Ghoddusi M, Horn M, Adamson M, Kelly W R, Schenkel R 2000 An apparent outbreak of cutaneous papillomatosis in Merino sheep in Patagonia, Argentina. Veterinary Research Communications 24: 197-202 [ Links ]
37. Van Dyk E, Oosthuisen M C, Bosman A-M, Nel P J, Zimmerman D, Venter E H 2009 Detection of bovine papillomavirus DNA in sarcoid affected and healthy free-roaming zebra (Equus zebra) populations in South Africa. Journal of Virological Methods 158: 141-151 [ Links ]
38. Williams J H, van Dyk E, Nel P J, Lane E, van Wilpe E, Bengis R G, de Klerk-Lorist L-M, van Heerden J 2011 Pathology and immunohistochemistry of papillomavirus-associated cutaneous lesions in Cape mountain zebra, giraffe, sable antelope and African buffalo in South Africa. Journal of the South African Veterinary Association 82: 97-106 [ Links ]
39. Wosiacki S R, Barreiro A B, Alfieri A F, Alfieri A A 2005 Semi-nested PCR for detection and typing of bovine papillomavirus type 2 in urinary bladder and whole blood from cattle with enzootic haematuria. Journal of Virological Methods 43: 5428-5434 [ Links ]
40. Yuan Z, Philbey A W, Gault E A, Campo M S, Nasir L 2007 Detection of bovine papillomavirus type 1 genomes and viral gene expression in equine inflammatory skin conditions. Virus Research 124: 245-249 [ Links ]
Received: February 2011.
Accepted: April 2011.
* Author for correspondence. E-mail: enettevd@gmail.com














