Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
Journal of the South African Veterinary Association
On-line version ISSN 2224-9435
Print version ISSN 1019-9128
J. S. Afr. Vet. Assoc. vol.81 n.2 Pretoria Jan. 2010
ARTICLE ARTIKEL
Evaluation of PCR assays for the detection of Campylobacter fetus in bovine preputial scrapings and the identification of subspecies in South African field isolates
T SchmidtI,*; E H VenterII; J A PicardII
IAllerton Provincial Veterinary Laboratory, Private Bag X2, Cascades, 3202 South Africa
IIDepartment of Veterinary Tropical Diseases, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, Onderstepoort, 0110 South Africa
ABSTRACT
As a result of the high lability and slow growth of Campylobacter fetus subspecies, the laboratory diagnosis of bovine genital campylobacteriosis has always been difficult. This is especially true under South African conditions, where farms are far apart, laboratories are only present in major centres and there are high ambient temperatures. In order to overcome the shortcomings associated with traditional diagnostic methods, the implementation of a molecular assay was sought. This work describes how a previously published PCR assay (MG3F/ MG4R primers) was adapted, optimised and applied in the diagnostic laboratory to test preputial samples directly for the presence of Campylobacter fetus. Field evaluation of the assay revealed an analytical sensitivity and specificity of 85.7 % and 99 %, respectively. Subsequent genotyping and phenotyping of a diverse collection of South African field isolates revealed that South Africa has an unexpected and previously unreported high incidence of Campylobacter fetus subsp. venerealis biovar intermedius strains. These strains were not identified correctly by the subspecies-specific primer set evaluated. Until such time that cost-effective genotyping methods are available to diagnostic laboratories in South Africa, and other countries with these atypical Campylobacter fetus subsp. venerealis strains, the need for bacterial culture will persist. Identification to subspecies level of isolates at present remains dependent upon a single phenotypic criterion, namely tolerance to 1 % glycine.
Keywords: bovine genital campylobacteriosis, Campylobacter fetus fetus, Campylobacter fetus venerealis biovar intermedius, culture, polymerase chain reaction.
INTRODUCTION
Genital campylobacteriosis is a transmissible genital disease of cattle caused by the bacterium Campylobacter fetus subsp. venerealis (Cfv). The disease is characterised by temporary infertility of female cattle, early embryonic mortality, irregular oestrus cycles, delayed conception and infrequent abortions6. Most cows recover from infection and oestrus cycles will recommence after a few months but the herd experiences a decreased pregnancy rate and a prolonged calving season8. Infected bulls usually show no clinical signs of infection but become carriers and infect females at service2.
Campylobacteriosis is of considerable economic importance to the cattle industry worldwide1 and is considered to be one of the most important infectious causes of poor calving rates in southern Africa. Regional veterinary laboratories in South Africa and Namibia have reported between 0 and 12 % prevalences based on bacterial isolations made from bull sheath wash submissions. One particular study conducted on bulls which were communally grazed, reported a substantially higher prevalence of almost 29 %14.Itis believed that the availability of vaccines and artificial insemination practices have decreased the prevalence of the disease15 but many researchers still consider the prevalence of bovine venereal campylobacteriosis to be underestimated as a consequence of the lack of sensitive and reliable techniques for diagnosis4.
Currently diagnosis of genital campylobacteriosis relies on the cultivation and identification of the causative organism1. The success hereof is dependent upon the collection and maintenance of the fragile C. fetus bacterium, which has limited viability outside the host8. Most of the cattle-breeding farms in South Africa tend to be extensive in nature and located long distances from diagnostic laboratories. In many cases the consulting veterinarian may need to travel long distances to sample animals and the task of ensuring that samples reach the laboratory timeously becomes a logistical dilemma. In order to address these problems, alternative diagnostic methods have been sought.
Polymerase chain reaction (PCR) assays present a number of advantages over more traditional methods and the technology is becoming increasingly more accessible to diagnostic laboratories. Apart from exhibiting high specificities and sensitivities, the ability of these types of assays to detect non-viable organisms could also potentially resolve problems encountered in the field with the preservation of organism viability for subsequent bacteriological culture22.
A number of C. fetus-species-and subspecies-specific PCR assays have been described. Of these the multiplex PCR assay initially evaluated by Hum and colleagues5 has been described extensively. This assay, which is designed to identify and differentiate the 2 C. fetus subspecies, has been evaluated by various researchers and been shown to be both sensitive and specific13,20. Good correlation of subspecies identification results, albeit not 100 %, was achieved when the PCR method was compared with traditional biochemical and genotyping methods, including random amplification polymorphic DNA (RAPD-PCR); pulsed field gel electrophoresis (PFGE) and amplified fragment length polymorphism (AFLP)10,13.
The aim of this investigation was to assess the suitability of the C. fetus-specific primer set, initially evaluated by Hum and colleagues5, to detect C. fetus directly in bovine preputial specimens. The multiplex assay was used in conjunction with traditional phenotyping tests to establish the suitability of the assay to assign South African C. fetus field isolates to subspecies.
MATERIALS AND METHODS
Bacterial and protozoan cultures
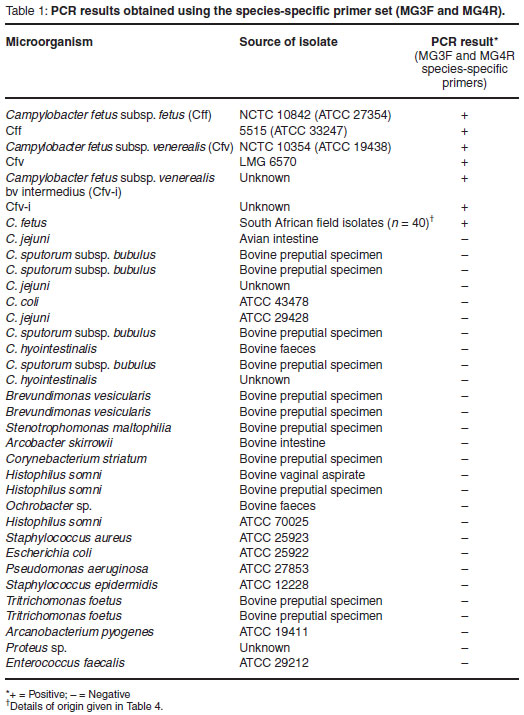
Forty-six Campylobacter fetus isolates and 27 bacterial and protozoan isolates, genomically similar to C. fetus or occupying a similar biological niche, were obtained from reference collections or were isolated from diagnostic material (Table 1). All bacterial isolates were grown on Columbia agar (Oxoid) supplemented with 5 % sheep blood, at 37 ºC under microaerophilic atmospheric conditions. Microaerophilic atmospheric conditions were generated in anaerobic jars using Campylobacter gas generating kits (Oxoid). The Tritrichomonas foetus isolates were maintained in Trichomonas medium (Oxoid) at 37 ºC.
PCR template preparation
DNA was isolated from diagnostic specimens and bacterial suspensions using a commercial kit (Qiagen® DNeasy blood and tissue kit). Extraction was carried out according to the manufacturer's instructions except when processing diagnostic specimens where 500 µℓ of sample was subjected to extraction and final elution of DNA was done using only 100 µℓ of the elution buffer.
PCR protocol
The C. fetus-specific uniplex PCR using the MG3F(5'-GGTAGCCGCAGCTGCTAA GAT-3') and MG4R (5'-TAGCTACAATAACGACAACT-3') primers was carried out as described by Schulze and colleagues16. For the processing of diagnostic specimens, the volume of DNA template added to the reaction mixture was 0.2 µℓ and the volume of Taq polymerase incorporated into the reaction mixture was increased to 0.6 µℓ (Roche 1 IU/µℓ ). The multiplex PCR was used to identify bacterial isolates to both species (MG3F and MG4R primer set) and subspecies level (VenSF (5-CTTAGCAGTTTGCGATATTGCCATT-3) and VenSR (5-GCTTTTGAGATAACAATAAGAGCTT-3) primer set). The test was performed as described previously with no modifications5. PCR products were analysed by electrophoresis using 1.5 % agarose gels in 1 X Tris-Borate-EDTA (TBE) buffer at 100 V. Following staining in ethidium bromide (0.5 µg/mℓ ), the DNA bands were visualised using a gel documentation system and sized against a 100-bp DNA ladder (Fermentas).
Evaluation of the specificity of the PCR
The specificity of the C. fetus-specific uniplex PCR was determined by testing C. fetus reference and South African field isolates (n=46) in addition to a collection of other Campylobacter species (n=10) and organisms which may be encountered in the genital tract of cattle (n=18). Furthermore, amplicons from one of the Cfv reference strains and a field isolate were sent for sequencing.
Evaluation of the sensitivity of the PCR and the influence of potential inhibitors
Dilutions of a Cfv reference strain (ATCC 19438), prepared in modified Weybridge transport medium7, were subjected to the PCR. Aliquots of each dilution were also plated out on blood agar and incubated at 37 °C in a microaerophilic atmosphere for 72 h to determine bacterial counts. The detection limit of the PCR assay was taken as the lowest bacterial inoculum (cfu/mℓ ) that yielded a positive PCR result. Once the detection limit was established, the 2 dilutions immediately above the limit were spiked with varying concentrations (1 %, 2 %, 5 %, 10 %, 20 % and 50 % (v/v)) of blood, urine, faeces and semen. Aliquots of each spiked sample were subjected to PCR.
Evaluation of the sensitivity of PCR and culture assays using spiked bovine preputial specimens
Preputial specimens were collected from bulls younger than 2 years of age and which had previously been tested and found to be negative for C. fetus. Between 4 and 8 animals were sampled and the preputial material inoculated into Weybridge medium. Samples were pooled in the laboratory before being divided into aliquots and inoculated with dilutions of the Cfv reference strain (ATCC 19438) prepared in phosphate buffered saline containing 0.02 % Tween 80. Inoculated aliquots were kept at room temperature for 72 h. During this time, material was removed from each aliquot at 0, 24, 48 and 72 h post-inoculation for PCR and culture. For bacteriological culture, 100 µℓ of each aliquot was spread plated onto Skirrow's agar (Oxoid) and approximately 300 µℓ filtered (0.65 µm cellulose acetate filter) onto blood agar supplemented with FBP Campylobacter enrichment supplement (Oxoid). Following incubation at 37 ºC under microaerophilic conditions, all plates were examined for colonies typical of C. fetus, Gram-stained and counted11.
Field evaluation
All bovine preputial scrapes (n = 212) submitted in Weybridge transport medium to Allerton Provincial Veterinary Laboratory, KwaZulu-Natal, South Africa between June 2007 and March 2008 were tested using the uniplex PCR assay as well as the traditional bacteriological culture method. It was decided to only test preputial scrapes in modified Weybridge transport medium as this has been proven to give the highest sensitivity when culturing is used8,17. All samples were processed within 36 hours of collection. Following incubation, all plates were examined for the presence of small, smooth, shiny colonies having a slightly grey to pink appearance. Suspect colonies were Gram-stained and identified as C. fetus if they were oxidase positive, grew at 25 ºC, were resistant to nalidixic acid and sensitive to cephalothin and did not produce hydrogen sulphide in triple sugar iron agar11.
Identification of subspecies
Campylobacter fetus reference and field isolates were identified to subspecies level using the multiplex PCR and 2 phenotyping tests namely: tolerance to 1 % glycine (BDH) and hydrogen sulphide production using lead acetate paper11. Additionally the ability to reduce selenite and susceptibility to metronidazole and cefoperazone was determined11,16.
RESULTS
PCR specificity and sensitivity
The MG3F and MG4R C. fetus-specific primers were shown to be highly specific yielding a single, distinct PCR amplicon of approximately 750-bp when tested against all C. fetus isolates. No amplicons were observed when any of the other Campylobacter strains or the collection of genital-associated microbes which were tested. A summary of the results is illustrated in Table 1.
The specificity of the primer pair was further demonstrated by DNA sequencing and alignment of the PCR amplification products from 1 of the Cfv reference strains (LMG 6570) and a field isolate (Inqaba Biotec). A BLAST search, using the assembled sequencing data, revealed 100 % similarity between the 2 sequences obtained and C. fetus sequences loaded in GenBank (Accession Numbers: CP000487.1; AY158814.1 and AY 158813.1).
The determination of sensitivity of the PCR revealed a detection limit of 615 cells/mℓ Weybridge medium, or 6.15 cells per PCR reaction. The 2 Cfv dilutions above the detection limit were spiked with varying concentrations of faeces, blood, urine and semen. Significant inhibition of the PCR reaction was noted for samples spiked with faeces; a 10-fold decrease in sensitivity was observed with aliquots spiked with 1 % faeces and complete inhibition observed with samples spiked with 2 % faecal material. Conversely, blood, urine and semen seemingly had no impact on the PCR results at the concentrations tested even when the test aliquots were spiked with as much as 50 % of the potential inhibitor.
Evaluation of the sensitivity of PCR and culture assays using spiked bovine preputial specimens
The evaluation of sensitivity of the PCR on spiked bovine specimens were carried out in triplicate using preputial material collected from different herds. The detection limit of both the culture and PCR assays were monitored over a period of 72 hours and recorded as the lowest Cfv inoculum (Cfv/mℓ) that yielded a positive result. Results are summarised in Table 2.
The 1st trial run was hampered by the presence of Pseudomonas aeruginosa in the samples. All Skirrow's plates were overgrown with Ps. aeruginosa preventing the detection and recovery of any C. fetus colonies.
Field evaluation
During the investigation period 212 bull preputial scrapes were received and processed using PCR and bacteriological culture. The results obtained using both methods are presented in Table 3.
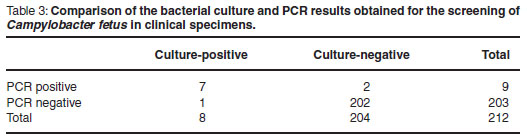
From this data the analytical sensitivity and specificity of the PCR assay was calculated using guidelines12. A sensitivity of 85.7 % and a specificity of 99 % were obtained.
Identification of subspecies
The multiplex PCR was evaluated using 6 reference C. fetus isolates representing both sub species as well as C. fetus subsp. venerealis biovar intermedius (Cfv-i). A single PCR amplicon of approximately 750-bp was obtained with all 6 cultures. A 2nd amplicon of approximately 180-bp was obtained with both of the Cfv isolates but not the Cff or the Cfv-i reference isolates.
Phenotyping and PCR subspecies identification results for the 6 C. fetus reference cultures and 40 field isolates are summarised in Table 4. Based on the pivotal glycine tolerance test, 5 of the 40 field isolates were identified as Cff. All of the remaining isolates were unable to grow on blood agar supplemented with 1 % glycine; by virtue of the fact that all of these isolates produced hydrogen sulphide in L-cysteine supplemented medium, they were classified as Cfv-i.
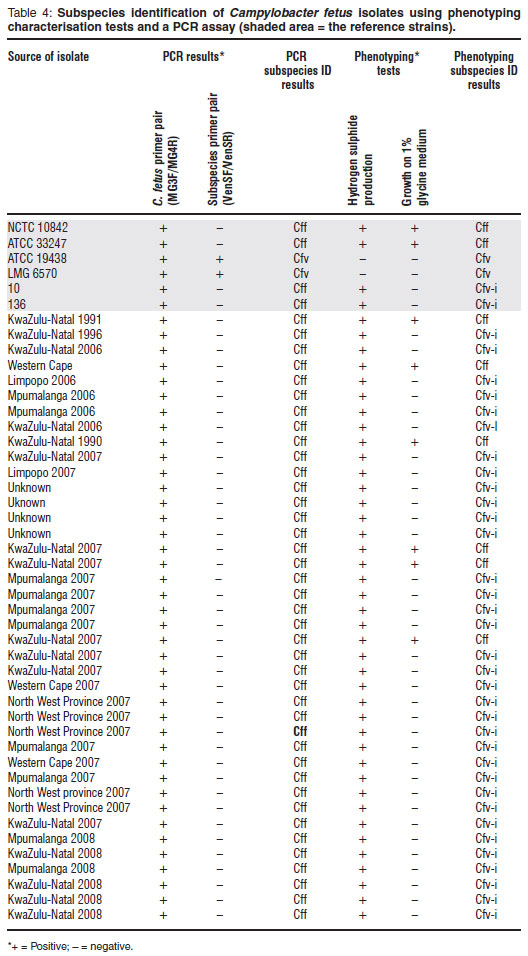
Using the multiplex PCR assay all isolates yielded the same results: species-specific amplicons were observed but no subspecies-specific amplicons. The VenSF and VenSR subspecies-specific primer set used in the assay targets a segment of the Cfv genome. The generation of an amplicon of approximately 182-bp is indicative that the test organism is Cfv. Campylobacter fetus subsp. fetus is identified by exclusion. Based on the PCR test results obtained, all the field isolates should be classified as Cff.
DISCUSSION
In veterinary diagnostic laboratories the culture and isolation of C. fetus is the traditional method for the diagnosis of bovine campylobacteriosis. The inherent limitations of this methodology prompted this investigation to identify and evaluate a more sensitive means of detecting C. fetus. Molecular techniques, particularly PCR, are becoming more practical and affordable for diagnostic laboratories to incorporate into their test portfolio. The specificity and sensitivity of assays, combined with the speed at which samples can be processed and results obtained, are appealing attributes22.
The multiplex PCR initially evaluated by Hum and colleagues5 is the most extensively described assay for the identification of C. fetus at species and subspecies level. In the multiplex format many investigators have used the PCR for the direct identification of subspecies of bacterial isolates. No publications to date have reported on the use of this PCR for the direct screening of clinical specimens. The ability to screen clinical specimens directly could potentially improve the overall sensitivity and accuracy of diagnostic results as many of the shortcomings conventionally encountered with the culture method would be eliminated. As a possible screening assay only the species-specific primer set was evaluated. It was reasoned that as a result of the reportedly low prevalence of C. fetus in South Africa6 and the improved test sensitivity achieved with uniplex test formats16 it would be more feasible to screen samples using only the C. fetus-specific primer set.
The specificity of the species-specific uniplex PCR was demonstrated by testing a collection of reference and field C. fetus isolates as well as bacteria taxonomically related to C. fetus or known to exist in similar microbiological niches. All C. fetus isolates yielded a single PCR amplicon of approximately 750-bp in size when tested with the uniplex PCR (Table 1). None of the other isolates tested produced amplicons, indicating that the test has an analytical specificity of 100 %. It is interesting to note that initial publications5,13,20 reported the generation of a PCR amplicon of about 960-bp in size. The amplicon has, however, been more recently reported and sequenced as 750-bp9,12,23. The reason for the discrepancy of almost 200-bp in size has not been explained.
During sample collection preputial material can quite easily become contaminated with urine, semen, faeces and/or blood. All 4 of these components have been documented as containing elements which may interfere with PCR tests24.It was consequently deemed necessary to evaluate the influence of each of the components on the sensitivity of the PCR assay. The detection limit of the assay was seemingly unaffected when samples spiked with up to 50 % of urine, semen or blood were tested. Faecal contamination, however, was shown to have a major inhibitory effect on the assay even at concentration levels as low as 1 %. The use of an alternative DNA extraction protocol could be investigated as a means of overcoming this potential problem but the inclusion of an internal control in the PCR assay would probably be the most feasible way of ensuring that false negative results are not reported as a result of PCR failure due to the presence of inhibitors in the samples.
The spiking of preputial material was carried out in triplicate using material collected by different practitioners and from animals in different herds. Sampling criteria were instituted to ensure as far as possible that only animals negative for C. fetus were sampled. The 1st run of spiked preputial material clearly illustrated the advantage of PCR over bacterial culture as a means of detecting C. fetus. The presence of Pseudomonas in the preputial material dramatically impeded the recovery of Cfv. Pseudomonas colonies, resistant to the antibiotics in the Weybridge transport and Skirrow's media, readily grew on the culture plates, obscuring any Cfv colonies which were plated out. Using the PCR assay however, Cfv was detected at 77 Cfv/mℓ (Table 2) at both 0 h and 24 h. Surprisingly, no amplicons were obtained when the 72 h spiked aliquots was tested. In view of the fact that PCR amplicons were visualised when the 72 h spiked aliquots from both subsequent test runs were analysed, it is hypothesised that the failure to detect Cfv is a result of the presence of Pseudomonas and possibly other contaminants that had increased in number in the pooled sample.
Certain strains of Pseudomonas such as Pseudomonas aeruginosa are known to produce the enzyme DNase3. The activity of this enzyme could have resulted in the degradation of DNA present in the sample. Problems resulting from the contamination of preputial cultures by Pseudomonas spp. have been reported8. Itis interesting to note, however, that none of the diagnostic samples tested during field evaluation exhibited such extensive Pseudomonas contamination as was noted with this test run, suggesting that in practice this type of contamination may be a rare occurrence.
Owing to geographical and other logistical reasons, private practitioners in South Africa typically face difficulties in getting specimens to the laboratory within the 6-hour viability period of the bacterium. The use of a transport enrichment medium (TEM) is a prerequisite as it permits a more lenient window period between sample collection and laboratory testing to be observed. As shown by the test results obtained here, the detection limit of the culture assay decreased with time, even when use was made of a TEM. Nonetheless, Cfv was still recoverable from samples up to 72 hours post-inoculation. Comparatively, the sensitivity of the PCR did not appear to be affected by time, with the detection limit remaining unchanged between 0 and 72 hours. The greater sensitivity achieved with PCR at 48 and 72 hours post-inoculation of samples (Table 2) is of particular interest as these are the time intervals more likely to be experienced in the field.
Field evaluation of the PCR assay was carried out over a 10-month period using diagnostic specimens submitted to Allerton Provincial Veterinary Laboratory. All bovine sheath scrapes submitted in Weybridge transport medium were tested using both the culture method and the PCR assay. Of the 212 samples received and tested, 4.2 % were found to be positive using the PCR assay while 3.8 % were found to be positive using the culture method. A high degree of agreement was observed between the PCR and the culture results (Table 3). Discrepancies between results were only noted on 2 occasions. In the 1st instance 1 of the samples was found to be positive by PCR but negative on culture. Both tests were repeated with the same results being obtained. This discrepancy is possibly a demonstration of the higher sensitivity of the PCR assay; illustrating the advantages of the PCR in detecting C. fetus in low numbers where the culture could possibly fail, especially in the presence of large numbers of contaminant bacteria or where non-viable bacteria are present. The converse situation was observed with the 2nd anomalous result where a negative PCR result was obtained but the sample was positive on culture. This result is harder to explain. The presence of trace amounts of PCR inhibitors cannot be ruled out. Neither can the possibility that the small volume (500µℓ) of the aliquot used for for DNA extraction and subsequent dilution of possible DNA units in the PCR reaction mix could have resulted in the diluting out of the target DNA. In spite of the 2 anomalous results obtained, it is important to note that neither of the tests under evaluation failed to detect a positive herd when repeat testing was carried out.
The high degree of agreement between the PCR and culture results was surprising, particularly since the shortcomings associated with the culture method are well documented8. The success achieved with culturing is attributed in part to the fact that the laboratory imposed rigorous sampling guidelines; only sheath scrape samples submitted in Weybridge transport medium were accepted. Sheath scrapes are preferable to sheath washes, a technique still used by many local practitioners, since the scraping technique has been shown to recover larger numbers of Cfv and provide 'cleaner' samples with less bacterial contamination17. In addition to sampling criteria, the culture procedure in the laboratory was enhanced by ensuring that all samples were plated out onto 2 different media; 1 selective and 1 FBP-enriched medium. This practice is often compromised, particularly in some of the smaller regional veterinary laboratories where budget constraints are prevalent.
In order to evaluate the described subspecies-specific primer set, VenSF and VenSR, a collection of South African C. fetus field isolates were identified to subspecies using the multiplex PCR as well as traditional phenotyping techniques. Currently, tolerance to 1 % glycine is the only internationally accepted phenotypic test prescribed for the differentiation of the 2 C. fetus subspecies11. The reproducibility of this assay is, however, poor and the test can give ambiguous results19. Other biochemical tests have been described for the purpose of determining subspecies but contradictory results have been obtained both in this study (results not shown) as well as by Schulze and colleagues16.
The C. fetus isolates tested represented a geographically diverse collection which was sourced from different veterinary laboratories across the country. Furthermore, some of the field isolates were recovered from culture collections and represent a genetic gene pool that existed up to 2 decades ago. With the exception of 5 of the field isolates, which were identified as Cff, all the isolates were identified as Cfv-i using phenotyping methods. The PCR results, however, indicated that all field isolates were Cff. The lack of correspondence obtained was somewhat startling. Previous reports have only intimated minor incongruencies when comparing these 2 approaches to identifying subspecies5,9 although a recent investigation23 alluded to greater problems when a collection of United Kingdom isolates were tested. These investigators found that only 3 isolates, out of a group of 19, which were initially phenotyped as Cfv, gave the same results for subspecies identification using this specific multiplex PCR assay. After further analysis the researchers proposed that the discrepancies were the result of an unusual Cfv clone circulating within the cattle population in the United Kingdom (UK) and that the multiplex PCR assay was consequently unsuitable for the subspecific identification of UK isolates.
Unfortunately the initial evaluation of the multiplex PCR did not take Cfv-i strains into consideration5. In fact, many of the phenotyping and genotyping investigations which have been carried out in this field have seemingly ignored the existence of this particular group of bacteria. This is to some extent understandable since the initial description of this group of isolates was very vague and reference isolates were not readily available18.Recently, however, with the availability of high resolution genotyping tools such as AFLP and multilocus sequence typing (MLST), isolates that have been phenotyped as Cfv biovar intermedius, have been more specifically characterised. These techniques have supported phenotyping results and have in fact provided evidence for the reclassification of Cfv-i as a separate subspecies19.
The apparent absence of classical Cfv field strains and the widespread distribution of Cfv-i in South Africa render the multiplex PCR unsuitable for local use. In order to resolve the subspecies identification dilemma, efforts need to be redirected towards the sequence analysis of the Cfv-i genome. The identification of unique gene sequences, or lack of specific sequences, may allow for the development of a primer set which can be used to differentiate these strains from classical Cfv and Cff isolates. Until such time, the reliance on bacteriological culture for isolation, identification and subspecies differentiation of C. fetus will persist.
Despite the practical drawbacks typically associated with bacteriological culture the relatively high success rate achieved during this study has been particularly encouraging. In order to ensure optimal recovery it is advocated that preputial scrapes are collected and submitted for laboratory examination in an appropriate transport medium such as Weybridge. Samples should be kept cool and reach the laboratory within 36 hours of collection. Culturing onto at least 2 different culture media (one selective and 1 enriched) is advised. As far as identification to subspecies of isolates is concerned, diagnosticians are advised to ensure that the pivotal glycine tolerance test is carried out in triplicate for all isolates being tested in order to ensure the accuracy of the test results.
ACKNOWLEDGEMENTS
We wish to sincerely express our gratitude to all the colleagues who submitted samples and made this study possible.
REFERENCES
1. Brooks B W, Devenish J, Lutze-Wallace C L, Milnes D, Robertson, R H, Berlie-Surujballi G 2004 Evaluation of a monoclonal antibody-based enzyme-linked immunosorbent assay for detection of Campylobacter fetus in bovine preputial washing and vaginal mucus samples. Veterinary Microbiology 103: 77-84 [ Links ]
2. Garcia M M, Eaglesome M D, Rigby C 1983 Campylobacters important in veterinary medicine. Veterinary Bulletin 53: 793-811 [ Links ]
3. Gilardi G L 1985 Pseudomonas. In Lenette E H, Balows A, Hausler, W J, Shadomy, H J (eds) Manual of clinical microbiology (4th edn). American Society of Microbiology, Washington: 350-372 [ Links ]
4. Hum S, Quinn K, Kennedy D 1994 Diagnosis of bovine venereal campylobacteriosis by ELISA. Australian Veterinary Journal 71: 140-143 [ Links ]
5. Hum S, Quinn K, Brunner J, OnSLW 1997 Evaluation of a PCR assay for identification and differentiation of Campylobacter fetus subspecies. Australian Veterinary Journal 75: 827-831 [ Links ]
6. Irons P C, Schutte A P, van der Walt M L, Bishop G C 2004 Genital campylobacteriosis in cattle. In Coetzer J A W, Tustin R C (eds) Infectious diseases of livestock with special reference to southern Africa Vol. 3 (2nd edn), Oxford University Press, Cape Town: 1459- 1468 [ Links ]
7. Lander K P 1990 The development of a transport and enrichment medium for Campylobacter fetus. British Veterinary Journal 146: 327-333 [ Links ]
8. Monke H J, Love B C, Wittum T E, Monke D R, Byrum B A 2002 Effect of transport enrichment medium, transport time, and growth medium on the detection of Campylobacter fetus subsp. venerealis. Journal of Veterinary Diagnostic Investigation 14: 35-39 [ Links ]
9. Muller W, Hotzel H, Schulze F 2003 [Identification and differentiation of Campylobacter fetus subspecies by PCR] Deutsche Tierärzliche Wochenschrift 110: 55-59 (In German) [ Links ]
10. Newell D G, Duim B, van Bergen M A P, Grogono-Thomas R, Wagenaar J A 2000 Speciation, subspeciation and subtyping of Campylobacter spp. associated with bovine infertility and abortion. Cattle Practice 8: 421-425 [ Links ]
11. OIE 2008 Bovine genital campylobacteriosis. In Manual of diagnostic tests and vaccines for terrestrial animals (mammals, birds and bees) (6th edn). Office International desÉpizooties, Paris: 661-670 [ Links ]
12. OIE 2008 Validation and quality control of polymerase chain reaction methods used for the diagnosis of infectious diseases. In: Manual of diagnostic tests and vaccines for terrestrial animals (mammals, birds and bees)(6th edn). Office International des Épizooties, Paris: 46-55 [ Links ]
13. On SL W, Harrington C S 2001 Evaluation of numerical analysis of PFGE-DNA profiles for differentiating Campylobacter fetus subspecies by comparison with phenotypic, PCR and 16S rDNA sequencing methods. Journal of Applied Microbiology 90: 285-293 [ Links ]
14. Pefanis S M, Herr S, Venter C G, Kruger L P, Queiroga C C, Amaral L 1988 Trichomoniasis and campylobacteriosis in bulls in the Republic of Transkei. Journal of the South African Veterinary Association 59: 139-140 [ Links ]
15. Schmidt-Dumont A M A 1984 The incidence of Campylobacter fetus and Tritrichomonas foetus in cattle in South West Africa/Namibia. Proceedings of the Tenth International Congress on Animal Reproduction and Artificial Insemination, Urbana Champaign, Illinois, June 1984: 10-14 [ Links ]
16. Schulze F, Bagon A, Muller W, Hotzel H 2006 Identification of Campylobacter fetus subspecies by phenotypic differentiation and PCR. Journal of Clinical Microbiology 44: 2019-2024 [ Links ]
17. Tedesco L F, Errico F, Del Baglivi L P 1977 Comparison of three sampling methods for the diagnosis of genital vibriosis in the bull. Australian Veterinary Journal 53: 470-472 [ Links ]
18. Van Bergen M A P, Linnane S, van Putten J P, Wagenaar J A 2005 Global detection and identification of Campylobacter fetus subsp. venerealis. Revue Scientifique et Technique,Office International des Épizooties 24: 1017-1026 [ Links ]
19. Van Bergen M A P, Simons G, van der Graaf-van Bloois L, van Putten J P M, Rombout J, Wesley I, Wagenaar J A 2005 Amplified fragment length polymorphism based identification of genetic markers and novel PCR assay for differentiation of Campylobacter fetus subspecies. Journal of Medical Microbiology 54: 1217-1234 [ Links ]
20. Vargas A C, Costa M M, Vainstein M H, Kreutz L C, Neves J P 2003 Phenotypic and molecular characterization of bovine Campylobacter fetus strains isolated in Brazil. Veterinary Microbiology 93: 121-132 [ Links ]
21. Wagenaar J A, van Bergen M A P, Newell D G, Grogono-Thomas R, Duim B 2001 Comparative study using amplified fragment length polymorphism fingerprinting, PCR genotyping, and phenotyping to differentiate Campylobacter fetus strains isolated from animals. Journal of Clinical Microbiology 39: 2283-2286 [ Links ]
22. Willoughby K 2003 The ABC of PCR. In Practice 25: 140-145 [ Links ]
23. Willoughby K, Nettleton P F, Quire M, Maley M A, Foster G, Toszeghy M, Newell D G 2005 A multiplex polymerase chain reaction to detect and differentiate Campylobacter fetus subspecies fetus and Campylobacter fetus - species venerealis: use on UK isolates of C. fetus and other Campylobacter spp. Journal of Applied Microbiology 99: 758-766 [ Links ]
24. Wilson I G 1997 Inhibition and facilitation of nucleic acid amplification. Applied and Environmental Microbiology 63: 3741-3751 [ Links ]
Received: February 2010.
Accepted: April 2010.
* Author for correspondence. E-mail: tracy.schmidt@kzndae.gov.za














