Serviços Personalizados
Artigo
Indicadores
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares em Google
Similares em Google
Compartilhar
South African Journal of Animal Science
versão On-line ISSN 2221-4062
versão impressa ISSN 0375-1589
S. Afr. j. anim. sci. vol.46 no.4 Pretoria 2016
http://dx.doi.org/10.4314/sajas.v46i4.4
SHORT COMMUNICATION
Identification of genetic variation in the major histocompatibility complex gene region in Turkish sheep breeds
F. Ilhan#; I. Keskin; A. Tozluca
Department of Animal Science, Faculty of Agriculture, Selcuk University, 42075, Konya, Turkey
ABSTRACT
The major histocompatibility complex (MHC) in sheep, Ovar-Mhc, remains poorly characterized relative to other domestic animals. However, its basic structure is similar to that of other mammals, comprising class I, II and III regions. In this study, the Ovine MHC class II DRB1 and DRB3 genes were amplified by polymerase chain reaction in eight sheep breeds reared in Turkey. Informative restriction fragment length polymorphisms (RFLPs) were obtained with five restriction enzymes for DRB1 and with two restriction enzymes for DRB3. The digestion of DRB1 exon 2 with NciI, SacI, SacII, Hin1I each produced three genotypes and two alleles (viz., a and b) with frequencies of 0.69 and 0.31; 0.65 and 0.35; 0.91 and 0.09; 0.57 and 0.43, respectively. The digestion of DRB1 exon 2 with DcCeI produced four genotypes and three alleles (viz., a, b and c) with frequencies of 0.62, 0.28 and 0.10, respectively. On the other hand, the digestion of DRB3 exon 2 with NcteII and BsaI each produced three genotypes and two alleles (viz., a and b) with frequencies of 0.72 and 0.28; 0.96 and 0.04, respectively. This study presents the genetic profiles of the exon 2 region of the MHC DRB1 and DRB3 genes in native Turkish sheep breeds.
Keywords: DRB1, DRB3, polymerase chain reaction (PCR), restriction fragment length polymorphism(RFLP)
The major histocompatibility complex is a large genomic region or gene family that is found in most vertebrates, which encodes MHC molecules. MHC molecules play an important role in the immune system and auto-immunity (Tizard, 2004). The MHC was discovered during tissue transplantation studies in mice and was first characterized for its role in histocompatibility. Subsequently, its roles in immune regulation and several other functions were discovered. The primary function of the MHC is to code for specialized antigen-presenting receptor glycoproteins, known as histocompatibility molecules or MHC molecules (Dukkipati et al., 2006). The major histocompatibility complex in sheep is poorly characterized when compared with other domestic animals. However, its basic structure is similar to that of mammalian MHC molecules, comprising class I, II and III regions. The MHC genes of sheep are known as ovar and are located on chromosome 20. Ovar class II genes encode polymorphic glycoproteins composed of heterodimers of noncovalently linked alpha (α) and beta (β) subunits, which play a pivotal role in the initiation of the immune response to pathogen-derived peptide antigens (Brujeni et al., 2009). The schematic structure of the ovine MHC is illustrated in Figure 1.
Class I MHC glycoproteins are expressed on the surface of all nucleated somatic cells. They are found in highest concentration on lymphocytes and macrophages, and consist of heterodimers of and/or heavy chain noncovalently linked to a light β2-microglobulin chain. Class I genes include eight exons and seven introns. The DRB locus is highly polymorphic among the class II MHC genes (Anderson & Rask, 1988). This locus encodes heterodimeric peptide-binding proteins and proteins that modulate antigen loading onto the MHC class II. Relative to other parts of the MHC, the class III region has the highest gene density, with the least number of pseudogenes (Kulski et al., 2002). However, some of the genes located in this region are not involved in immune system functions. The MHC class III region encodes other immune components including the complement system (e.g., C2, C4 or factor B) and cytokines (e.g., TNF-α).
Studies of genetic variation in Ovar-Mhc class II genes have shown that the expressed DRB locus is highly polymorphic (Schwaiger & Epplen, 1995; Schwaiger et al., 1996; Jugo & Vicario, 2000; Konnai et al., 2003; Ballingall et al., 2008; Nikbakht et al., 2012; Lotfi et al., 2012; Shen et al., 2014; Takeshima et al., 2014). In particular, a high polymorphism level is present in exon 2, which encodes the antigen-binding site (Escayg et al., 1997; Konnai et al., 2003). There is little knowledge of MHC polymorphism in Turkish sheep. Only one study has been performed by Bozkaya & Kurar (2005) until today. Therefore, in this study, the ovine MHC, class II DRB1 and DRB3 genes were analysed by polymerase chain reaction and restriction fragment length polymorphism (PCR-RFLP) in eight Turkish sheep breeds.
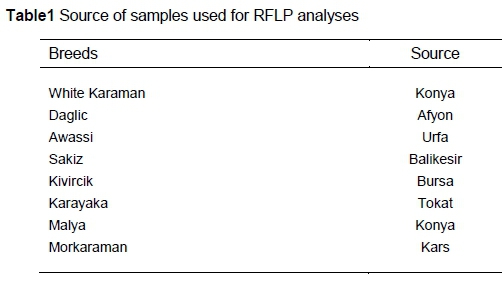
Genetic variation was analysed in sheep from eight breeds reared in Turkey, including the White Karaman, Daglic, Awassi, Sakiz, Kivircik, Karayaka, Malya and Morkaraman (15 to 25 sheep per breed) (Table 1).

Blood samples were obtained from Ankara University Faculty of Agriculture. Genomic DNA was isolated from blood samples according to the method describe in Miller et al. (1988).
In this study, genetic variability in the ovine MHC, class II DRB1 and DRB3 genes were analysed using the PCR-RFLP technique.
OLA-DRB1 was amplified according to Konnai et al. (2003). Nested PCR was used to amplify the second exon of the DRB1 gene. The first round of PCR was performed with primers OLA-ERB1 (5'-ccggaattcccgtctctgcagcacatttctt-3') and HL031 (5'-tttaaattcgcgctcacctcgccgct-3'). The PCR reactions were carried out in a total volume of 25 μΙ containing the following reaction mixture: 2 μΙ of 10* reaction buffer with KCl, 1.5 mM MgCl2, 1.2 mM of dNTP mix, 20 μΜ of each primer, 2.5 U of Taq polymerase and 20 ng of purified sheep DNA. The following amplification profile was used: initial denaturation at 94 °C for 5 min, 15 cycles at 94 °C for 30 s, annealing at 50 °C for 30 s and extension at 72 °C for 1 min, followed by a final extension step at 72 °C for 10 min. The second round of PCR amplification was carried out using 5 μ! of the resultant mixture, with the addition of primers OLA-ERB1 and OLA-XRB1 (5'-gctcgagcgctgcacagtgaaactc-3'). The thermal cycle included an initial denaturation at 94 °C for 5 min, 30 cycles at 94 °C for 30 s, annealing at 60 °C for 30 s and extension at 72 °C for 1 min, followed by a final extension step at 72 °C for 10 min. The amplified PCR products were electrophoresed on 1% agarose gels to verify the fragment sizes. The amplified second exon of the DRB1 gene was digested with five restriction enzymes to verify the expected recognition sites. The informative restriction enzymes used for the DRB1 PCR fragments were NciI, SacI, SacII HinII and DdeI.
OLA-DRB3 was amplified according to Amills et al. (1996). PCR reactions were performed using primers DRB1.1 (5'-tatcccgtctctgcagcacatttc-3') and DRB1.2 (5'-tcgccgctgcacactgaaactctc-3'). Amplification was performed in a 25-μl reaction volume containing 2.5 μl of 10* reaction buffer with KCl, 1.5 mM MgCl2, 1.0 mM of dNTP mix, 10 μΜ of each primer, 1.5 U of Taq polymerase and 20 ng of purified sheep DNA. The thermal cycle was programmed for an initial denaturation at 94 °C for 5 min, followed by 30 cycles of denaturation at 94 °C for 1 min, annealing at 60 °C for 30 s and extension at 72 °C for 1 min, followed by a final extension step at 72 °C for 10 min. The amplified DRB3 PCR products were digested with two restriction enzymes NdeII and BsaI.
The informative restriction enzymes were analysed using 15 to 25 individuals from each population. The digested fragments were separated electrophoretically on 2 or 3% agarose gels in 1x TBE buffer, stained with ethidium bromide and photographed using a Vilber Lourmat gel imaging system.
Gene and genotypic frequencies were estimated by direct counting. Deviations from Hardy-Weinberg (HW) equilibrium were estimated by chi-square (χ2) (Duzgunes et al., 1983).
In the present study, polymorphism in exon 2 of the Ovar-DRBI and Ovar-DRB3 loci was analysed by PCR-RFLP of DNA samples obtained from eight Turkish sheep breeds. The average size of the PCR-amplified Ovar-DRB1 region observed by agarose gel electrophoresis for all the populations studied was 296 bp. Five restriction enzymes generated informative digestion profiles in DRB1 exon 2. The NciI, SacI, SacII, Hin11 and Ddel restriction enzymes generated different digestion profiles of DNA samples from eight Turkish sheep breeds (Table 1). The average size of the PCR-amplified Ovar-DRB3 region for all the populations studied was 285 bp. The NdeII and BsaI restriction enzymes generated informative digestion profiles in Ovar-DRB3 exon 2 (Table 2).
The enzymatic digestion of exon 2 of DRB1 with NciI, SacI, SacII, Hin1I each generated three genotypes and two alleles, and DdeI digestion generated four genotypes (aa, bb, ab and ac) and three alleles. The enzymatic digestion of exon 2 of DRB3 with NdeII and BsaI each generated three genotypes and two alleles.
Allele a was obtained from the NciI, SacI, SacII and Dde\ digestions of DRB1 exon 2 and was found to be more frequent in all of these sheep breeds. On the other hand, allele b obtained from the digestion of DRB1 with HinII was more frequent in the Kivircik and White Karaman breeds (Table 3). The DRB1 exon 2 showed high heterozygosity by digestion with HinII. The lowest heterozygosity was obtained by digestion of DRB1 exon 2 with SacII and NciI. The HW test showed that the studied most of the breeds fit the theoretical proportions for the NciI, SacI, SacII and DdeI digestions of exon 2 of DRB1 (Table 4) (P <0.05). However, White Karaman, Sakiz, Karayaka and Malya don't fit the theoretical proportions for the Hin1I digestions of this gene (Table 4) (P <0.05).
The digestion of DRB3 exon 2 with NdeII and BsaI each resulted in three genotypes and two alleles. The NdeII- and BsaI-generated allele a of exon 2 of DRB3 was more frequent in all of the studied sheep breeds (Table 5). The DRB3 gene had a high heterozygosity for NdeII digestion of exon 2. The HW test showed that the studied most of the breeds fit the theoretical proportions for the NdeII and BsaI digestions of the exon 2 region of DRB3 (Table 4) (P <0.05). Table 6 shows the allele frequencies of various patterns in the exon 2 of DRB1 and DRB3 after RFLP analysis.
This study presents the first insights and RFLP profiles of the DRB1and DRB3 genes in native sheep breeds in Turkey. Alleles found in this study were similar to those previously identified and reported by Konnai et al. (2003) and Amills et al. (1996). Although novel alleles were not identified, further detailed DNA sequence analysis of the exon 2 regions of Ovar-DRB1 and Ovar-DRB3 from Turkish sheep breeds will be carried out.
Several methods have been employed for typing Ovar-DRB1 genes in various sheep breeds and have revealed extensive polymorphism at these loci. Among these methods, PCR-RFLP analysis has been suggested for the effective typing of DRB1 alleles of farm animals (Amills et al. 1996; Konnai et al. 2003; Dongxiao & Yuan 2004; Gruszczynska et al. 2005; Brujeni et al. 2009). To date, 106 Ovar-DRB1 alleles have been identified by DNA sequencing of exon 2 from various breeds of sheep (Schwaiger et al., 1995; Schwaiger et al., 1996; Jugo & Vicario, 2000; Konnai et al., 2003; Ballingall et al., 2008). The results of the studies have shown that MHC polymorphism in animals correlates with immunity and the immune response.
The NdeII b allele has been correlated with the occurrence of Ile 37, 67 substitutions. The positions are expected to be involved in the formation of one region of the antigen recognition site (ARS). The resolving power of this method allows the detection of amino acid substitutions at the ARS of the DR molecule (a MHC class II cell surface receptor) may help to understand the genetic basis of disease resistance (Amills et al. 1996).
There is little knowledge of MHC polymorphism in ruminants compared with that of human beings and mice. Therefore, similar studies should be extended to ruminants, including the analysis of additional MHC regions. Allelic variation in the MHC region and its association with various traits in immunity has yet to be analysed in ruminants. The current study is a preliminary analysis of the polymorphism of the MHC DRB1 and MHC DRB3 genes in Turkish sheep breeds, which provides a basis for further studies. Knowledge of MHC polymorphism in Turkish sheep breeds allows the development of sheep by helping to understand the genetic basis of their resistance to disease. A larger sample size and additional molecular analysis are required to verify these results and determine the association between the MHC DRB genes and disease resistance in ruminants.
Acknowledgments
This study was performed by Fatma Ilhan in partial fulfilment of the Ph.D. degree in Biometry and Genetics, Animal Science Dept., Selcuk University. This project was supported by a grant from The Scientific Research Project Coordinating Office of Selcuk University. (Project No: 10101036).
Authors' Contributions
Conception, design, data collection, analyses and drafting of paper - FI; Critical revision - IK; Data collection, final approval of version to be published - AT.
Conflict of Interest Declaration
There are no conflicts of interest.
References
Amills,M., Francino, O. & Sanchez, A., 1996. A PCR-RFLP typing method for the caprine MHC class II DRB gene. Vet. Immunol and Immunopathol, 55, 255-260. [ Links ]
Andersson, L. & Rask. L., 1988. Characterization of the MHC class II region in cattle. The number of DQ genes varies between haplotypes, Immunogenet 27, 110-120. [ Links ]
Ballingall, K.T., Fardoe, K. & McKeever, D.J., 2008. Genomic organisation and allelic diversity within coding and non- coding regions of the Ovar-DRB1 locus. Immunogenet 60(2),95-103. [ Links ]
Bozkaya, F. & Kurar, E., 2005. Linkage disequilibrium between MHC-linked microsatellite loci in White Karaman, Awassi and Merinolandschaf sheep breeds. Firat Un. Med. J. of Health Sci. 19(1 ),57-61. [ Links ]
Brujeni, N.G.H., Emam, M., Mahmoudzadeh, H., Hamedmonfared, E., Talebnia Jahromi, R. & Rezaei, H., 2009. Typing of Ovar-DRB1 second exon with PCR-RFLP technique in Iranian Shaul Sheep. Iranian J. of Vet Research, Shiraz Un. Vol. 10, No. 3, Ser. No. 28. [ Links ]
Dongxiao, S. & Yuan, Z., 2004. Polymorphisms of the second exon of MHC-DRB gene in Chinese local sheep and goat. Biochemical Genetics. 42(9-10),385-390. [ Links ]
Dukkipati, V.S.R., Blair, H.T., Garrick, D.J. & Murray, A., 2006. Ovar-MHC- ovine major histocompability complex: structure & gene polymorphisms. Genetics and Molecular Research 5 (4),581-608.3 [ Links ]
Duzgunes, O., Kesici, T. & Gurbuz, F., 1983. Methods of statistic. Ankara Un. Fac. of Agr. Vol. 229. [ Links ]
Escayg, A.P., Hickford, J.G.H. & Bullock, D.W., 1997. Association between alleles of the ovine major histocompatibility complex and resistance to footrot. Res. in Vet. Sci. 63, 283-7. [ Links ]
Gruszczynska, J., Brokowska, K., Charon, K.M. & Swiderek, W.P., 2005. Restriction fragment length polymorphism of exon 2 Ovar-DRB1 gene in Polish Heath Sheep and Polish Lowland Sheep. J Appl Genet 46(3), 311-314. [ Links ]
Jugo, B.M. & Vicario, A., 2000. Single-strand conformational polymorphism and sequence polymorphism of MHC-DRB in Latxa and Karrantzar sheep: implications for caprinae phylgeny. immunogenetics, 51, 887-897. [ Links ]
Konnai, S., Nagaoka, Y., Takesima, S., Onuma, M. & Aida, Y., 2003. Technical note: DNA typing for ovine MHC-DRB1 using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP). J. Dairy Sci. 86,3362-3365. [ Links ]
Kulski, J.K., Shiina, T., Anzai, T., Kohara, S. & Inoko, H., 2002. Comparative genomic analysis of the MHC: The evolution of class I duplication blocks, diversity and complexity from shark to man. Immunol. Rev. 190, 95-122. [ Links ]
Lotfi, M., Nassiri, M.T.B., Roshanfekr, H. & Fayazi,J., 2012. Polymorphism of Ovar-DRB1 Second Exon with PCR-RFLP Technique in Arabi Sheep Population of Khuzestan Province. J. Anim. and Vet. Advances, 11: 343-345. [ Links ]
Miller, S.A., Dykes, D.D. & Polesky, H.F., 1988. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Research. 16 (3), 1215. [ Links ]
Nikbakht, G., Rezaii, H., Stear, M.J., Talebi, M.A. & Mahmoudzadeh, H. 2012. Allelic polymorphism in the second exon of Ovar-DRB1 in fat-tailed sheep. Vet. J. 2012 Jun; 192(3), 547-9. [ Links ]
Schwaiger, F.W. & Epplen, J.T., 1995. Exonic MHC-DRB polymorphisms and intronic simple repeat sequences: Janus' faces of DNA sequence evolution. Immunol Rev 143,199-224. [ Links ]
Schwaiger, F.W., Maddox, J., Ballingall, K. & Buitkamp, J., 1996. The ovine major histocompatibility complex. In: The major histocompatibility complex region of domestic animal species (Schook LB and Lamont SJ, eds.). CRC Press, Inc., Boca Raton, Fla, USA, 121-176. [ Links ]
Shen, H., Han, G., Jia, B., Jiang, S. & Du, Y., 2014. MHC-DRB1/DQB1 Gene polymorphism and its association with resistance/susceptibility to cystic echinococcosis in Chinese Merino Sheep. Hindawi Publishing Corporation J. of Parasitology Research, Article ID 272601. [ Links ]
Takeshima, S.N., Miyasaka, T., Polat, M., Kikuya, M., Matsumoto, Y., Mingala, C.N., Villanueva, M.A., Salces, A.J., Onuma, M. & Aida, Y., 2014. The great diversity of major histocompatibility complex class II genes in Philippine native cattle. Meta. Gene. 2, 176-190. [ Links ]
Tizard, I.R., 2004. Acquired immunity: Antigen-Presenting Receptors. In: Vet Immunology: an Introduction. Elsevier, Philadelphia, Pa, USA, 67-77. [ Links ]
Received 14 March 2016
Accepted 7 September 2016
Published 31 October 2016
# Corresponding author: Fatma ILHAN, E-mail: fatmailhan@selcuk.edu.tr