Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Animal Science
On-line version ISSN 2221-4062
Print version ISSN 0375-1589
S. Afr. j. anim. sci. vol.44 n.2 Pretoria Feb. 2014
Association of polymorphism in the alpha-1-antitrypsin gene with milk production traits in Holstein dairy cows
M. Heihavand-Kheiripour; A.H. Mahdavi#; H.R. Rahmani; M. Soltani-Ghombavani; M.A. Edriss
Department of Animal Sciences, College of Agriculture, Isfahan University of Technology, Isfahan 8415683111, Iran
ABSTRACT
Alpha-1-antitrypsin (A1AT) as a strong protease inhibitor plays a major role in the protection of tissues against proteolytic destruction by neutrophil elastase. Existence of this protein in the mammary gland may increase the survival of milk proteins such as lactoferrin and lysozyme. The biological role of A1AT in tissues such as the mammary gland and results from previous quantitative trait loci (QTL) studies have provoked an investigation of A1AT as a candidate gene influencing milk production traits. The present study assessed the association of a single nucleotide polymorphism (SNP) at position 5504 of the A1AT gene with 305-day milk yield, milk fat and protein percentage and somatic cell score (SCS) in 408 Iranian Holstein cows. The polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) technique was used for genotyping. The effect of the genotype on the traits of interest was analysed using the general linear models (GLM) procedure of SAS software. The frequencies of alleles A and B were 0.633 and 0.367, respectively, and genotypes showed a significant deviation from Hardy-Weinberg equilibrium, which can be a result of selection for milk production traits. Results of association analysis showed a significant relationship between the SNP in the A1AT gene, and fat and protein percentage. Cows of genotype AB had higher milk fat percentage than those of genotype AA. Cows with genotype AA showed a lower milk protein percentage than those carrying genotype AB. When the association of this polymorphism with fat and protein percentage is considered, the SNP could be implemented as a marker in breeding programmes for these traits.
Keywords: A1AT gene, candidate gene, mammary gland, PCR-RFLP
Introduction
Quantitative traits in dairy cattle are controlled by a large number of genes and environmental factors. Several genes that were selected based on their biological actions or that are located in genome regions containing previously identified QTLs have been regarded as candidate genes affecting milk production traits and SCS in dairy cattle. Genes, markers or QTLs can be considered for selection of desirable genotypes at an early age in the life of an animal. Marker-assisted selection (MAS) as an efficient tool in animal breeding is believed to increase the annual rate of genetic gain in livestock breeding programmes (Meuwissen & Goddard, 1996; Ruane & Colleau, 1996). The expected genetic gain through MAS is estimated to be 44.7% to 99.6% (Edwards & Page, 1994). For effective use of individual markers or genes in animal breeding programmes, existence of an association between them and traits of interest is inevitable.
A1AT belongs to the superfamily of serpins, or serine proteinase inhibitors with a molecular mass of 52 kDa, which contains 15% carbohydrate (Carrell et al., 1982). The primary role of A1AT is to protect tissues against proteolytic digestion by neutrophil elastase (Blank & Brantly, 1994). Possible roles of A1AT in the immune response include inhibition of lymphocyte toxicity and of chemotaxis (Blank & Brantly, 1994). A1AT is synthesized primarily in the liver, but is also expressed by blood monocytes and macrophages (Perlmutter et al., 1985), intestinal epithelium cells (Molmenti et al., 1993), breast carcinoma cells (Finlay et al., 1993) and the cornea (Twining et al., 1994). In addition, it has been reported that A1AT protein is produced by the human mammary gland and may increase the survival of milk proteins such as lactoferrin and lysozyme (Chowanadisai & Lonnerdal, 2002). Hence, A1AT may affect the susceptibility of dairy cattle to mastitis through its possible role in the protection of lactoferrin from proteolytic degradation in the mammary gland. Mastitis is caused by the invasion and multiplication of pathogenic microorganisms in the mammary gland, leading to reduced synthetic capacity of the mammary gland and compositional changes in the milk produced by infected cows (Harmon et al., 1994). Lactoferrin has been demonstrated to inhibit in vitro growth of mastitis-causing pathogens, and the most potent activity of the protein is against Esherichia coli and Pseudomonas aeruginosa (Kutila, 2004; Chaneton et al., 2008). A direct inhibitory activity of lactoferrin has been demonstrated against Staphylococcus aureus, both in vitro and in vivo (Lacasse et al., 2008). The mean concentration of lactoferrin in the milk of cows with subclinical mastitis has been reported to be higher than in milk of normal cows (Hagiwara et al., 2003). Reported concentrations of A1AT in human milk range from 0.1 to 0.4 g/L in early lactation, with a subsequent decrease as lactation progresses (Lindberg et al., 1979; McGilligan et al., 1987; Davidson et al., 1990). The bovine A1AT gene is located on chromosome 21 (Khatib et al., 2004) and consists of five exons, spanning approximately 9 kb of genomic DNA, which encodes a protein of 416 amino acids (Li et al., 2010). Five SNPs (single nucleotide polymorphisms) at positions 164, 269, 284, 407 and 989 of the bovine A1AT gene have been identified in different cattle populations (Khatib et al., 2004).
Owing to previous QTLs affecting production traits and SCS identified on bovine chromosome 21 and the biological role of A1AT protein in human (Li et al., 2010), the A1AT gene has been proposed as a candidate affecting these traits (Khatib et al., 2005). Heyen et al. (1999) reported a putative QTL affecting milk yield and protein yield at position 56 cM from the centromere of the chromosome. In addition, Rodriguez-Zas et al. (2002) found a QTL located at 56 cM associated with SCS and milk protein yield. Mosig et al. (2001) mapped a QTL at 67.3 cM affecting milk protein percentage. Khatib et al. (2005) found a significant association between the A1AT gene and milk yield, fat yield and protein percentage in North American Holstein population. In a similar study, Li et al. (2010) reported the relationship of this gene with 305-day milk yield, and fat and protein percentage in Chinese Holstein. The present study was designed to investigate the possible association of an SNP at position 5504 of the A1AT gene with milk-related traits and SCS in Iranian Holstein cows.
Materials and Methods
A total of 408 Iranian Holstein cows that had registered phenotypic data for 305-day milk yield, milk fat and protein percentage and SCS were randomly selected from five dairy herds in Isfahan province. Blood samples of the animals were collected using vacuum venoject tubes containing EDTA and stored at -20 °C for DNA extraction. Data for 305-day milk yield, milk fat and protein percentage and SCS were obtained from Vahdat Industrial Agriculturists & Dairymen Cooperative of Isfahan.
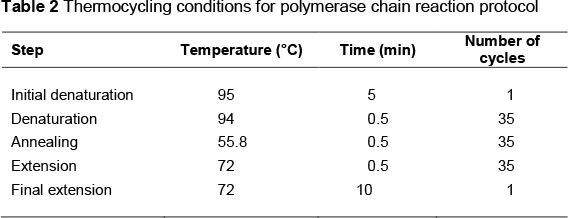
Genomic DNA extraction was carried out by the salting out method (Miller et al., 1988). Genotypes were determined using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) technique. PCR was used to amplify the 448-bp DNA fragments containing the SNP at position 5504 of the A1AT gene. Each PCR reaction was carried out in a total volume of 20 µL containing 50 ng genomic DNA, 10 pmoL of each primer, 2 µL 10X PCR buffer, 2 mM MgCl2, 200 µM dNTPs and 2.5 units of Taq DNA polymerase. Sequences of the primers used in PCR were previously reported by Khatib et al. (2005). The sequences of the primers were as follows:
F: 5'-ATGGCACTCTCCATCACGCG-3'
R: 5'-CCACTAGCTTTGCACTCTCA-3'
The PCR products were digested with 5 U of SphI enzyme (Fermentas, Germany) in 20 µL of reaction volume at 37 °C for 16 h. The digested products were separated by horizontal electrophoresis on 1.2% agarose gel in 0.5 x TBE stained with ethidium bromide and visualized under UV light. The A allele (uncut) was indicated by a band at 448 bp and the B allele (cut) resulted in two bands at 313 bp and 135 bp (Figure 1).
The allele and genotype frequencies were estimated by direct counting, and deviation from Hardy-Weinberg equilibrium was examined using POPGENE v. 1.31 (Yeh et al., 1997). For the association studies, the traits of interest were analysed using the least square method of the general linear model (GLM) procedure of SAS software (SAS, 1994) according to the following model:
Yijk= µ + Gi + HYSj + b1 (Wijk- W) + b2 (Nijk - N) b3 (Zijk-Z) + eijk
where Yijk is value for each trait; µ is overall mean; Gi is the effect of the ith genotype (AA, AB and BB); HYSj is the combined effect of herd, year and season of parturition; b1 is the linear regression coefficient of milk yield; Wijk is milk yield; W is mean milk yield; b2 is the linear regression coefficient of open days; Nijk is open days; N is mean open days; b3 is the linear regression coefficient of days in milk; Zijk is days in milk; Z is mean days in milk; eijk is the random residual effect.
Results
Using the PCR-RFLP technique, both alleles (A and B) and three genotypes (AA, BB, and AB) of the locus of interest were identified in all five herds. Genotype AA was characterized by a fragment of 448 bp, while genotype BB was determined by two fragments of 313 and 135 bp. AB individuals were distinguished by three fragments of 448, 313 and 135 bp. The genotype and allele frequencies of the polymorphism in Iranian Holstein cows are listed in Table 2. The result of the test for Hardy-Weinberg equilibrium for the polymorphism showed that the genotypes in herds 1, 4 and 5 did not show significant deviation from Hardy-Weinberg equilibrium proportions (P >0.05), while herds 2, 3 and the overall population were not distributed according to Hardy-Weinberg equilibrium (P <0.01).
The results of the analysis of association between the A1AT gene and the traits of interest and allele substitution effects (of allele B) are presented in Table 3.
Association analysis in the present study revealed a statistically significant relationship between the SNP in the A1AT gene and fat percentage. Cows with genotype AB showed higher milk fat percentage in comparison with those of genotype AA (P <0.05). As shown in Table 3, substituting the A allele with the B allele increased the percentage of milk fat by 0.07%. Additionally, there was a significant association in the present study between the polymorphism in the A1AT gene and protein percentage. Cows carrying genotype AA exhibited lower protein percentages than those with genotype AB (P <0.05). The substitution of the A allele for the B allele increased the percentage of milk protein by 0.02%. No significant association of the A1AT polymorphism with 305-day milk yield and SCS was evident in the current study (P >0.1).
Discussion
The PCR-RFLP method was useful for detecting an SNP at position 5504 of the A1AT gene and determining the association between this polymorphism and milk production traits and SCS in Iranian Holstein cows. The allele frequencies observed in our study were in line with the results of Li et al. (2010) who reported frequencies of 0.51, 0.49 for alleles A and B, respectively, in Chinese Holstein population.
Association of the SNP with fat percentage revealed in the present study was in agreement with the findings of Li et al. (2010) who showed a lower fat percentage in the AA genotype compared with the BB genotype in Chinese Holstein. In contrast, Khatib et al. (2005) reported no association of the A1AT gene with fat percentage in North American Holstein population. The association of the polymorphism in the A1AT gene with protein percentage found in this study is in accordance with the conclusions of Khatib et al. (2005) and Li et al. (2010). Analysing the association of five SNPs at positions 164, 269, 284, 407 and 989 of the A1AT gene with milk production traits, Khatib et al. (2005) reported an association of the gene with protein percentage in North American Holstein population. Li et al. (2010) showed that an SNP at position 8178 (but not at position 5504) of the gene was associated with protein percentage in Chinese Holstein population. The SNP in the present study is a silent mutation that does not cause alteration in the amino acid sequence and the function of A1AT protein (Li et al., 2010). Hence, the SNP is not a functional mutation, but may be in linkage disequilibrium with a functional polymorphic site in the A1AT or other closely linked gene(s) influencing milk-related traits (Khatib et al., 2005).
Lack of an association between the A1AT gene and SCS in the present study is in line with the results from Li et al. (2010). In contrast, Khatib et al. (2005) found an effect of this gene on SCS in North American Holstein cattle. The results of our study did not show any relationship between the A1AT gene and 305-day milk yield. Contrary to our findings, Khatib et al. (2005) and Li et al. (2010) revealed an effect of this gene on breeding value for milk yield and 305-day milk yield, respectively. Furthermore, Khatib et al. (2005) found an association of the A1AT gene with fat yield. Differences between the results in the present study and those of other researchers might be due to these aspects: i) Different environments to which populations were exposed (genotype by environment interaction). Effect or expression of genes varies depending on the environmental conditions such as feeding, climate and management, which may differ in various countries; ii) Different genetic background of the animals and possible interactions between the A1AT polymorphism and background genes. Dairy cattle breeders in Iran put more emphasis on increasing milk yield than on milk composition, whereas in other countries both milk yield and composition are emphasized. Different breeding goals and selection criteria in different countries will lead to differences in genetic background of cattle populations over generations; iii) Different statistical models used to analyse the data. For the traits of interest in this study, phenotypic values were available. So, to adjust the data, the researchers included some effects in the statistical model which may be different from those included by others. The differences in statistical models can lead to different results. According to the findings of the present study and reports from other authors, it can be postulated that the A1AT gene may be the actual QTG (quantitative trait gene) or in close linkage disequilibrium with the QTG underlying QTLs affecting milk production traits in cattle (Khatib et al., 2005).
Conclusion
In summary, the researchers found an association between polymorphism in position 5504 of the A1AT gene and fat and protein percentage in Iranian Holstein cows. These findings support the results of similar studies by Khatib et al. (2005) and Li et al. (2010) in North American and Chinese Holstein cattle populations, respectively. In conclusion, the polymorphism in this gene could be used as a marker in breeding programmes for selection of dairy cattle with increased fat and protein percentage. However, further association studies of various polymorphisms in different regions of the A1AT gene with these traits are necessary in other breeds and populations of cattle in order to verify the effects of this gene as a candidate on milk production traits and SCS.
Acknowledgement
The authors gratefully acknowledge Isfahan University of Technology for supporting this research financially. The authors also thank Vahdat Industrial Agriculturists & Dairymen Cooperative of Isfahan province of Iran for providing data for milk-related traits.
References
Blank, C.A. & Brantly, M., 1994. Clinical features and molecular characteristics of alpha 1- antitrypsin deficiency. Ann. Allergy 72, 105-120. [ Links ]
Carrell, R.W., Jeppsson, J.O., Laurell, C.B., Brennan, S.O., Owen, M.C., Vaughan, L. & Boswell, D.R., 1982. Structure and variation of human alpha-1-antitrypsin. Nature 298, 329-334. [ Links ]
Chaneton, L., Tirante, L., Maito, J., Chaves, J. & Bussmann, E., 2008. Relationship between milk lactoferrin and etiological agent in the mastitic bovine mammary gland. J. Dairy Sci. 91, 1865-1873. [ Links ]
Chowanadisai, W. & Lönnerdal, B., 2002. al-Antitrypsin and antichymotrypsin in human milk: Origin, concentrations, and stability. Am. J. Clin. Nutr. 76, 828-833. [ Links ]
Davidson, L.A. & Lönnerdal, B., 1990. Fecal alpha-1-antitrypsin in breast-fed infants is derived from human milk and is not indicative of enteric protein loss. Acta Paediatr. Scand. 79, 137-141. [ Links ]
Edwards, M.D. & Page, N.J., 1994. Evaluation of marker-assisted selection through computer simulation. Theor. Appl. Genet. 88, 376-382. [ Links ]
Finlay, T.H., Tamir, S., Kadner, S.S., Cruz, M.R., Yavelow, J. & Levitz, M., 1993. Alpha-1-antitrypsin- and anchorage-independent growth of MCF-7 breast cancer cells. Endocrinology 133, 996-1002. [ Links ]
Hagiwara, S., Kawai, K., Anri, A. & Nagahata, H., 2003. Lactoferrin concentrations in milk from normal and subclinical mastitic cows. J. Vet. Med. Sci. 65, 319-323. [ Links ]
Harmon, R.J., 1994. Physiology of mastitis and factors affecting somatic cell counts. J. Dairy Sci. 77, 2103-2112. [ Links ]
Heyen, D.W., Weller, J.I., Ron, M., Band, M., Beever, J.E., Feldmesser, E., Da, Y., Wiggans, G.R., VanRaden, P.M. & Lewin, H.A., 1999. A genome scan for QTL influencing milk production and health traits in dairy cattle. Physiol. Genomics 1, 165-175. [ Links ]
Kashi, Y., Hallerman, E. & Soller, M., 1990. Marker-assisted selection of candidate bulls for progeny testing program. Anim. Prod. 51, 63-74. [ Links ]
Khatib, H., 2004. Monoallelic expression of the protease inhibitor gene in humans, sheep, and cattle. Mamm. Genome 16, 50-58. [ Links ]
Khatib, H., Heifetz, E. & Dekkers, J.C.M., 2005. Association of the Protease Inhibitor Gene with production traits in Holstein dairy cattle. J. Dairy Sci. 88, 1208-1213. [ Links ]
Kutila, T., Suojala, L., Lehtolainen, T., Saloniemi, H., Kaartinen, L., Tähiti, M., Seppälä, K. & Pyörälä, S., 2004. The efficacy of bovine lactoferrin in the treatment of cows with experimentally induced Escherichia coli mastitis. J. Vet. Pharmacol. Ther. 27, 197-202. [ Links ]
Lacasse, P., Lauzon, K., Diarra, M.S. & Petitclerc, D., 2008. Utilization of lactoferrin to fight antibiotic-resistant mammary gland pathogens. J. Anim. Sci. 86, 66-71. [ Links ]
Li, Q.L., Zhang, Z.F., Wang, C.F., Yang, H., Wang, H.M., Li, J.B., Huang, J.M. & Zhong, J.F., 2010. Association of polymorphism of the alpha 1-antitrypsin gene with milk production traits in Chinese Holstein. S. Afr. J. Anim. Sci. 40, 113-120. [ Links ]
Lindberg, T., 1997. Protease inhibitors in human milk. Pediatr Res. 13, 969-972. [ Links ]
McGilligan, K.M., Thomas, D.W. & Eckhert, C.D., 1987. Alpha-1-antitrypsin concentration in human milk. Pediatr Res. 22, 268-270. [ Links ]
Meuwissen, T.H. & Van Arendonk, J.A., 1992. Potential improvements in rate of genetic gain from marker-assisted selection in dairy cattle breeding schemes. J. Dairy Sci. 75, 1651-1659. [ Links ]
Meuwissen, T.H.E. & Goddard, M.E., 1996. The use of marker haplotypes in animal breeding schemes. Genet. Sel. Evol. 28, 161-176. [ Links ]
Miller, S.A., Dykes, D.D. & Polesky, H.F., 1988. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16, 1215. [ Links ]
Molmenti, E.P., Perlmutter, D.H. & Rubin, D.C., 1993. Cell-specific expression of a1-antitrypsin in human intestinal epithelium. J. Clin. Invest. 92, 2022-2034. [ Links ]
Mosig, M., Lipkin, E., Khutoreskaya, G., Tchourzyna, E., Soller, M. & Friedmann, A., 2001. A whole genome scan for quantitative trait loci affecting milk protein percentage in Israeli-Holstein cattle, by means of selective milk DNA pooling in a daughter design, using an adjusted false discovery rate criterion. Genetics 157, 1683-1698. [ Links ]
Nonnecke, B.J. & Smith, K.L., 1984. Inhibition of mastitic bacteria by bovine milk apo-lactoferrin evaluated by in vitro microassay of bacterial growth. J. Dairy Sci. 67, 606-613. [ Links ]
Perlmutter, D.H., Cole, F.S., Kilbridge, P., Rossing, T.H. & Colten, H.R., 1985. Expression of the a1-proteinase inhibitor gene in human monocytes and macrophages. Proc. Natl. Acad. Sci. USA. 82, 795-799. [ Links ]
Rodriguez-Zas, S.L., Southey, B.R., Heyen, D.W. & Lewin, H.A., 2002. Interval and composite interval mapping of somatic cell score, yield, and components of milk in dairy cattle. J. Dairy Sci. 85, 3081-3091. [ Links ]
Ruane, J. & Colleau, J.J., 1996. Marker-assisted selection for a sex-limited character in a nucleus breeding population. J. Dairy Sci. 79, 1666-1678. [ Links ]
SAS, 1994. Statistical Analysis Systems User's Guide. Statistics Version 6. SAS Inst., Inc. Cary, N.C., USA. [ Links ]
Twining, S.S., Fukuchi, T., Yue, B.Y., Wilson, P.M. & Boskovic, G., 1994. Corneal synthesis of alpha 1-proteinase inhibitor (alpha 1-antitrypsin). Invest. Ophthalmol. Vis. Sci. 35, 458-462. [ Links ]
Yeh, F.C., Yang, R.C., Boyle, T.B.J., Ye, Z.H. & Mao, J.X., 1997. Popgene, the user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Center, University of Alberta, Canada. [ Links ]
Received 20 November 2013
Accepted 14 February 2014
First published online 11 June 2014
# Corresponding author: mahdavi@cc.iut.ac.ir














