Serviços Personalizados
Artigo
Indicadores
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares em Google
Similares em Google
Compartilhar
South African Journal of Animal Science
versão On-line ISSN 2221-4062
versão impressa ISSN 0375-1589
S. Afr. j. anim. sci. vol.43 no.4 Pretoria 2013
Genetic diversity of four protected indigenous chicken breeds in China using microsatellite markers
Lin WeiI, II; Bin ChenII, #; Xiao-ying LiI; Sheng-gui LiuI; Jing-jing WangII
IThe Department of Life Science, Huaihua University, Huaihua 418008
IICollege of Animal Science and Technology, Hunan Agriculture University; Key Laboratory of Hunan Province for Livestock and Poultry Breeds Improvement, Changsha 410128, China
ABSTRACT
The genetic diversity of four protected indigenous chicken breeds was evaluated with 25 microsatellite markers. Polymorphism information content (PIC), heterozygosity with the estimator of genetic differentiation FST and Nei's genetic distance were evaluated. The results showed that these four protected local chicken populations showed high levels of diversity. The proportion of inter-population subdivision among the four protected local chicken populations was 16.0%. The average heterozygosity was 0.514, 0.581, 0.567 and 0.589 in Dongan, Xuefeng black-bone, Xianghuang and Taoyuan chickens, respectively, while the average PIC estimates were 0.455, 0.581, 0.557 and 0.576. A phylogenetic tree was constructed using genetic distance and the neighbour-joining method. Its topology reflects the general pattern of genetic differentiation among the four chicken breeds. The results also showed high genetic diversity and genetic variation among all the breeds. The information about the four local breeds estimated by microsatellite analysis may be useful as an initial guide for the effective conservation of chicken genetic diversity and developing conservation strategies.
Keywords: Native chicken; genetic variation; microsatellite DNA
Introduction
Chicken is one of the most widely distributed livestock in China. It plays a significant role as a source of income and high-quality protein for humankind. Indigenous chickens appear to possess enormous genetic diversity, especially in adaptive traits, and the ability to survive harsh conditions and under minimum feeding regimens (Qu et al., 2006). Successful preservation and utilization of these local types depend on accurate assessment of genetic diversity and genetic structure. Four indigenous chicken breeds, namely Dongan, Xianghuang, Taoyuan and Xuefeng black-bone chickens were included in the National Poultry Genetic Resource Directory, and play an important role in socio-economic development and ecological values in Hunan Province. In addition, their precious values as genetic resources are used for the genetic improvement of chicken breeds. Dongan are special meat-type chickens, and have good meat characteristics, high nutritional value and other excellent traits, such as yellow feathers, yellow skins and yellow shanks (Qu et al, 2006). The Xuefeng black-bone chicken is a meat and egg-type chicken, whose qualities include black meat, black bone, black beak and black feet, which are rich in nutritive and medicinal value, and were formed through long-term natural selection in the Xuefeng mountainous region in Hunan Province (Wei et al., 2008). Xianghuang and Taoyuan chickens are excellent meat and egg-type local chicken breeds, which come mainly from Changsha and Taoyuan counties in Hunan Province, respectively (Gao et al., 2008). So far, limited genetic diversity research has been conducted on these four chicken breeds, except for the study of Wei et al. (2008) on Xuefeng black-bone chickens.
Many molecular markers have become excellent means for the study of genetic variation (Chang et al, 2005; Chen et al., 2003), such as random amplified polymorphic DNAs (RAPD), amplified fragment length polymorphisms (AFLP), microsatellite DNA, and sequence-related amplified polymorphism (SRAP) (Zietkiewicz et al., 1994; Li et al., 2001). Among these DNA markers, microsatellites are widely used since they are numerous, randomly distributed in the genome, highly polymorphic, and with co-dominant inheritance (Groen et al., 1994; Kavaca et al., 1999). Many microsatellites have been mapped in chickens, and are used to study the genetic relationships among breeds (Tadano et al, 2007; Kaya et al, 2008). The purpose of this study is to estimate the level of genetic differentiation and phylogenetic relationships among the four indigenous chicken breeds in China. This information can contribute to the conservation and utilization of local chicken breeds.
Materials and Methods
Samples were obtained from 179 unrelated individuals, representing four indigenous chicken breeds in Hunan Province. These included 50 Dongan chickens (DA) from Dongan county, 50 Xianghuang chickens (XH) from Liuyang county, 29 Taoyuan chickens (TY) from Taoyuan county and 50 Xuefeng black-bone chickens (XF) from Hongjiang city. Blood samples (3 mL) were collected with syringes from the wing vein into a tube containing DNA preservation solution as an anti-coagulating agent. All samples were stored at -80 °C for further analysis.
Genomic DNA was isolated from blood using a phenol/chloroform extraction method (Sambrook, 2002). The DNA was quantified with a spectrophotometer, comparing band intensities with known standards of DNA marker on 1.5% agarose gel. The working solution of DNA (approx. 25 ng/µL) was dissolved in sterile double-distilled water. In a preliminary experiment, 40 SSR primers were tested on four random individuals from each breed. Based on the amplification result, 25 microsatellite loci were further investigated, which were listed in Table 1. All primers were synthesized by ShenggongBio-Tech. Co., Ltd. Shanghai, China.
Polymerase chain reaction (PCR) amplifications were performed on PTC-200 thermal cyclers. A total reaction volume of 8 µL with 1 µL of 10 x buffer, 0.6 µL of 25 mmol MgCl2, 0.2 µL of 10 mmol dNTPs, 0.1 µL of 5 U/µL Taq DNA polymerase, 0.3 µL of 10 pmol/µL each primer, and approximately 50 ng of genomic DNA were used. The reaction was carried out by initial denaturation at 94 °C for 3 min, and then denaturing at 94 °C for 30 s, annealing at the temperature optimized for each primer pair for 30 s and extending at 72 °C for 30 s for 35 cycles, followed by an extra extension step at 72 °C for 5 min. The optimized annealing temperatures of different primer pairs are listed in Table 1. The amplification products were separated by electrophoresis on 12% non-denaturing polyacrylamide gels and visualized by silver staining (Su et al., 2006). The images data were analysed with Kodak Digital Science ID Image Analysis Software.
Based on microsatellite genotyping and allele frequencies, the number of alleles, effective number of alleles (Ne), observed heterozygosity (Ho), (Nei, 1987) expected heterozygosity (He), and Wright's (1978) fixation index (Fis) were estimated using the computer software package PopGene version 1.31 (Yeh et al., 1997). Allele frequencies obtained from the microsatellite genotypes were used to calculate PIC (polymorphism information content) values (Botstein et al, 1980) using the computer software package Cervus 3.0 (Marshall et al., 1998; Kalinowski et al., 2007) in order to measure the information obtained by a microsatellite. Based on microsatellite genotyping, Nei's (1978) unbiased genetic distance between breeds was estimated. Software FSTAT (version 2.9.3.2) was used to test pairwise linkage equilibriums at all loci over any two groups to calculate the pairwise genetic differentiation FST (Weir & Cockerham, 1984). These results were used to construct phylogenetic trees by neighbour-joining cluster analysis with the appropriate options of computer software Mega Version 4.0 (Tamura et al., 2007).
Results
The genetic diversity and differentiation among the four local chicken breeds at the 25 microsatellite loci were estimated. The numbers of alleles per locus and the size range of alleles are listed in Table 1. All loci were polymorphic in the four breeds. The observed numbers of alleles varied from 3 (MCW0330) to 9 (MCW0094) and the mean number of alleles across all loci was 5.84. The observed heterozygosity (Ho) ranged from 0.268 (LEI0254) to 0.726 (LEI0098) (Table 3). The expected heterozygosity (He) was quite high, ranging from 0.448 (LEI0254) to 0.861 (MCW0224) (Table 2). The PIC among loci was highest for MCW0224 (0.854) and lowest for ADL0210 (0.447) (Table 2).
There was highly significant genetic divergence across the four breeds for every locus. The FST values ranged from 0.058 (MCW0330) to 0.243 (MCW0097). Using the multilocus FST, approximately 16.0% of the total genetic variation can be explained by breed differences, and the remaining 84.0% was owing to the differences among individuals (Table 2).
The estimated multilocus heterozygosities varied from H = 0.514 in DA to H = 0.589 in TY. The mean effective allele number was between 3.20 (XF) and 3.72 (TY). Among breeds, the mean PIC value was 0.455, 0.581, 0.557 and 0.576 for DA, XF, XH and TY, respectively. Wright's fixation index (Fis) values ranged from -0.16 (XF) to 0.14 (DA). Deviation from Hardy-Weinberg equilibrium (HWE) between loci and breeds was tested with FSTAT (version 2.9.3, Goudet, 2001). Significance levels were adjusted using Bonferroni correction for multiple testing. Significant deviations (P <0.05) from HWE were observed at the breed level (DA) (Table 3).
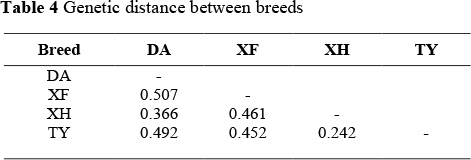
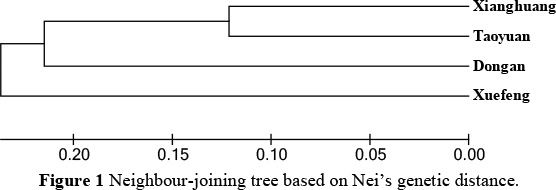
Using Nei's (1978) unbiased genetic distance (Table 4) and the neighbour-joining method, a phylogenetic tree was constructed for the four chicken breeds. The smallest genetic distance, between Xianghuang and Taoyuan chickens, was 0.242. The largest genetic distance, between Dongan and Xuefeng black-bone chickens, was 0.507. The neighbour-joining dendrogramme in Figure 1 was drawn using the genetic distances given in Table 4. The Xianghuang, Taoyuan chickens and Dongan chickens and Xuefeng black-bone breeds were clustered as two groups, in order to support the reliability of this analysis.
Discussion
Choosing scientific microsatellite markers is essential to analyse genetic diversity. In this study, the 25 microsatellite markers with extensive coverage on main genome and high polymorphism worked very well for this purpose. They also demonstrated their utility as informative molecular markers in the four indigenous chicken breeds. The mean number of alleles for all loci was similar among the four breeds (Table 3). The mean effective number of alleles in this research for overall loci was 3.404 (Table 2). Compared with a previous study (Wei et al., 2008), the present research revealed the same microsatellite allele variation in Xuefeng black-bone chickens (Ne, 3.43). The result is similar to that of other studies. For instance, Emara et al. (2002) examined 41 microsatellite markers in three commercial broiler pure lines and reported an average number of alleles per marker of 3.5, 2.8 and 3.1 for each of the lines. Hillel et al. (2003) reported that the mean number of alleles was 3.5 within 52 breeds. Shahbazi et al. (2007) reported a mean number of alleles of 4.5 per locus in Iranian native chickens.
Heterozygosity estimates within the breeds were based on a set of markers showing substantial number of detected alleles and polymorphic information content. This result showed that genetic diversity in the four indigenous chicken breeds was high. A similar result (0.45 - 0.67) was reported by Wimmers et al. (2000) for African, Asian and South American local chickens. However, the mean He recorded in this research is lower than that reported by Zhang et al. (2002) in Chinese native chickens (0.63 - 0.86) and by Shahbazi et al. (2007) in Iranian native chickens (0.62 - 0.74). It was also higher than Hillel et al. (2003), who reported that the average gene diversity within 52 breeds across all 22 loci was 0.47. The variation of expected heterozygosity may be adduced to differences in location, sample size, breed structure and microsatellite markers. The high mean heterozygosity values may be attributed to the low level of inbreeding, low selection pressure and large number of alleles present in one breed. In this study, the high gene diversity value is a reflection of a high intra-breed genetic variation among these chicken breeds in Hunan province in China.
The mean PIC was an ideal index to measure the polymorphism of allele fragments. The mean PIC among loci was 0.663 (Table 2), and almost all markers were highly informative in the four indigenous chicken breeds. The FIS represents a degree of nonrandom mating (deviation from HWE). A positive value for FIS indicates deviation from HWE. In this study, causes of deviations from HWE, such as shortage of samples, selective mating, low levels of polymorphism, were not the major concerns in the breeds. The presence of inbreeding may be a reasonable explanation for the observed lack of agreement with HWE. Significant deviations from HWE were observed for 17 loci in Dongan chickens. These results indicate that the level of gene flow among breeds is restricted. The observed divergence probably reflects the human selection and the bottleneck effect (Maak et al., 2003).
The degree of genetic differentiation among these breeds and the high levels of significance for the inter-breed FST estimations indicate a relatively low gene flow among the four local chicken breeds and a relatively high reproductive isolation. The mean FST value of 0.16 indicates that approximately 16.0% of the total genetic variation is caused by breed differences, whereas the remaining 84.0% is due to differences among individuals within breeds. Chen et al. (2006) reported a mean FST value of 0.16 from 12 chicken breeds using 29 microsatellite markers, and Zhang et al. (2008) reported a mean FST value of 0.142 from seven chicken breeds using 29 microsatellite markers.
The genetic distance among the four protected chicken breeds varied from 0.242 to 0.507, and the mean genetic distance between any given breeds was 0.42, reflecting that these breeds are genetically isolated from each other. Hillel et al. (2003) emphasized that genetic distance measures based on gene frequencies were in good agreement with the genetic diversity of these breeds, indicating that these approaches fit the history of domesticated chickens well. The genetic differentiation found among the four protected local chicken breeds in the neighbour-joining dendrogramme (Figure 1) was confirmed by their breeding origin and evolution.
The information about the four protected local chicken breeds estimated by microsatellite analysis may be useful as an initial guide to defining objectives for designing future investigations of genetic variation and developing conservation strategies. Microsatellite data in this study indicate that the four protected local chicken breeds showed a high within-breed genetic variation, which is a favourable factor when planning conservation and improvement programmes. The effective conservation of genetic diversity in the chicken gene pool relies essentially on the understanding of genetic diversity patterns of chicken breeds in a conservation region, including the levels and distribution of the diversity (Chen et al., 2006). It is particularly important to conserve the chicken's genetic diversity on farm management (or onfarm conservation), because the combination of farmers' diverse needs with the breeds in different ecosystems has created and accumulated wide genetic variation. This study revealed a high genetic diversity within the four protected local chicken breeds. The results from this study indicated that the genetic variation of chicken breeds is still remarkably rich. The considerably rich genetic diversity of chicken breeds in China can be attributed to its complicated local geographical conditions where different farming practices and agro-ecosystems exist. The diversity may also be significantly associated with its rich culture diversity that promotes miscellaneous needs and applications of chicken breeds. Relative isolation of the various areas has probably played a considerable role in reducing the exchange with modern improved chicken breeds. Such a factor has played an essential role in maintaining the genetic diversity of Chinese chicken breeds. In addition, it is important to carry out more studies in future, which can provide us with useful information for the effective conservation of chicken genetic diversity and a roadmap for conservation strategies of chicken genetic resources.
Conclusion
These four protected indigenous chicken breeds throughout Hunan Province in China are resources of considerable socio-economic value. Their genetic diversity was evaluated with 25 microsatellite markers in this study. The result demonstrated that these breeds showed high polymorphism. A phylogenetic tree was constructed using genetic distance and the neighbour-joining method. The four breeds were clustered as two groups, in order to support the reliability of this analysis. The information about the four local breeds that was estimated by microsatellite analysis may be useful as an initial guide to defining objectives for designing future investigations of genetic variation and developing conservation strategies
Acknowledgements
We are grateful to breeders' associations for their help in collecting samples, and thank Xianwei Shi for his comments on the draft paper. This work was supported by Hunan Provincial Natural Science Foundation of China (No: 07JJ6044), the Scientific Research Fund of Hunan Provincial Education Department (No: 06C635) and the Foundation of Key Discipline Construction Projects of Huaihua University.
References
Botstein, D., White, R.L., Skolnik, M. & Davis, R.W., 1980. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32, 314-331. [ Links ]
Chang, G.B., Chang, H., Liu, X.P., Wang, H.Y., Xu, W., Zhao, W.M. & Wang, Q.H., 2005. Study on genetic diversity of wild quail in China with microsatellite DNA markers. Acta Genet. Sin. 32, 795-803. [ Links ]
Chen, H.J., Yue, Y.S., Fan, X.Z., Zhang, C.S. & Du, L.X., 2003. Analysis of genetic diversity of shandong indigenous chicken breeds using microsatellite marker. Acta. Genet. Sin. 30, 855-860. [ Links ]
Chen, G.H., Ji, C.L., Wang, M.Q. & Weigend, S., 2006. Analysis of genetic structure and diversity of 12 Chinese indigenous chicken breeds. Acta Veterinaria et Zootechnica Sinica 37, 105-111. [ Links ]
Cheng, H.H., Levin, I., Vallegjo, R.L., Khatip, H., Dodgson, J.B., Crittenden, L.B. & Hillel, J., 1995. Developing of a genetic map of the chicken with markers of high utility. Poult. Sci. 74, 1855-1874. [ Links ]
Emara, M.G., Kim, H., Zhu, J., Lapierre, R.R., Lakshmanan, N. & Lillehoj, H.S., 2002. Genetic diversity at the major histocompatibility complex (B) and microsatellite loci in three commercial broiler pure lines. Poult. Sci. 81, 1609-1617. [ Links ]
Gao, H.L., Wang, Q.G., Tian, K.X. & Du, S.Q., 2008. Study on utilization and developing prospects of Taoyuan chicks. China Anim. Husb. Vet. Med. 35, 157-158. [ Links ]
Goudet, J., 2001. FSTAT, a program to estimate and test gene diversities and fixation indexes (version 2.3.9). http://www.unil.ch/izea/softwares/fstat.html. [ Links ]
Groen, A.F., Crooijmans, R.P.M.A., Van Kampen, A.J.A., Van der Beek, S., Van der Poel, J.J. & Groenen, M.A.M., 1994. Microsatellite polymorphism in commercial broiler and layer lines, in: Proceedings of the 5th World Congress on Genetics Applied to Livestock Production, Guelph, 7-12 August, 1994, Vol. 21, University of Guelph, Ontario, 94-97. [ Links ]
Hillel, J., Groenen, M.A., Tixier Boichard, M., Korol, A.B., David, L., Kirzhner, V.M., Burke, T., Barre-Dirie, A., Crooijmans, R.P., Elo, K., Feldman, M.W., Freidlin, P.J., Mäki-Tanila, A., Oortwijn, M., Thomson, P., Vignal, A., Wimmers, K. & Weigend, S., 2003. Biodiversity of 52 chicken populations assessed by microsatellite typing of DNA pools. Genet. Sel. Evol. 35, 533-557. [ Links ]
Kalinowski, S.T., Taper, M.L. & Marshall, T.C., 2007. Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 16, 1099-1006. [ Links ]
Kavaca, M., Karaca, F.G., Patel, C. & Emara, M.G., 1999. Preliminary analysis of microsatellite loci in commercial broiler chickens. In: Plant and Animal Genome VII Conference, 17-21 January, San Diego, CA, USA. [ Links ]
Kaya, M. & Yildiz, M.A., 2008. Genetic diversity among Turkish native chickens, Denizli and Gerze, estimated by microsatellite markers. Biochem. Genet. 46, 480-491. [ Links ]
Li, G. & Quiros, C.F., 2001. Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: Its application to mapping and gene tagging in Brassica. Theor. Appl. Genet. 103, 455-461. [ Links ]
Maak, S., Wimmers, K., Weigend, S. & Neumann, K., 2003. Isolation and characterization of 18 microsatellites in the Peking duck (Anas platyrhynchos) and their application in other waterfowl species. Mol. Ecol. Notes 3, 224-227. [ Links ]
Marshall, T.C., Slate, J., Kruuk, L.E.B. & Pemberton, J.M., 1998. Statistical confidence for likelihood-based paternity inference in natural populations. Mol. Ecol. 7, 639-655. [ Links ]
Nei, M., 1978. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89, 583-590. [ Links ]
Nei, M., 1987. Molecular Evolutionary Genetics. Columbia University Press, New York. [ Links ]
Qu, L. J., Li, X.Y., Xu, G.F., Chen, K.W., Yang, H.J., Zhang, L.C., Wu, G.Q, Hou, Z.C., Xu, G.Y. & Yang, N., 2006. Evaluation of genetic diversity in Chinese indigenous chicken breeds using microsatellite markers. Science in China Series C: Life Sciences 49, 332-341. [ Links ]
Shahbazi, S., Mirhosseini, S.Z. & Romanov, M.N., 2007. Genetic diversity in five Iranian native chicken populations estimated by microsatellite markers. Biochem. Genet. 45, 63-75. [ Links ]
Su, Y., Chen, G.H., Long, R.J., Bao, W.B. & Wan, J.H., 2006. Microsatellite analysis of genetic diversity and phylogenetic relationship of meat-type and dual-purpose duck breeds in China. Anim. Biotechnol. Bull. 10, 199-205. [ Links ]
Tadano, R., Nishibori, M., Nagasaka, N. & Tsudzuki, M., 2007. Assessing genetic diversity and population structure for commercial chicken lines based on forty microsatellite analyses. Poult. Sci. 86, 2301-2308. [ Links ]
Tamura, K., Dudley, J., Nei, M. & Kumar, S., 2007. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596-1599. [ Links ]
Wei, L., Liu, S.G. & Shi, X.W., 2008. Genetic diversity of the Xuefeng black bone chicken based on microsatellite markers. Biodiv. Sci. 16, 503-508. [ Links ]
Weir, B.S. & Cockerham, C.C., 1984. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358-1370. [ Links ]
Wimmers, K., Ponsuksili, S., Hardge, T., Valle-Zarate, A., Mathur, P.K. & Horst, P., 2000. Genetic distinctness of African, Asian and South American local Chickens. Anim. Gen. 31, 159165. [ Links ]
Wright, S., 1978. Variability Within and Among Natural Populations. vol 4. The University of Chicago Press, Chicago, USA. [ Links ]
Yeh, F.C., Yang, R.C., Boyle, T.B.J., Ye, Z.H. & Mao, J.X., 1997. PopGene, the user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Centre, University of Alberta, Canada. http://www.ualberta.ca/~fyeh. [ Links ]
Zhang, X., Leung, F.C., Chan, D.K.O., Yang, G. & Wu, C., 2002. Genetic diversity of Chinese native chicken breeds based on protein polymorphism, randomly amplified polymorphic DNA, and microsatellite polymorphism. Poult. Sci. 81, 1463-1472. [ Links ]
Zhang, X.Y, Chen, G.H., Han, W., Zhu, Y.F. & Lu, J.H., 2008. Genetic differentiation and clustering analysis of 7 Chinese indigenous chicken breeds. J. Yunnan Agricultural Univ. 23, 225-229. [ Links ]
Zietkiewicz, E., Rafalski, A. & Labuda, D., 1994. Genome fingerprinting by simple sequence repeat(SSR) anchored polymerase chain-reaction amplification. Genomics 20, 176-183. [ Links ]
Sambrook, J. & Russell, D.W.: Translated by Huang, P.T., Wang, J.X., Zhu, H.C., Zhang, Z.S., Chen, H.P., Fan, M., Yu, W.Y. & He, F.C., 2002. Molecular Cloning: A Laboratory Manual. 3rd ed. Beijing, Science Press, 461-512. [ Links ]
Received 10 December 2012
Accepted29 September 2013
First published online 18 December 2013
# Corresponding author: hnchenbin@163.com