Servicios Personalizados
Articulo
Indicadores
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares en Google
Similares en Google
Compartir
South African Journal of Animal Science
versión On-line ISSN 2221-4062
versión impresa ISSN 0375-1589
S. Afr. j. anim. sci. vol.42 no.5 Pretoria ene. 2012
Estimation of genetic parameters for growth traits in Brangus cattle
F.W.C. NeserI, #; J.B. van WykI; M.D. FairI; P. LuboutII; B.J. CrookIII
IDepartment of Animal, Wildlife and Grassland Sciences, University of the Free State, PO Box 339, Bloemfontein 9300, South Africa
IIBrangus Cattle Breeders' Society, P.O. Box 12465, Brandhof, Bloemfontein, 9324, South Africa
IIIAgricultural Business Research Institute, UNE Armidale NSW 2351, Australia
ABSTRACT
A combination of multiple trait and repeatability models were used to estimate genetic parameters for birth weight (BW), weaning weight (WW), yearling weight (YW), eighteen month weight (FW) and three measurements of mature weight (MW) using 23 768 records obtained from the South African Brangus Cattle Breed Society. The data covered a period of 7 generations from 1985 to 2010. Direct heritability estimates obtained were 0.21 ± 0.024, 0.23 ± 0.021, 0.22 ± 0.025, 0.29 ± 0.029 and 0.24 ± 0.019 for BW, WW, YW, FW and MW, respectively. Maternal heritability estimates for birth weight and weaning weight were 0.05 ± 0.01 and 0.11 ± 0.001, respectively. The direct genetic correlations between the different traits were all positive, ranging from moderate (0.43 ± 0.081) between YW and MW to high (0.99 ± 0.043) between WW and FW.
Keywords: (Co)variance components and ratios, repeatability models, South African Brangus cattle
Introduction
The prediction of breeding values requires knowledge of the magnitude of the (co) variances of random effects in a model. Incorrect (co)variance components could lead to biased breeding values, especially in multiple trait analysis of growth traits, where erosion of records takes place over time because of selection and culling of animals. It could also lead to an incorrect measurement of the effectiveness of genetic selection. As there is limited information available on the genetic parameters for growth traits in Brangus cattle, the aim of the study was to estimate (co)variance components specific for the South African Brangus population which will be used in the national genetic evaluation of the breed.
Material en Methods
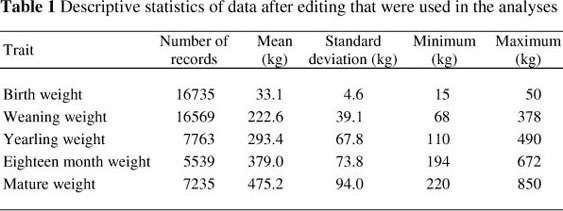
Data from 73 676 animals for birth weight (BW; n = 41 572), weaning weight (WW; n = 23 104), yearling weight (YW; n = 9 114) eighteen month weight (FW; n = 6 450) and mature weight (MW; n = 9 258) were originally available to estimate (co)variance components and subsequent genetic parameters in the breed. All incomplete records, as well as records outside four standard deviations from the mean, were disregarded. Herds with less than three years of recording, as well as contemporary groups with less than five records were also removed from the final data set used for analyses. The final dataset consisted of 53 841 weight records obtained from 23 768 animals in 78 herds, collected over a period of 26 years (1985 to 2010). A total of 1 623 sires and 415 sires of sires, as well as 18 491 dams with 1 028 sires of dams and 5 729 dams of dams were present in the data. The pedigree file consists of 58 973 animals born over a period of 8 generations.
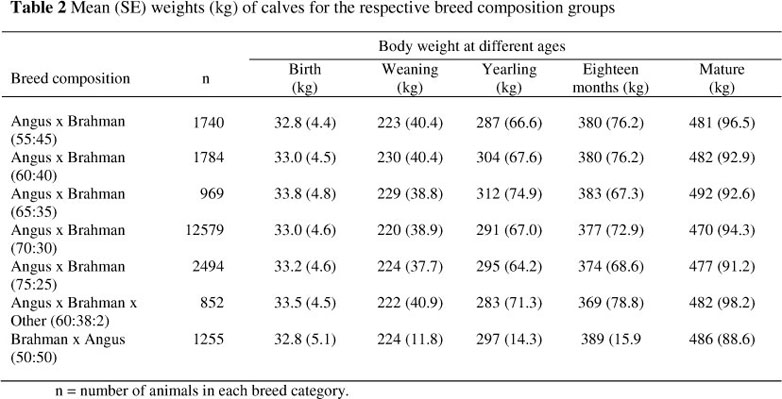
The Brangus is a composite breed and in South Africa, an open register system is used. As different breed combinations are involved, it also means that heterosis can play a role in the estimation of genetic parameters. It is therefore important to include breed composition in the analysis. There were 1 398 different breed combinations that were consolidated into seven broad combinations according to likeness in breed composition and weaning weight performance.
Other fixed effects fitted were a concatenation of herd-year-season, sex and damage. Two distinct calving seasons were identified: from September to March was classified as season one, while April to August was classified as season two. Age of dam was expressed in years, starting with dams of two years and younger. All dams older than six years were grouped together. The following traits were corrected for weighing age to simplify the analysis: weaning-, yearling- and eighteen month weight.
Estimates of (co)variance components were obtained using the ASREML program (Gilmour et al., 2009) fitting single trait animal models. Mature cow weight, defined as the weight of the cow when the calf is weaned, was fitted as a repeated trait, with up to three weights possible per cow.
The following final models were fitted:
BW & WW: y = Xb + Z1a+Z2m+Z3hysxs+e with cov(a,m)=0
YW & AW: y = Xb + Z1a+e
MW: y1,2,3 = Xb + Z1a+e
Where: y = a vector of observations (BW, WW, YW, AW)
y1,2,3 = a vector of repeated observations for mature weight
b = a vector of fixed effects
a = a vector of direct additive genetic effects
m = a vector of maternal additive effects
hysxs = a vector of herd-year-season x sire interaction
e = a vector of residuals
X, Z1, Z2 and Z3 = incidence matrices relating observations to their respective fixed and random effects
It was assumed that:
V(a) = Aσ2a; V(m) = Aσ2m; V(hysxs) = Iσ2hysxs; V(e) = Iσ2e
Where I is an identy matrix, σ2a σ2m> σ2hysxs, and σ2e is the direct additive-, maternal additive-, HYSxS -and environmental variance respectively.
The significance of random effects was tested using the log likelihood ratio tests after inclusion of one random effect in the model. A random effect was considered significant when its inclusion in the model caused a significant improvement in the log likelihood ratio. A chi-square distribution of α = 0.05 at one degree of freedom was used as a test statistic (3.841). When -2 times the difference between the log likelihoods was greater than this critical value, the inclusion of the particular random effect was considered to significantly improve the fit of the model (Swalve, 1993). The estimates obtained in the single trait analyses also act as starting values for the multiple trait analysis.
Results and Discussion
All non-genetic factors tested were significant (P <0.01) for all the traits under consideration and were thus included in the subsequent (co)variance analyses. A summary of the (co)variance components and ratios obtained in the analysis is presented in Table 3.
Direct and maternal heritability estimates for all the traits are within the parameter range described in the literature, albeit in the lower sector. Literature values for direct and maternal heritability estimates for BW vary from 0.07 (direct) and 0.04 (maternal) (Diop & Van Vleck, 1998) to 0.62 (direct) (Schoeman & Jordaan, 1999) and 0.24 (maternal) (Mohiuddin, 1993). The same values for WW vary from 0.07 (direct) (Plasse et al., 2002) and 0.06 (maternal) (Haile-Mariam & Kessa-Mersa, 1995) to 0.57 (direct) (Schoeman & Jordaan, 1999) and 0.21 (maternal) (Diop & Van Vleck, 1998). Direct heritability estimates for the other weight traits from the literature are: YW = 0.16 (Meyer, 1992) to 0.41 (Mohiuddin, 1993), FW = 0.13 (Plasse et al., 2002) to 0.22 (Meyer, 1992; Mostert et al., 1998) and MW = 0.33 (Crook et al., 2010) to 0.50 (Koots et al., 1994a).
The reason for the inclusion of HYSxS is well documented (Meyer, 1987; Neser et al., 1996; Van Niekerk et al., 2004). The high value of the HYSxS ratio for weaning weight (0.10) was, however, surprising, indicating that some re-ranking of sires might occur over different contemporary groups. This is higher than the value obtained by Neser et al. (1996) (0.08) in Bonsmara cattle as well as Pico et al. (2004) (0.06) in Brahman cattle.
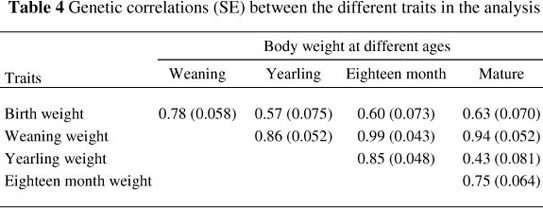
As shown in Table 4, the direct genetic correlations between the different traits were all positive, ranging from moderate (0.44 between YW and MW) to high (0.99 between WW and FW). These results correspond to results obtained by Koots et al. (1994b), Van Niekerk et al. (2004) in Nguni cattle, Pico et al. (2004) in Brahman cattle and Van Niekerk et al. (2006) in Limousin cattle.
Conclusion
Direct heritability estimates were moderate (above 0.21) for all live weight traits while direct genetic correlations between the different traits were all positive, ranging from moderate (0.43) to high. This means that in animal breeding no decision can be taken in isolation. Selection on one trait will thus have consequences on all other traits as well. The estimation of (co)variance components should be seen as the first step in developing a proper breeding objective for the breed.
References
Crook, B.J., Neser, F.W.C. & Bradfield, M.J., 2010. Genetic parameters for cow weight at calving and at calf weaning in South African Simmental cattle. S. Afr. J. Anim. Sci. 40, 140-144. [ Links ]
Diop, M. & Van Vleck, L.D., 1998. Estimates of genetic parameters for growth traits of Gobra cattle. Anim. Sci. 66, 349-355. [ Links ]
Gilmour, A.R., Cullis, B.R., Welham, S.J. & Thompson, R., 1999. ASREML Reference Manual. NSW Agriculture Biometric Bulletin No.3 NSW Agriculture, Orange Agriculture Institute, Forest Road, Orange 2800 NSW, Australia. [ Links ]
Haile-Mariam, M. & Kassa-Mersa, H., 1995. Estimates of direct and maternal covariance components of growth traits in Boran cattle. J. Anim. Breed. 113, 43-55. [ Links ]
Koots, K.R., Gibson, J.P. & Wilton, J.W., 1994a. Analyses of published genetic parameter estimates for beef production traits. 1. Heritability. Anim. Breed. Abstr. 62, 309-388. [ Links ]
Koots, K.R., Gibson, J.P. & Wilton, J.W., 1994b. Analyses of published genetic parameter estimates for beef production traits. 1. Phenotypic and genetic correlations. Anim. Breed. Abstr. 62, 825-853. [ Links ]
Meyer, K., 1987. Estimates of variance due to sire x herd interactions and environmental co-variances between paternal half-sibs for first lactation dairy production. Livest. Prod. Sci. 17, 95-114. [ Links ]
Meyer, K., 1992. Variance components due to direct and maternal effects for growth traits of Australian beef cattle. Livest. Prod. Sci. 31, 179-204. [ Links ]
Mostert, B.E., Groeneveld, E., Rust, T. & Van der Westhuizen, J., 1998. Multitrait variance component estimation of South African beef breeds for growth traits. Proc 6th Wrld Congr. Genetics Appl. Livest. Prod. 23, 145-152. [ Links ]
Mohiuddin, G., 1993. Estimates of genetic and phenotypic parameters of some performance traits in beef cattle. Anim. Breed. Abstr. 61, 495-522. [ Links ]
Neser, F.W.C., Konstantinov, K.V. & Erasmus, G.J., 1996. The inclusion of herd-year-season by sire interaction in the estimation of genetic parameters in Bonsmara cattle. S. Afr. J. Anim. Sci. 26, 75-78. [ Links ]
Pico, B.A., Neser, F.W.C. & Van Wyk, J.B., 2004. Genetic parameters for growth traits in South African Brahman cattle. S. Afr. J. Anim. Sci. 34 (supplement 2), 44-46. [ Links ]
Plasse, D., Verde, O., Fossi, H., Romero, R., Hoogesteijn, R., Bastidas, P. & Bastardo, J., 2002. (Co)variance components, genetic parameters and annual trends for calf weights in a pedigree Braham herd under selection for three decades. J. Anim. Breed. Genet. 119, 141-153. [ Links ]
Schoeman, S.J. & Jordaan, G.F., 1999. Multitrait estimation of direct and maternal (co)variances for growth and efficiency traits in a multibreed beef cattle herd. S. Afr. J. Anim. Sci. 29, 124-136. [ Links ]
Swalve, H.H., 1993. Estimation of direct and maternal (co)variance components for growth traits in Australian Simmental beef cattle. J. Anim. Breed. Genet. 110, 241-252. [ Links ]
Van Niekerk, M. & Neser, F.W.C., 2006. Genetic parameters for growth traits in South African Limousin cattle. S. Afr. J. Anim. Sci. 36, 5 (Suppl 1), 6-9. [ Links ]
Van Niekerk, M., Neser, F.W.C. & Van Wyk, J.B., 2004. (Co)variance components for growth traits in the Nguni cattle breed S. Afr. J. Anim. Sci. 34 (Issue 6, Suppl. 2), 113-115. [ Links ]
Copyright resides with the authors in terms of the Creative Commons Attribution 2.5 South African Licence. See: http://creativecommons.org/licenses/by/2.5/za/ Condition of use: The user may copy, distribute, transmit and adapt the work, but must recognise the authors and the South African Journal of Animal Science
# Corresponding author: neserfw@ufs.ac.za














