Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Animal Science
On-line version ISSN 2221-4062
Print version ISSN 0375-1589
S. Afr. j. anim. sci. vol.41 n.3 Pretoria Jan. 2011
Genetic parameter estimates for live animal ultrasound measures of carcass traits in South African Angus cattle
M.M. SerobaI,II; A. MaiwasheII,*; K.A. NephaweIII; D. NorrisIV
ILimpopo Department of Agriculture, Private Bag X45, Tzaneen 0850, South Africa
IIAgricultural Research Council, Animal Production Institute, Private Bag X2, Irene 0062, South Africa
IIILimpopo Department of Agriculture, Private Bag X9487, Polokwane 0700, South Africa
IVUniversity of Limpopo, Private Bag X1106, Sovenga 0727, South Africa
ABSTRACT
Genetic parameters were estimated for body weight at scanning (SCW) and live animal ultrasound measures of rump fat thickness (P8FT), rib fat thickness (RBFT), rib eye-muscle area (REA) and intramuscular fat percentage (IMF) for South African Angus beef cattle. A multiple-trait animal model was used to estimate genetic parameters using the Residual Maximum Likelihood procedure (REML). Ultrasound data was used on 1 122 animals that participated in the South African National Beef Cattle Improvement Scheme. The fixed effects that were significant included contemporary group, sex and age of the animal at scanning. Random effects considered were the direct additive genetic effect and the residual. Heritability estimates were 0.38 ± 0.08, 0.34 ± 0.06, 0.33 ± 0.05, 0.32 ± 0.06 and 0.38 ± 0.06 for SCW, P8FT, RBFT, REA and IMF respectively. Genetic correlation between SCW and REA was high (0.79 ± 0.07). Genetic correlations with SCW were 0.24 ± 0.14 for P8FT, 0.25 ± 0.12 for RBFT and 0.29 ± 0.11 for IMF. Measures of body fat were genetically highly correlated with one another, with correlations ranging from 0.71 to 0.99. The magnitude of heritability estimates obtained in the current study indicates that genetic improvement for ultrasound indicators of carcass traits should be effective. However, the indirect selection of carcass traits using ultrasound data, should consider the antagonistic genetic relationship between subcutaneous and intramuscular fat.
Keywords: Heritability, genetic correlations, beef cattle, real-time ultrasound
Introduction
Traditionally, national beef cattle genetic evaluation for carcass traits relied on data collected primarily from organised carcass progeny testing programmes. However, progeny testing is expensive to implement and the number of sires that could be reliably tested are therefore limited. Progeny testing programmes in lowly reproductive species such as beef cattle are characterised by low selection intensity and a prolonged generation interval which in turn hinders genetic improvement. Live animal ultrasound measures of carcass traits were recently introduced to supplement progeny testing programmes or for usage as sole source of carcass data in beef cattle breeding programmes (Crews et al., 2004; MacNeil & Northcutt, 2008). Ultrasonic measures are cheaper to collect than actual carcass measurements and therefore could be useful in providing early genetic predictions for carcass traits. Also, animals with ultrasound data could be used for breeding in contrast to those with carcass measurements.
The heritability for ultrasound traits is moderate to high (Reverter et al., 2000; Kemp et al., 2002; Stelzleni et al., 2002). Computation on genetic relationships between ultrasonic measures and their corresponding carcass traits indicate strong genetic correlations between the two sets of traits (Bertrand et al., 2001; Kemp et al., 2002; Greiner et al., 2003). Crews et al. (2003) found that genetic correlations between carcass traits predicted by ultrasound and measured directly ranged from 0.74 to 0.83. These results indicate that ultrasound traits could be useful indicators of carcass traits.
The collection of data on ultrasound measures of carcass traits in Angus cattle in South Africa was first implemented during 2000. The project is run and managed by the National Beef Cattle Improvement Scheme (NBCIS) of the Agricultural Research Council. Against this background, the objective of this study was to estimate genetic parameters for P8FT, RBFT, REA and IMF as well as the SCW for South African Angus cattle.
Materials and Methods
Records on ultrasound traits and live weight at scanning (SCW) from 1 122 South African Angus animals were obtained from the South African Integrated Registration and Genetic Information System (INTERGIS). The ultrasound traits included fat thickness measured on the rump (P8FT) and between the 12th and 13th ribs (RBFT), rib-eye area (REA) and intramuscular fat percentage (IMF) collected between years 2000 and 2004. Animals considered in the current study participated in the post-weaning growth tests of the NBCIS (i.e. Phases B, C, or D). Phase B is a single-herd post-weaning growth test for heifers, bulls and oxen. In Phase C young bulls from different herds undergo post-weaning growth tests in privately or publicly-owned central testing stations. Phase D is a single herd or centralised multiple-herd on-farm post-weaning growth test for young bulls. A complete description of the growth tests can be obtained from the National Beef Cattle Improvement Scheme: Guidelines. Version 1, 01 June 2007. Trained field officers of the South African Agricultural Research Council collected the ultrasound measurements using a PIE100 scanner (ARC-AII, 2003). All animals, within a group in a given herd, were scanned by the same technician. Three measurements of IMF were taken and an average was used as an official measurement of IMF. Repeated measurements on IMF were carried out to improve the accuracy, since IMF was difficult to measure accurately. However, a single measurement on other ultrasound traits was taken since they were relatively easy to measure.
Standard data editing procedures for variance component estimation were used. The criteria specifically included: (1) exclusion of extreme records (i.e. records outside five standard deviations from either side of the overall mean), (2) single-sire contemporary groups, and (3) contemporary groups without variation. Contemporary group was a concatenation of a breeder-assigned code for animals that participated in Phase B or test number for C and D animals and date at scanning. The numbers of records available for each trait after editing are presented in Table 1. Animals used in the analysis were the progeny of 137 sires and 812 dams. The average and standard deviation for age of the animals at scanning were 404.2 days and 42.1 days, respectively.
Least square means for fixed effects were conducted using SAS 9.1. (Co)variance components were estimated using a multiple-trait animal model. The fixed effects included in the model were contemporary group, sex of the animal, age of the animal at scanning fitted as a quadratic regression, and random effects of animal and residual effects. It should be acknowledged that the model considered in the current study is not optimal, since other effects such as the maternal genetic effect and the covariance between direct and maternal effects were not included in the model. Preliminary analyses fitting a more comprehensive model, including these effects led, to convergence problems (results not shown). This could be attributed to insufficient information to estimate (co)variance components for some of these effects, since the data considered in the current study was of modest size. It is also common practice to include only the direct animal genetic effect in the analyses of ultrasound traits in beef cattle (e.g. Crews et al., 2003). The multiple-trait animal model considered in the current study can be represented in matrix notation as follows:
y = Xb + Zu + e
where y is a vector of observations (y' = [y'1 y'2 y'3 y'4 y'5]), y'i is a vector of observations for traits i, b is a vector of unknown fixed effects, u is a vector of unknown random effects, and e is a vector of residuals. X and Z are the incidence matrices relating fixed and random effects to observations in y. The expected values for random effects were assumed to be null. The following covariance structures were assumed for the random effects (u and e):
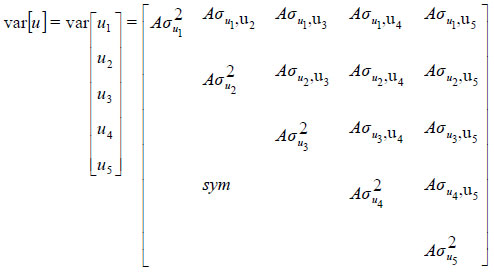

and
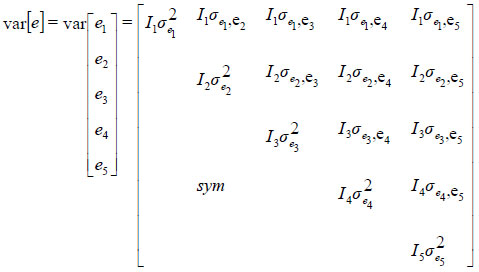
where A is the additive genetic relationship matrix and Ii is the identity matrix of the order appropriate to the number of records for the ith trait; 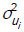 and
and 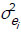 are the additive genetic and residual variances for the ith trait, respectively; σui, ui and σei, ei are the corresponding covariances for i# j.
are the additive genetic and residual variances for the ith trait, respectively; σui, ui and σei, ei are the corresponding covariances for i# j.
A three-generation pedigree (n = 6 346) was created by tracing ancestors of animals with records. The pedigree is required in the construction of the inverse of the additive relationship matrix necessary in setting up the mixed-model equations. The model presented above was used to estimate genetic parameters using the VCE package version 5.1 of Kovac & Groeneveld (2003) which employs the Restricted Maximum Likelihood (REML) procedure.
Results and Discussion
The summary statistics for all the traits are presented in Table 1. Performance on ultrasound traits is a function of many variables (e.g. sex of the animal, level of feeding and age at scanning) and therefore differences exist among estimates from different studies. The least square means for the effect of sex are presented in Table 2. The mean P8FT of 4.8 mm obtained in the current study is within the range of literature estimates. Robinson et al. (1993) reported average P8FT of 6.5 mm and 4.1 mm on heifers and bulls, respectively, on Angus and Hereford cattle at an average age of 500 d. Similarly, Reverter et al. (2000) reported average P8FT of 6.6 mm and 4.2 mm on Angus heifers and bulls, respectively. The average RBFT of 3.5 mm in this study was generally lower than values reported in other studies (Moser et al., 1998; Stelzleni et al., 2002; Crews et al., 2003).
The estimate of 62.2 cm2 for mean REA obtained in this study is consistent with the mean of 56.7 cm2 reported by Reverter et al. (2000). Crews et al. (2004) reported a mean REA of 66.1 cm2 for yearling Simmental heifers. Higher values of average REA above 70 cm2 were reported in other studies (Williams et al., 1997; Kemp et al., 2002; Crews et al., 2004). The mean IMF of 3.2% observed in this study is comparable to 2.7% and 3.4% reported by Crews et al. (2003) on yearling Simmental bulls and heifers, and also to 3.0% and 3.9% reported by Reverter et al. (2000) on yearling Angus bulls and heifers.
The estimates of variance components, heritabilities and genetic covariances are presented in Table 3. The heritability estimates for ultrasound traits obtained in the current study were moderate to high: 0.38 ± 0.08, 0.34 ± 0.06, 0.33 ± 0.05, 0.32 ± 0.06, and 0.38 ± 0.06 for SCW, P8FT, RBFT, REA and IMF respectively. The heritability estimate of 0.38 for SCW obtained in this study was in general agreement with estimates reported in the literature. Koots et al. (1994) reported unweighted and weighted mean heritability for yearling weight of 0.35 and 0.33, respectively, on various breed types while higher estimates of 0.40, 0.47 and 0.53 were reported by Moser et al. (1998), Crews et al. (2003) and Stelzleni et al. (2002) on yearling breeding bulls, Simmental steers and heifers, and Brangus cattle, respectively. A heritability estimate for SCW of 0.23 was reported by Tarouco et al. (2006) on replacement bulls and heifers of the Braford and Hereford breeds.
A heritability estimate of 0.34 obtained for P8FT in the current study falls within the range of estimates reported in the literature. Wilson et al. (1999) used a maternal grandsire model and obtained a heritability estimate for P8FT of 0.52 in the Angus breed. Reverter et al. (2000) reported heritability estimates of 0.55 and 0.31 for P8FT measures on Angus and Hereford cattle, respectively, using an animal model. Although reports on heritability estimates for ultrasound P8FT are limited, the high heritability estimate from these few studies suggests that selection for P8FT should result in appreciable genetic progress.
The heritability estimate was moderate for rib fat thickness obtained in the current study. Reverter et al. (2000) reported higher heritability estimates of 0.47 and 0.54 on Angus bulls and heifers, respectively. Likewise, Shepard et al. (1996) reported an estimate of 0.56 on Angus cattle. However, low heritability estimates for RBFT have also been reported in other studies (Johnson et al., 1993; Moser et al., 1998). For example, Johnson et al. (1993) reported an estimate of 0.14 on Brangus cattle. Moser et al. (1998) also reported a low heritability estimate of 0.11 on the Brangus breed. These authors reasoned that the low heritability estimates they obtained for RBFT were associated with the low phenotypic mean, probably because of age and level of nutrition.
There is considerable variation in the estimates of heritability for rib-eye area in the literature. Shepard et al. (1996) reported a low heritability estimate (0.11) from a small data set on the Angus breed using an animal model, while Kemp et al. (2002) and Reverter et al. (2000) reported a moderate (0.29) and a high (0.42) estimate, respectively, using data on the same breed. The estimate of heritability obtained in the current study is on the high end of the literature estimates.
The heritability estimate of 0.38 for IMF found in the current study is consistent with the average estimate of 0.41 reported by Bertrand et al. (2001) from two studies on various breed types. Crews et al. (2003) reported higher heritability estimates of 0.52 and 0.47 on Simmental heifers and bulls, respectively. Reverter et al. (2000) and Devitt & Wilton (2001) reported moderate heritability estimates (0.23 and 0.29). Stelzleni et al. (2002) reported a low heritability estimate of 0.16 from data on Brangus cattle. Reverter et al. (2000) identified two factors that are implicated in the different heritability estimates for IMF, namely (1) genetic difference between animals; this can be expressed to a greater degree if IMF mean at measurement is high and (2) the prediction of IMF from ultrasound images that may be less accurate resulting in a larger residual variance. Devitt & Wilton (2001) further stated that the differences in heritability could be explained by breed differences for genetic expression of IMF. The results in the current study contradict the observation by Stelzleni et al. (2002) that IMF was substantially influenced by other factors than additive genetic effects when obtaining a heritability estimate of 0.16.
The estimates of genetic correlations are presented in Table 3. The genetic correlation between scan weight and measures of fat was found to be low. The genetic correlation between SCW and P8FT was weak (P >0.05) and barely important for SCW and RBFT (P <0.05). These results suggest that selection for growth should have little, if any, impact on the amount of body fat.
In contrast, the genetic correlation of 0.79 between REA and SCW was higher than genetic correlations reported in other studies (Table 3). Arnold et al. (1991), Moser et al. (1998) and Tarouco et al. (2006) reported moderate genetic correlations between REA and SCW of 0.38, 0.42 and 0.40, respectively. These correlations suggest that selection for SCW would increase the size of the eye muscle area.
There was a weak (P >0.05) genetic correlation between REA and RBFT. Arnold et al. (1991) reported a similar estimate of 0.23 in Hereford cattle. Lower genetic correlations of 0.12 and 0.13 were reported in Brangus cattle by Johnson et al. (1993) and Moser et al. (1998), respectively. Stelzleni et al. (2002) reported a low but negative estimate of -0.07 in the same breed. In contrast, a higher genetic correlation (0.39) than in the current study on Angus cattle was reported by Kemp et al. (2002).
Studies on the genetic correlation between IMF and other ultrasound traits are limited and therefore only a few reports are available for comparison with the results of the current study. The genetic correlation of 0.29 between REA and IMF obtained in this study is similar to the genetic correlation of 0.25 on Brangus cattle, reported by Stelzleni et al. (2002). Bertrand et al. (2001) reported a genetic correlation of 0.07 from a summary of values on various breed types.
There was a high and positive genetic correlation among measures of fat (Table 3). The strong genetic relationship between P8FT and RBFT is consistent with results from other studies. Robinson et al. (1993) and Wilson et al. (1999) found that the two external fat traits (P8FT and RBFT) were highly correlated (0.86 and 0.82, respectively). These results suggest that the two traits are largely controlled by similar genes and that selection for one trait will lead to a concomitant increase in the other trait.
Conclusions
Estimates of heritability obtained in the current study indicate that live animal ultrasound indicators of carcass traits are moderately heritable. Selection for these traits should thus be effective. The weak genetic correlation observed between rib-eye area and measures of subcutaneous and intramuscular fat imply that attempts to increase red meat yield will be associated with little, if any, change in body fat. The strong and positive genetic correlation between measures of fat is unfavourable, since selection for reduced subcutaneous fat will be associated with a decrease in intramuscular fat. Results from the current study indicate that selection for ultrasound traits is possible but that genetic antagonism between traits should be considered in genetic improvement programmes.
Acknowledgements
This research was jointly funded by the Agricultural Research Council and the Red Meat Research and Development Trust of South Africa. Helpful comments and suggestions from two anonymous reviewers are greatly appreciated.
References
ARC-AII., 2003. Body sites for RTU measurements. In: Manual for participants in the National Beef Cattle Improvement Scheme. ARC-LNR. Livestock Business Division. Private Bag X2, Irene, 0062, South Africa. [ Links ]
Arnold, J.W., Bertrand, J.K., Benyshek, L.L. & Ludwig, C., 1991. Estimate of genetic parameters for live animal ultrasound, actual carcass data and growth traits in beef cattle. J. Anim. Sci. 69, 985-992. [ Links ]
Bertrand, J.K., Green, R.D., Herring, W.O. & Moser, D.W., 2001. Genetic evaluation for beef carcass traits. J. Anim. Sci. 79 (Suppl.), E190-E200. [ Links ]
Crews Jr, D.H., Pollak, E.J., Weaber, R.L., Quaas, R.L. & Lipsey, R.J., 2003. Genetic parameters for carcass traits and their live animal indicators in Simmental cattle. J. Anim. Sci. 81, 1427-1433. [ Links ]
Crews Jr, D.H., Pollak, E.J. & Quaas, R.L., 2004. Evaluation of Simmental carcass EPD estimated using live and carcass data. J. Anim. Sci. 82, 661-667. [ Links ]
Devitt, C.J. & Wilton, J.W., 2001. Genetic correlation estimates between ultrasound measurements on yearling bulls and carcass measurements on finished steers. J. Anim. Sci. 79, 2790-2797. [ Links ]
Greiner, S.P., Rouse, G.H., Wilson, D.E., Cundiff, L.V. & Wheeler, T.L., 2003. The relationship between ultrasound measurements and carcass fat thickness and longissimus muscle area in beef cattle. J. Anim. Sci. 81, 676-682. [ Links ]
Johnson, M.Z., Schalles, R.R., Dikeman, M.E. & Golden, B.L., 1993. Genetic parameter estimates of ultrasound-measured longissimus muscle area and 12th rib fat thickness in Brangus cattle. J. Anim. Sci. 71, 2623-2630. [ Links ]
Kemp, D.J., Herring, W.O. & Kaiser, C.J., 2002. Genetic and environmental parameters for steer ultrasound and carcass traits. J. Anim. Sci. 80, 1489-1496. [ Links ]
Koots, K.R., Gibson, J.R., Smith, C. & Wilton, J.W., 1994. Analyses of published genetic parameter estimates for beef production traits. 1. Heritability. Anim. Breed. Abstr. 62, 309-338. [ Links ]
Kovac, M. & Groeneveld, E., 2003. VCE-5 User's Guide and Reference Manual Version 5.1. Neustadt, Germany. [ Links ]
MacNeil, M.D. & Northcutt, S.L., 2008. National cattle evaluation system for combined analysis of carcass characteristics and indicator traits recorded by using ultrasound in Angus cattle. J. Anim. Sci. 86, 2518-2524. [ Links ]
Moser, D.W., Bertrand, J.K., Misztal, I., Akriese, L. & Benyshek, L.L., 1998. Genetic parameter estimates for carcass and yearling ultrasound measurement in Brangus cattle. J. Anim. Sci. 76, 2542-2548. [ Links ]
National Beef Cattle Improvement Scheme: Guidelines. Version 1, 01 June 2007. ARC-LNR. Livestock Business Division. Private Bag X2, Irene, 0062, South Africa. [ Links ]
Reverter, A., Johnston, D.J., Graser, H.-U., Wolcott, M.L. & Upton, W.H., 2000. Genetic analysis of live animal ultrasound and abattoir carcass traits in Australian Angus and Hereford cattle. J. Anim. Sci. 78, 1786-1795. [ Links ]
Robinson, D.L., Hammond, K. & McDonald, C.A., 1993. Live animal measurement of carcass traits: estimation of genetic parameters for beef cattle. J. Anim. Sci. 71, 1128-1135. [ Links ]
SAS, 2009. SAS User's Guide : Statistics, Release 9.1. SAS Institute Inc., Cary, North Carolina, USA. [ Links ]
Shepard, H.H., Green, R.D., Golden, B.L., Hamlin, K.E., Perkins, T.L. & Diles, J.B., 1996. Genetic parameter estimates of live animal ultrasonic measures of retail yield indicators in yearling breeding cattle. J. Anim. Sci. 74, 761-768. [ Links ]
Stelzleni, A.M., Perkins, T.L., Brown, A.H., Pohlman, F.W., Johnson, Z.B. & Sandelin, B.A., 2002. Genetic parameter estimates of yearling live animal ultrasonic measurements in Brangus cattle. J. Anim. Sci. 80, 3150-3153. [ Links ]
Tarouco, J.U., Silva, S.L., Ferraz, J.B.S., Balieiro, J.C.C., Roso, V.M., Mourão, G.B., Mattos, E.C., Tarouco, A.K. & Eler, J.P., 2006. Genetic analysis of live ultrasound measurements on replacement bulls and heifers of Braford. 8th World Cong. Gen. Appl. Livest. Prod., August 13-18, Belo Horizonte, MG, Brasil. [ Links ]
Williams, R.E., Bertrand, J.K., Williams, S.E. & Benyshek, L.L., 1997. Biceps femoris and rump fat as additional ultrasound measurements for predicting retail product and trimmable fat in beef carcasses. J. Anim. Sci. 75, 7-13. [ Links ]
Wilson, D.E., Rouse, G.H. & Hays, C., 1999. Carcass EPD's for yearling Angus bulls using real time ultrasound measures. Available at: http://www.exnet.iastate.edu/Pages/ansci/beefreports/bodycomp99.html. Accessed October 15, 2007. [ Links ]
Copyright resides with the authors in terms of the Creative Commons Attribution 2.5 South African Licence.
See: http://creativecommons.org/licenses/by/2.5/za
Condition of use: The user may copy, distribute, transmit and adapt the work, but must recognise the authors and the South African Journal of Animal Science
* Corresponding author: norman@arc.agric.za














