Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Animal Science
On-line version ISSN 2221-4062
Print version ISSN 0375-1589
S. Afr. j. anim. sci. vol.39 n.5 Pretoria Jan. 2009
Association of the polymorphism in the 5' flanking region of the ovine IGF-I gene with growth traits in the Baluchi sheep
Mojtaba Tahmoorespur#; Mehdi Vafaye Valeh; Mohammad Reza Nassiry; Alireza Heravi Moussavi; Maziar Ansary
Department of Animal Science, Faculty of Agriculture, Ferdowsi University of Mashhad, P.O. Box 91775-1163, Mashhad, I.R. Iran
ABSTRACT
The insulin-like growth factor 1 (IGF-I) gene has been described in several studies as a candidate gene for growth traits in farm animals. The present preliminary study attempts to establish associations between growth traits and genetic polymorphisms at the 5' flanking region s IGF-I in the Baluchi sheep. The DNA of 102 sheep of the indigenous Iranian Baluchi breed was evaluated. Polymerase chain reaction-single strand conformation polymorphism (PCR-SSCP) analysis of the 5' flanking region (Exon1) of the ovine IGF-I gene revealed three banding patterns (genotypes) named as A/A, A/B and B/B. The evaluation of an association effect between these SSCP patterns with birth weight (BW), weaning weight (WW), and average daily gain from birth to weaning (GBW), weaning to six month (GWS) and from six month to yearling age (GSY) suggest a positive effect of pattern (Genotype) A/B with GBW and weaning weight (WW). Pattern (Genotype) B/B had a superior birth weight when compared to those of individuals with other patterns. Moreover, the A/A pattern (Genotype) appeared favourable for live weight at yearling age. These results confirmed the potential usefulness of this gene in marker-assisted selection programmes for sheep breeding.
Keywords: Baluchi sheep, Insulin-like growth factor I, growth, Single strand conformation polymorphism
Introduction
In Iran lamb and mutton are traditional sources of protein and the consumption levels are high in comparison with meat from cattle and goats. Due to the fact that the supply of meat from sheep does not cover the demand, a breeding scheme is needed to increase the efficiency of sheep production by improving litter size, body conformation, lamb weight and milk yield (Yazdi et al., 1997). Profitability of sheep production for meat depends to a great extent on lamb weight, so the selection objective should concentrate on these traits (Tosh & Kemp, 1994). Quantitative traits are often controlled by a great number of genes. Localization of QTL's (Quantitative Trait Loci) can be done by linkage disequilibrium analysis or by candidate gene approach. Candidate genes have known biological functions related to the development or physiology of an important trait (Rothschild et al., 1997). Insulin-like growth factors one and two (somatomedins- IGF-I and IGF-2) are structurally related proteins, playing a key role in cell differentiation, embryogenesis, growth and regulation of metabolism (Siadkowska et al., 2006). Due to their role in regulation of cell proliferation and animal growth, the IGF-I and its gene are considered as candidate markers for growth rate and meat production traits in mammalia. In spite of the functional importance of IGF-I in the regulation of GH, no reports have been published on the effects of the IGF-I gene polymorphism on growth traits in sheep breeds. The objectives of this study were to search for the same polymorphism in the 5'flanking region of IGF-I in Iranian Baluchi sheep that was found by Yilmaz et al. (2005) in purebred Polypays and crossbreds consisting of the Hampshire, Targhee, Rambioullet, Dorset and Suffolk breeds using a non-radioactive SSCP protocol and to investigate the relationship between these polymorphisms and growth traits of Baluchi sheep. This is intended to be a first step of a more in-depth study of the "Baluchi" breed in order to establish a breeding programme based on marker-assisted selection.
Material and Methods
Blood samples were randomly collected from 102 purebred Baluchi sheep, progeny of 18 rams, at the Baluchi Breeding Station located north-east of Mashhad, Iran. The data set and pedigree information used in this research were pre- and post-weaning body weights, birth weight (BW), weaning weight (WW), 6-month weight (6MW) and yearling weight (YW)) collected from 2000 to 2003 from the sheep at the Breeding Station of Baluchi. Lambs were weighed and ear-tagged at birth time. Weaning was ca. three months of age. All lambs were weaned at the same day, though necessarily at the same age. Moreover, lambs were weighed monthly on the same day. Blood samples (10 mL) were obtained by jugular venipuncture, using vacuum tubes treated with 0.25% ethylenediaminetetracetic acid (EDTA). DNA was extracted from 100 µL of blood as described by Boom et al. (1990). After measuring the DNA concentration and its purity by spectrophotometry, DNA was diluted to a final concentration of 50 ng/µL in water and stored at 4 ºC. The 265-bp fragment of the 5' flanking region of the ovine IGF-I gene was amplified. Based on the ovine IGF-I gene sequence (GenBank accession no. AY803775), one pair of oligonucleotide primers were designed (Table 1) to amplify this fragment using the Primer Premier 5.0 Software (www.primerbiosoft.com).
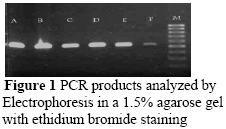
To obtain the 265 bp fragment of IGF-I exon1 the following PCR mix was used: Each tube contained 1.5 units of Taq DNA polymerase, 10 mM of Tris-HCl (pH 9), 50 mM of KCl, 1.5 mM of MgCl2 and 200 mM of each dNTP. To this mixture, 1 µL of each primer (50 ng/µL), 22 µL of water, 1 µL of DNA (50 ng/µL) and two drops of mineral oil were added. Amplification was carried out for IGF-I in a Biometra T-Personal Ver: 1.11 thermocycler. Following a hot start (95 ºC for 3 min), 35 cycles were carried out (95 ºC for 45 seconds, 62 ºC for 40 seconds, 72 ºC for 50 seconds), ending with a 10 min final extension at 72 ºC. Amplification was verified by electrophoresis on 1.5% (w/v) agarose gel in 1 x TBE buffer (2 mM of EDTA, 90 mM of Tris-Borate, pH 8.3), using a 50bp ladder as a molecular weight marker for confirmation of the length of the PCR products. Gels were stained with ethidium bromide (1 µg/mL).
For single-strand conformation polymorphism (SSCP) analysis, several factors were tested to optimize the methodology: Amount of PCR product (4 - 15 µL), dilution in denaturing solution (20 - 85%), denaturing solution (A: 95% of formamide, 10 mM NaOH, 0.05% xylene-cyanol and 0.05% bromophenol blue; B: same as A, plus 20 mM of EDTA), acrylamide concentration (6 - 14%), percentage of cross linking (1.5 to 5%), presence (10%) or absence of glycerol, voltage (100 - 350 V), running time (2-12 h) and running temperature (4, 6, 10 and 15 ºC). Each PCR reaction was diluted in denaturing solution, denatured at 95 ºC for 5 min, chilled on ice and resolved on non-denaturing polyacrylamide gel. The electrophoresis was carried in a vertical unit (Paya pajoohesh VEU-7350, 160×140×0.75 mm), in 1× TBE buffer. The gels were stained with silver.
Prior to statistical analyses, birth weight (BW), weaning weight (WW), 6-month weight (6MW), and yearling weight (YW) traits were adjusted for age of lamb. Average daily gain (ADG) was calculated as the difference between the initial and final body weights divided by the number of days of the test. The relationship structure of the sample was taken into account by including sire and dam-within-sire effects in the models. The effects of genotypes on the traits studied were analyzed by the least-squares method as applied in the general linear model (GLM) procedure of JMP 7.0 (SAS, 2007) according to the following statistical model:
Yijklmn = µ + Gi + Sj + b1(Bijklmn- 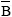 ) + Srk + D1(Srk) + Ym + eijklmn
) + Srk + D1(Srk) + Ym + eijklmn
Where: Yijklmn = phenotypic value of trait, µ = the overall mean, Gi = the fixed effect of IGF-I genotypes (i = 1..., 3), Sj = the fixed effect of sex (j = 1, 2), b1 = the linear regression coefficient of trait on birth weight, Srk = fixed effect of sire, D1(Srk) = dam nested within sire as random effect, Ym = the fixed effect of year; eijklmn= the random residual error.
Least squares means of the genotypes were compared by the Tukey test. Significance was determined at P <0.05. The non-significant effects and corresponding interactions were discarded for the final analysis. Of the 102 animals, four were excluded of the statistical approach due to missing values or unreliable data.
Results
A total of 102 sheep were genotyped for polymorphism in the IGF-I gene. The PCR amplified a 265bp fragment from exon1 of the ovine IGF-I gene (Figure 1). After optimization of the parameters that affect the detection of SSCP's, we analyzed the PCR products from 102 animals, with the conditions described in Table 2. The PCR-SSCP analysis of exon1 IGF-I revealed three distinct patterns (Figure 2). Yilmaz et al. (2005) found the same patterns that corresponded with three genotypes of A/A, A/B, and B/B in mixed breed sheep. The frequencies were 45 for pattern (A/A), 48 for pattern (A/B) and nine for pattern (B/B) (Table 3).
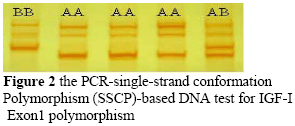

In the tested Baluchi sheep population, significant statistical results were found in average daily gain from birth to weaning (GBW) between individuals with genotype AB and BB (P <0.05). Individuals with genotype AB of IGF-I gene had a superior average daily gain from birth to weaning (GBW) when compared to those of individuals with other genotype (P <0.05). However, no significant association of different genotypes with other traits was detected (P >0.05) (Table 3).
Discussion
A common assumption in animal breeding is that variation in quantitative traits such as growth or lactation is controlled by many genes. Usually, each of these genes has a small effect. However, the major gene model suggests that few genes may account for a relatively large proportion of genetic variation. Genes involved in the biology of a trait of interest are candidates for association studies, and can be considered as "candidate genes". The SSCP analysis of genes, whose product is associated with production traits, could be a valuable alternative approach for the establishment of the allelic variants useful as markers to aid selection. Up till now no papers have been found in the literature concerning effects of IGF-I polymorphism on growth traits in sheep. In the present study associations were searched between a SNP (single nucleotide polymorphism), in the 5' flanking region of the ovine IGF-I gene and growth traits in Baluchi sheep. The polymorphism in question was first reported in Angus cattle by Ge et al. (1997) as SSCP. This polymorphism was then identified as T/C transition, also recognizable as RFLP-SnaBI (Ge et al., 2001). For a fragment of 265bp in the 5' flanking region of the ovine IGF-I gene, 467 to 732bp upstream from the 5' end of Exon1, we observed three conformational patterns. Yilmaz et al. (2005) found the same patterns that corresponded with three genotypes of A/A, A/B, and B/B in mixed breed sheep. Also, the present report shows an association of the single nucleotide polymorphism in the 5'-flanking region of the IGF-I gene with growth traits in Baluchi sheep. The results of the GLM analysis of association between the IGF-I gene and growth traits in Baluchi sheep are summarized in Table 3. The effect of the IGF-I gene was significant (P <0.05) only for average daily gain from birth to weaning (GBW). There was a significant difference between A/B and B/B genotypes. No other association between genotypes and traits studied was found. In addition the results demonstrate the superiority of the heterozygous A/B genotype for weaning weight, and of genotype B/B for birth weight and A/A genotype for yearling weight. Ge et al. (2001) reported an effect of the IGF-I/SnaBI polymorphism, located in the regulatory region of the IGF-I gene, on growth traits in Angus cattle, also suggested a direct action of this polymorphism on gene transcription and, consequently, on phenotypic traits. Moody et al. (1996) observed an effect of STR polymorphism on body weight at birth and at weaning, and with the growth rate In Hereford cattle, Siadkowska et al. (2006) reported a correlation between the polymorphism in the 5'-non coding region of the IGF-I and meat and milk production traits in Polish HF cattle. Finally for the reason that quantitative traits are regulated by many genes and affected by interactions among them, and thus, a candidate gene associated with a trait in one population may have a different effect, or show no effect at all, in another population due to negative effects of other genes and epistatic interactions of the candidate gene with other genes in the population (Pomp, 1994).
Conclusions
SSCP polymorphisms can be rapidly and inexpensively detected in a population of farm animals and their associations with growth traits studied. We have applied this approach to look for associations between polymorphisms observed in exons1 of the IGF-I gene from the autochthonous Iranian ovine Baluchi breed and growth traits. Average daily gain from birth to weaning (GBW) appears to be under the influence of polymorphisms detected in exons1. The positive influence of exon1 in average daily gain from birth to weaning (GBW) established in this study confirms that this region of the ovine IGF-I gene is particularly important and deserves further studies. Also, our data showed that these polymorphisms contributed to the variation in the traits analyzed and reinforce the possibility of using these polymorphisms in molecular marker-assisted selection and breeding programmes. Additional research with a bigger population of the same breed in order to have more animals of each genotype is required. Moreover, a study with other commercial breeds is also needed to determine the general effects of these polymorphisms.
Acknowledgements
This research was supported by the Centre of Excellence in Animal Science, Agricultural College, Ferdowsi University of Mashhad.
References
Boom, R., Sol, C.J.A. & Salimans, M., 1990. Rapid and simple method for purification of nucleic acids. J. Clin. Microbial. 28, 495-503. [ Links ]
Ge, W., Davis, M.E. & Hines, H.C., 1997. Two SSCP alleles detected in the 5'-flanking region of bovine IGF-I gene. J. Anim. Genet. 28, 155-156. [ Links ]
Ge, W., Davis, M.E., Hines, H.C., Irvin, K.M. & Simmen, R.C., 2001. Association of a genetic marker with blood serum insulin-like growth factor-I concentration and growth traits in Angus cattle. J. Anim. Sci. 79, 1757-1762. [ Links ]
Moody, D.E., Pomp, D., Newman, S. & MacNeil, M.D., 1996. Characterization of DNA polymorphisms in three populations of Hereford cattle and their associations with growth and maternal EPD in line 1 Herefords. J. Anim. Sci. 74, 1784-1793. [ Links ]
Pomp, D., 1994. Biotechnology and beef cattle improvement: Myths and realities. In: Proc. Beef Improv. Fed; 26th Res. Symp. Ann. Mtg: Des Moines, IA. pp. 236-241. [ Links ]
Rothschild, M.F., Messer, L.A. & Vincent, A., 1997. Molecular approaches to improved pig fertility. J. Rep. Fertil. 52, 227-236. [ Links ]
Siadkowska, E., Zwierzchowski, L., Oprządek, J., Strzałkowska, N., Bagnicka, E. & Krzyżewski, J., 2006. Effect of polymorphism in IGF-I gene on production traits in Polish Holstein-Friesian cattle. J. Anim. Sci. 24, 225-237. [ Links ]
Tosh, J.J. & Kemp, R.A., 1994. Estimation of variance components for lamb weights in three sheep populations. J. Anim. Sci. 72, 1184-1190. [ Links ]
Yazdi, M.H., Engstrom, G., Nasholm, A., Johansson, K., Jorjani, H., & Liljedahl, L.E., 1997. Genetic parameters for lamb weight at different ages and wool production in Baluchi sheep. J. Anim. Sci. 65, 247-255. [ Links ]
Yilmaz, A., Michael, E., Harold, C.H., Hines & Hoyoung Chung., 2005. Detection of two nucleotide substitutions and putative promoters in the 5' flanking region of the ovine IGF-I gene. J. Appl. Genet. 46, 307-309. [ Links ]
# Corresponding author. E-mail: me_va84@yahoo.com














