Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Animal Science
On-line version ISSN 2221-4062
Print version ISSN 0375-1589
S. Afr. j. anim. sci. vol.39 n.4 Pretoria Jan. 2009
Genetic parameter estimates for tick resistance in Bonsmara cattle
M.A. BudeliI,III,IV; K.A. NephaweII; D. NorrisIV; N.W. SelapaIII; L. BerghIII; A. MaiwasheIII,#
ILimpopo Department of Agriculture, Private Bag X2247, Sibasa, 0970, South Africa
IILimpopo Department of Agriculture, Private Bag X9487, Polokwane, 0700, South Africa
IIIAgricultural Research Council, Animal Production Institute, Private Bag X2, Irene, 0062, South Africa
IVUniversity of Limpopo, Private Bag X1106, Sovenga, 0727, South Africa
ABSTRACT
The objectives of the study were to estimate genetic parameters for tick resistance and to evaluate the effect of the level of tick infestation on the estimates of genetic parameters for South African Bonsmara cattle. Field data of repeated tick count records (n = 11 280) on 1 176 animals were collected between 1993 and 2005 by 10 breeders participating in the National Beef Recording and Improvement Scheme. The distribution of tick count records were normalized using a Box-Cox transformation. Data were divided into seven sub-data sets based on the mean tick count per contemporary group, to facilitate the investigation of the effect of level of tick infestation on the derived genetic parameters. A repeatability animal model including the fixed effects of contemporary group and age of animal at tick counting and random effects of the direct additive genetic, permanent environmental and residual effects was used to estimate genetic parameters using REML procedures. The additive genetic variance for tick count ranged from 0.01 to 0.08. The animal permanent environmental variance ranged from 0.00 to 0.03. Phenotypic variance decreased with increasing mean tick count level while additive genetic variance increased with mean tick count level. The heritability also increased with mean tick count level until a mean tick count level of >30. The highest heritability estimate obtained in the current study was 0.17 for data with mean tick count level >25. These results suggest that sufficient genetic variation for tick count exists in the Bonsmara cattle. Genetic selection for tick resistance is feasible even though genetic progress may be slow.
Keywords: tick count, variance components, heritability estimates, beef cattle, external parasites
Introduction
Tick-borne diseases are major constraints to livestock production throughout the tropics and sub-tropics. Losses in livestock production due to external parasites have long been a major concern to livestock producers in the tropics and subtropics (Seifert, 1984a; b). One million cattle are estimated to have died of East Coast fever in sub-Saharan Africa during 1989 alone (Mukhebi et al., 1992). In South Africa, tick-borne diseases have been estimated to cost the livestock industry about R 70 to R 200 million per year (Bigalke, 1980).
Various tick-borne disease control methods have been employed in South Africa (Bigalke et al., 1976; De Vos, 1979; Purnell & Schroder, 1984). Historically, the earliest tick control trials with dipping agents in South Africa started in 1889. These trials were prompted by the discovery in the USA in 1893 that ticks transmit the causal piroplasm of redwater in cattle (Hayward, 1981). Increase in acaricides prices and drug resistance pose challenges in the application of these methods because the application processes are associated with an increase in input costs, while it takes time to develop new drugs.
Hayward (1981) stated that the best solution to high acaricides prices and resistance to drugs might be the identification of naturally resistant breeds and the encouragement of their use. Natural disease resistance refers to the inherent capacity of an animal to resist disease when exposed to pathogens, without prior exposure or immunization (Adams & Templeton, 1998). Utech et al. (1978) defined tick resistance as the ability of cattle to limit the number of ticks that survive to maturity.
Although some of the observed variation in natural tick resistance is related to environmental factors, a significant component of variation in natural disease resistance appears to be of genetic origin (Adams & Templeton, 1998). Several studies have been conducted on genetic determination of tick resistance (Utech et al., 1978; Spickett et al., 1989; Rechav et al., 1990). Tick resistance has been shown to be heritable (Hewetson, 1972). Davis (1993) reported a heritability estimate of 34% for tick resistance, indicating that genetic improvement through selection should be effective. Information on resistance status within the various breeds of cattle is needed to provide a basis for selection, by either breeding from animals with resistance, or culling cattle with low tick resistance, or both.
The South African National Beef Recording and Improvement Scheme of the Agricultural Research Council initiated a tick count data collection pilot project in 1993 in conjunction with the South African Bonsmara Cattle Breeders Society. The short-term objective of the project was to collect data that could be used as a management tool (i.e. to determine when to dip the animals) and for phenotypic selection. The long-term objective was to collect data that will ultimately form the basis for the development and implementation of a genetic improvement programme for tick resistance. The primary objective of the current study was therefore to assess the level of genetic variation for tick resistance in South African Bonsmara cattle by estimating genetic parameters for tick count. The secondary objective was to evaluate the effect of the level of tick infestation on the genetic parameter estimates for tick count.
Materials and Methods
Data used in the current study were obtained from the National Beef Recording and Improvement Scheme (NBRIS). The data included tick count records on Bonsmara cattle from 10 stud herds that participated in the tick count data collection pilot project. The breeders participating in the project were located in Limpopo, North-West and Western Cape provinces of South Africa and their participation in the project ranged from one to nine years (1993 to 2005). Participants in the project (i.e. the breeders) were responsible for collection of tick count data following the guidelines of the NBRIS (2008). Briefly, the guidelines state that animals should not be dipped during the testing period, in case where dipping is necessary tick count records must be collected prior to dipping. A minimum period of three weeks must be allowed between two dates of tick counting. Frisch & O'Neill (1998) and Burns et al. (1997) used the same period between counts in their studies on tick resistance. The guidelines further state that the same person should record tick count in a given herd and date. It is also important that the person recording tick counts must be experienced or under the supervision of an experienced person. In the study by Seifert (1971) it was reported that errors due to observer were shown to be heterogeneous and were greater when a temporary engaged novice made duplicate counts than when they were made by an experienced person.
The following information was recorded at tick counting: animal identification, tick count and the sex of animal. In some herds tick counts were taken throughout the year while others concentrated on certain months (i.e. when ticks were prevalent). The original data set consisted of 11 280 repeated measurements of tick counts on 1 176 animals. Other information necessary for genetic parameter estimation such as national animal identification, date of birth and pedigree information were obtained from the Integrated Registration and Genetic Information System (INTERGIS) database.
Data were edited to exclude tick count records (1) on animals younger than 250 days, and (2) extreme tick count records, i.e. more than five standard deviations above the mean. Furthermore, contemporary groups (the concatenation of herd, sex and year, month and day of tick counts) with less than five animals were excluded. Preliminary analysis of the data showed that variation in tick counts among animals in a contemporary group depended on the level of tick infestation. That is, the coefficient of variation in tick count increased with the mean tick count in a contemporary group. In the study by Burrow (2001) a minimum of 15 ticks per side was required for the records to be considered useful. In the current study, to investigate the effect of level of tick infestation (i.e. mean tick count in a contemporary group) on genetic parameter estimates, a total of seven data sets were created. The seven data sets included records from contemporary groups with mean tick count > 5 (Data 1), > 10 (Data 2), > 15 (Data 3), > 20 (Data 4), > 25 (Data 5), > 30 (Data 6), and > 35 (Data 7). The number of contemporary groups per data set ranged from 36 to 210. Visual inspection of the frequency distributions revealed that tick count had a non-Gaussian or non-normal distribution. Thus, the data were normalized using a Box-Cox family of power transformations (Box & Cox, 1964). The skewness for data sets 1 to 7 before (after) transformation was as follows: 2.689 (-0.071), 2.333 (-0.193), 1.844 (-0.283), 1.565 (-0.364), 1.287 (-0.670), 1.130 (-0.919) and 0.846 (-1.209). The skewness of 0 indicates that the distribution is symmetric. In general, the transformation enhanced the normality of the data except for data set 7. The Box-Cox transformation is given by y'=(yk - 1)/k for k ≠ 0 or y' = log(y) for k = 0; where y and y' are the raw and transformed tick count data, respectively. The maximum likelihood estimate of the parameter k was obtained using an algorithm proposed by Hyde (1999). Silva et al. (2006) and Gasparin et al. (2006) also used a Box-Cox transformation in the analysis of faecal egg count in Angus cattle and tick count in an experimental population, respectively. Handlesman (2002) pointed out that a power transformation performs better than logarithmic transformation in normalising sperm concentration data. It should be noted that data transformation does not guarantee that the transformed data are normal. However, transformation may improve properties of estimates, predictions and inferences (Sonstegard et al., 2006). All results are presented on the transformed scale and all references to tick counts or scores imply transformed counts unless specific reference is made to raw data.
A repeatability animal model was used to analyse the data. The model included the fixed effects of the contemporary group and age of the animal at tick count data collection and random additive genetic, permanent environment and residual effects. The matrix representation of the model equation is as follows:
y = Xb + Za + Wc + e
where y is a vector of normalised tick count observations, b is a vector of fixed effects of contemporary group and age (fitted as a quadratic regression), a is a vector of random direct additive genetic effects of the animals, c is a vector of random permanent environmental effects of the animals, e is a vector of random residual effects unique to each observation and X, Z and W are known incidence matrices relating the fixed and random effects, respectively, to observations in y. The random effects were assumed to be normally distributed with the following first and second moments:
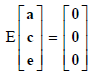
and
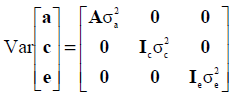
where A is the numerator relationship matrix, Ic is an identity matrix of the order equal to the number of animals with records, Ie is an identity matrix of the order equal to the number of records. The variance components 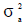 ,
, 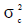 and
and 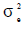 are the direct additive genetic, permanent environment and residual variances, respectively.
are the direct additive genetic, permanent environment and residual variances, respectively.
Variance components and their corresponding ratios to the phenotypic variance were estimated using the Variance Components Estimation version 6.0.2 package (VCE6) of Groeneveld et al. (2008). The VCE obtains estimates of variance components by the REML procedures using analytical gradients and Gibbs sampling. The method of analytical gradient was used in the current study. The software package for multivariate Prediction and Estimation (PEST Version 4.2) of Groeneveld et al. (1990) was used to format the data for VCE6. A three-generation pedigree was built around the data for each of the seven data sets considered. The pedigrees ranged from 1 583 to 2 747 animals in different data sets.
Results and Discussion
A comprehensive summary statistics of the final data sets is presented in Table 1. The smallest data set included repeated records of 514 animals while the largest data set included records of 1 137 animals. The number of sires and dams in the different data sets ranged from 75 to 130 and 364 to 754, respectively. The minimum and maximum number of herds in the different data sets was six and 10, respectively. The variation in the age of the animals at tick count data collection in the current study was considerable (i.e. from 250 to 5 521 days). The average age of the animals ranged from 864 to 964 days in the different data sets. Corbet et al. (2006) analysed tick count data from animals that had a narrower age range than considered in the current study (i.e. from 360 to 2 920 days). Burrow (2001) estimated genetic parameters for tick count using monthly records collected from weaning (at six months) to 18 months of age.
The average untransformed tick count ranged from 18.6 to 58.7 across the different data sets (Table 1). The level of tick infestation in the current study was comparable to what has been found in other studies. Corbet et al. (2006) found a mean of 37 ticks from 622 animals with tick count ranging from 1 to 150. Turner & Short (1972) compared tick infestation of different breeds and the mean tick count per side for the Afrikaner and Brahman breeds on natural infestation was 20 - 30 ticks whereas the Shorthorn breed carried 75 - 100 ticks per side. In the study by Regitano et al. (2006) the mean tick count after artificial infestation was 21.52.
The estimates of variance components and corresponding ratios for transformed tick count are presented in Table 2. The additive genetic variance increased with an increase in the mean tick count per contemporary group to a maximum at a mean tick count >30 after which a sharp decline was observed. The lowest estimate of genetic variance was observed for data with a mean tick count of >35. This decline in the additive genetic variance for Data set 7 (or mean tick count of >35) could be ascribed to the low number of animals in the data set. Variances for permanent environment decreased with an increase in mean tick count. This was expected since the number of records per animal decreased with an increase in the mean tick count per contemporary group (Table 1). Thus, the amount of information available to estimate the permanent environmental effect was limited in data sets with a high mean tick count per contemporary group. Phenotypic and residual variances decreased with an increase in mean level of tick count per contemporary group.
The heritability estimates for the different data sets are presented in Table 2. These estimates increased with the mean tick count per contemporary group and stabilised when the mean tick count per contemporary group was >25 and <31. The estimates ranged from 0.05 to 0.17 for data with mean tick counts of >5 and >30, respectively. The low estimate of heritability for data with a mean tick count of >35 corresponded with its lowest genetic variance. These results also indicate that as more cattle are infested with ticks, increased levels of genetic variation between cattle would be observed. Heritability estimates from this study are lower compared to results from other similar studies. A study on Belmont Red cattle in Australia reported a heritability estimate of a log-transformed tick count to be 0.42 (Burrow, 2001). Seifert (1971) evaluated variations in resistance of cattle tick between and within breeds of cattle and reported a heritability of 0.48 for the Shorthorn x Hereford line and an estimate of 0.82 for Zebu crossbreds. Davis (1993) reported a heritability estimate of 0.34 for log-transformed tick counts. Wharton et al. (1970) reported heritability estimates of 0.39 for dam-calf correlations and 0.49 for full sib correlations. In the same study heritability was further estimated based on winter and summer seasons. The heritability estimates for dam-calf were 0.42 and 0.07 whereas for full sibs were 0.64 and -0.17 for summer and winter, respectively. Separating data according to season resulted in increased heritability in summer and a low to zero heritability in winter. It was suggested that this might be due to either seasonal change in the intensity of expression of component of host resistance or a seasonal change in the sensitivity of ticks to some mechanism of host resistance. It is important to note that, at the time of the year when discrimination is poor, it is not a matter of the same variation in inherent susceptibility being obscured by some extra error of variation: the total variation (animal+error) is reduced at this time and inherent differences in resistance produce small effects. No suggestion was offered as to the cause of the seasonal effect on discrimination. It is therefore important that the existence of the effect should be defined under any circumstances where research into resistance or selection for resistance may be undertaken.
Wharton et al. (1970) suggested that the high heritability estimates from full-sib correlations compared to those from dam-calf correlations may be due to inflation by maternal effects. Apart from the possibility of short-lived transfer of passive immunity, the mechanism of maternal effects is not obvious.
Henshall (2004) reported heritability estimates of 0.41 for transformed data and emphasized that tick count should be recorded at the time when animals have had sufficient exposure to ticks to ensure that resistance has been acquired. In the study by Hewetson (1968) heritability estimates from five levels of artificial infestation of sires increased from 0.28 to 0.42 between the fourth and fifth infestation, whereas there was zero heritability at the first infestation. Hewetson (1968) emphasised the importance of acquired resistance in genetic analyses of tick counts.
The animal permanent environment variance ratios were lower for all ranges of mean tick counts considered in the current study, thus indicating that permanent environment had little or no effect on variation of tick counts. It was mentionable only in data set 4 (i.e. a mean tick count of >20) where it contributed only 6% of the total variation. In the study by Burrow (2001) the animal permanent environment accounted for 18% of the phenotypic variation, which was higher than in the present study. The residual variance ratios were higher for data sets considered in the current study indicating that there is still a lot of unknown phenotypic variation that was not accounted for by the effects in the model. The high proportion of residual variation observed in the current study could be due to differences in the data collection process among the different participating herds. Standardisation of the tick count data collection process should be considered in the National Beef Recording and Improvement Scheme to obtain data more suitable to estimating the heritability of tick counts.
Conclusions
The results from the current study indicate that sufficient levels of genetic variation for tick count exist in the South African Bonsmara population. Thus, selection for tick resistance using estimated breeding values for tick counts is a viable option even though genetic progress may be slow. Special attention should be given to the data collection process to enhance the quality of the data. It is recommended that records from contemporary groups with a mean tick count of at least 20 ticks per animal should be considered for genetic evaluation purposes. It is therefore important that under natural infestation tick count recording be done at the time when the tick population is high.
Acknowledgements
This research was jointly funded by the National Research Foundation Technology and Human Resources for Industry Programme (THRIP), the Red Meat Research and Development Trust of South Africa and the Limpopo Department of Agriculture.
References
Adams, L.G. & Templeton, J.W., 1998. Genetic resistance to bacterial diseases of animals. Rev. Sci. Technol. 17, 200-219. [ Links ]
Bigalke, R.D., 1980. The control of ticks and tick-bone diseases of cattle in South Africa. Zim. Vet. J. 11, 20-21. [ Links ]
Bigalke, R.D., De Vos, A.J. & Barrowman, P.R., 1976. The control of some tick-borne disease in S.A. Bullet. de l' off. Intl. des Epiz. 86, 89-100. [ Links ]
Box, G.E.P. & Cox, D.R., 1964. An analysis of transformations. J. R. Statist. Soc., B. 26, 211-246. [ Links ]
Burns, B.M., Reid, D.J. & Taylor, J.F., 1997. An evaluation of growth and adaptive traits of different cattle genotypes in a subtropical environment. Aust. J. Exp. Agric. 37, 399-405. [ Links ]
Burrow, H.M., 2001. Variances and covariances between productive and adaptive traits and temperament in a composite breed of tropical beef cattle. Livest. Prod. Sci. 70, 213-233. [ Links ]
Corbet, N.J., Shephard, R.K., Burrow, H.M., Van der Westhuizen, J., Strydom, P.E. & Bosman, D.J., 2006. Evaluation of Bonsmara and Belmont Red cattle breeds in South Africa. 1. Productive performance. Aust. J. Agric. Res. 46, 199-212. [ Links ]
Davis, P., 1993. Genetic parameters for tropical beef cattle in northern Australia: A Review . Aust. J. Agric. Res. 44, 179-198. [ Links ]
De Vos, A.J., 1979. Epidemiology of bovine babesiosis in South Africa. Onderstepoort J. Vet. Res. 50, 357-362. [ Links ]
Frisch, J.E. & O'Neill, C.J., 1998. Comparative evaluation of beef cattle breeds of African, European and Indian origins. 2. Resistance to cattle ticks and gastrointestinal nematodes. Anim. Sci. 67, 39-48. [ Links ]
Gasparin, G., Miyata, M., Coutinho, L.L., Martinez, M.L., Machado, M.A., Silva, M.V.G.B., Campos, A.L., Sonstegard, T.S. & Regitano, L.C.A., 2006. Detection of a QTL associated with tick resistance on bovine chromosome 7 (BTA7) using a F2 experimental population. Proc. 8th Wrld Congr. Genet. Appl. Livest. Prod. August 13-18, Belo Horizonte, MG, Brasil. [ Links ]
Groeneveld, E., Kovac, M. & Wang, T., 1990. PEST software for multivariate Prediction and Estimation, Version 4.2. Department of Animal Sciences, University of Illinois, USA. [ Links ]
Groeneveld, E., Kovac, M. & Mielenz, N., 2008. VCE User's Guide and Reference Manual Version 6.0. Institute of Farm Animal Genetics, Mariensee, Germany. [ Links ]
Handlesman, D.J., 2002. Optimal power transformations for analysis of sperm concentration and other semen variables. J. Andrology 23, 629-634. [ Links ]
Hayward, S., 1981. Opening address. In: Tick Biology and Control. Eds Whitehead, G.B. & Gibson, J.D., Tick Research Unit, Rhodes University, Grahamstown South Africa. pp. xiii -xiv. [ Links ]
Henshall, J.M., 2004. A genetic analysis of parasites resistance traits in a tropically adapted line of Bos taurus. Aust. J. Agric. Res. 55, 1109-1116. [ Links ]
Hewetson, R.W., 1968. Resistance of cattle to tick, Boophilus microplus. II. The inheritance of resistance to experimental infestations. Aust. J. Agric. Res. 19, 497-505. [ Links ]
Hewetson, R.W., 1972. The inheritance of resistance by cattle to cattle tick. Aust. Vet. J. 48, 299-303. [ Links ]
Hyde, S., 1999. Likelihood based inference on the Box-Cox family of transformations: SAS and MATLAB programs. MSc thesis, Montana State University, USA. [ Links ]
Mukhebi, A.W., Perry, B.D. & Kruska, R., 1992. Estimated cost of theileriosis control in Africa. Preventative Vet. Med. 12, 73-85. [ Links ]
NBRIS, 2008. National Beef Recording and Improvement Scheme Guidelines, Version 2008.3 ARC-Animal Production Institute, Irene, Pretoria, Republic of South Africa. [ Links ]
Purnell, R.E. & Schroder, J., 1984. Herd prophylaxis of tick-borne diseases in susceptible cattle in South Africa. Proc. XIIIth Wrld Congr. on Diseases of Cattle, 17-24 September, Durban. pp. 488-493. [ Links ]
Rechav, Y., Dauth, J. & Els, D.A., 1990. Resistance of Brahman and Simmentaler cattle to southern African ticks. Onderstepoort J. Vet. Res. 57, 7-12. [ Links ]
Regitano, L.C.A., Martinez, M.L. & Machado, M.A., 2006. Molecular aspects of bovine tropical adaptation. Proc. 8th Wrld Congr. Genet. Appl. Livest. Prod. August 13-18, Belo Horizonte, MG, Brasil . [ Links ]
Seifert, G.W., 1971. Variations between and within breeds of cattle in resistance to field infestations on the cattle tick (Boophilus microplus). Aust. J. Agric. Res. 22, 159-168. [ Links ]
Seifert, G.W., 1984a. Research and practical experience in selection for resistance to the cattle tick (Boophilus microplus) and gastrointestinal helminths in northern Australian beef cattle. Proc. 2nd Wrld Congr. Sheep and Beef Cattle Breeding. Pretoria, South Africa. pp. 149-159. [ Links ]
Seifert, G.W., 1984b. Selection of beef cattle in Northern Australia for resistance to the cattle tick (Boophilus microplus): Research and application. Preventative Vet. Med. 2, 553-558. [ Links ]
Silva, M.V.G.B., Van Tassell, C.P., Sonstegard, T.S., Cobuci, J.A. & Gasbarre, L., 2006. Estimates of genetic parameters using random regression models for gastrointestinal Helminthoid data in Angus cattle. Proc. 8th Wrld Congr. Genet. Appl. Livest. Prod. August 13-18, Belo Horizonte, MG, Brasil. [ Links ]
Sonstegard, T.S., Silva, M.V.G.B., Gasbarre, L.C. & Van Tassell, C.P., 2006. Use of Box-Cox transformation in analysis of fecal egg count data. Proc. 8th Wrld Congr. Genet. Appl. Livest. Prod. August 13-18, Belo Horizonte, MG, Brasil. [ Links ]
Spickett, A.M., De Klerk, D., Enslin, C.B. & Scholtz, M.M., 1989. Resistance of Nguni, Bonsmara and Hereford cattle to ticks in a bushveld region of South Africa. Onderstepoort J. Vet. Res. 56, 245-250. [ Links ]
Turner, H.G. & Short, A.J., 1972. Effects of field infestations of gastrointestinal helminths and of the cattle tick (Boophilus microplus) on growth of the three breeds of cattle. Aust. J. Agric. Res. 23, 177-193. [ Links ]
Utech, K.B.W., Wharton, R.H. & Kerr, J.D., 1978. Resistance to Boophilus microplus (Canestrini) in different breeds of cattle. Aust. J. Agric. Res. 29, 885-895. [ Links ]
Wharton, R.H., Utech, K.B.W. & Turner, H.G., 1970. Resistance to the cattle tick Boophilus microplus in a herd of Australian Illawarra Shorthorn cattle: Its assessment and heritability. Aust. J. Agric. Res. 21, 163-181. [ Links ]
# Corresponding author. E-mail: norman@arc.agric.za














