Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Animal Science
On-line version ISSN 2221-4062
Print version ISSN 0375-1589
S. Afr. j. anim. sci. vol.38 n.2 Pretoria Feb. 2008
SHORT COMMUNICATION
Leptin gene polymorphism in Indian Sahiwal cattle by single strand conformation polymorphism (SSCP)
P.P. Dubey I; Ashwani SharmaI; D.S. GourII; PrashantII; Anubhav JainII; C.S. MukhopadhyayI; Avtar SinghI; B.K. JoshiI; Dinesh KumarII
IDairy Cattle Breeding Division, National Dairy Research Institute, Karnal, Haryana -132001, India
IIGenes and Genetic Resources Molecular Analysis Lab, National Bureau of Animal Genetic Resources, Karnal -132001, India
ABSTRACT
Leptin, a 16-Kilo dalton protein produced by the obesity (ob) gene, plays an important role in the regulation of feed intake, energy metabolism, growth and reproduction in cattle. The genetic variation of the leptin gene in Sahiwal cattle (Bos indicus) was investigated using an optimized non-radioactive polymerase chain reaction-single strand conformation polymorphism (PCR-SSCP) analysis of 13 amplified fragments covering almost the entire gene. Twenty-eight SSCP band patterns were detected from 10 of these fragments in a sample of 202 Sahiwal cattle. Polymorphisms were detected in the samples, indicating that Sahiwal cattle have high genetic variability in the entire leptin gene. This result opens interesting prospects for future breeding programmes and conservation strategies. These leptin gene variants can be sequenced and screened in the entire population to develop single nucleotide polymorphisms (SNPs) for association studies with different productive and reproductive performances and marker assisted selection.
Keywords: Leptin gene, PCR-SSCP, genetic variability, dairy cattle
India is one of the few countries in the world that has contributed abundantly to the international livestock gene pool. Cattle breeds of Indian origin are being exploited in quite a few countries including Australia, South Africa, Latin America and the USA for developing major livestock economies (FAO, 1998). India has 30 well-defined breeds of cattle (Bos indicus) representing a wide spectrum of genetic variability. The Sahiwal is one of the best dairy breeds of zebu cattle (Nivsarkar et al., 2000). Its native breeding tract is erstwhile Montogomery district now the Sahiwal district (after the name of the breed) in Pakistian. Besides this, some herds are also found in India along the Indo-Pak border in the Ferozepur and Amritsar districts of Punjab and the Sri Ganganagar districts of Rajasthan (Prakash et al., 2005). Nucleotide variation in the coding region of a gene may lead to change in amino acids which alter expressed protein(s), and although intronic variation does not change the amino acid sequence of the protein it may play a significant role in marker assisted selection. In livestock such variation in DNA may also be associated with economic traits, which are governed by many genes each having a small effect (Gelderman, 1997). The obesity (ob) gene was discovered by the positional cloning technique (Zhang et al. 1994). The 167 amino acid protein product of the ob gene was named leptin and consists of three exons of which the first exon is not transcribed into the leptin protein of 16-Kilo dalton. The bovine leptin gene is located at BTA4q32 (Pfister-Genskow et al., 1996) and is involved in the growth and metabolism of animals, and plays an important role in the regulation of feed intake, energy metabolism, growth, production and reproduction of cattle (Ramasay & Cronwell, 1999; Zwrierchoski et al., 2002; Liefers et al., 2002; 2003a; 2003b). The leptin gene is a potential candidate gene for quantitative trait loci (QTL) studies. Some of the more frequently used methodologies for the identification of point mutations are Denaturing Gradient Gel Electrophoresis (DGGE) (Fisher & Lerman, 1980; Patino-Garcia et al., 1999), Temperature Gradient Gel Electrophoresis (TGGE) (Riesner et al., 1989) and Single Strand Conformation Polymorphism (SSCP) (Orita et al., 1989a). Single Strand Conformation Polymorphism is a simple and reliable technique, based on the assumption that changes in the nucleotide sequence of a polymerase chain reaction (PCR) product affect its single strand conformation. Molecules differing by as little as a single base substitution should have different conformers under non-denaturing conditions and migrate differently. Therefore, those differences can be detected as a shift in electrophoretic mobility (Hayashi, 1991). Early reports used radioactivity that limited its widespread use. To simplify and make SSCP analysis more efficient, alternative staining methods with ethidium bromide and with silver (Aninnsworth et al., 1991) have been described. The aim of the present investigation was to identify the genetic variations of the entire leptin gene in Sahiwal cattle using a non-radioactive PCR-SSCP technique.
Two hundred and two genetically unrelated Sahiwal cattle were selected from five different states (Uttar Pradesh, Uttarakhand, Punjab, Haryana and Chhattisgarh) covering the native breeding regions in the country. The unrelated animals were selected on the basis of their history (pedigree) records maintained at different farms. These cattle were considered as representative of the existing gene pool of the population. Blood samples (8 - 10 mL) were collected from the jugular vein, using vacutainers containing 15% ethylenediaminetetraacetic acid (EDTA) as an anticoagulant. Genomic DNA was isolated by the phenolchloroform extraction method (Sambrook & Russell, 2001) and purity was assessed by spectrophotometry. Samples showing an optical density (OD) ratio (260 nm/280 nm) of between 1.7 and 1.9 were used for further analysis while samples outside this range were reprocessed. After checking the quality and quantity of the DNA it was diluted to the final concentration of 50 ng/µL in water and stored at 4 °C. The PCR was carried out on about 50 - 100 ng of genomic DNA in 25 µL per reaction volume. Thirteen primers for PCR -SSCP covering the entire leptin gene (3956 bp long) were designed on nucleotide sequence of the Bos taurus leptin gene (GenBank accession no. BTU50365, Tellam, R. L. 2004) using Primer3 (v.0.4.0) online software (Table 1). Polymerase chain reaction amplification using Tris-HCl (pH 9.0), 0.1% all percentages (w/v) Triton X-100, 1.5 mM MgCl2, 0.75 unit of Taq DNA polymerase and 4 ng/µL of each primer was carried out in a PTC-200 machine (M J Research Inc., MA, USA) pre-programmed for the following conditions: initial denaturation for 5 min at 94 °C followed by 30 cycles (denaturation at 94 °C for 15 s, annealing at 62 - 65 °C for 30 s and extension at 72 °C for 1 min). Amplified products (5 µL) as verified by agarose (1.5%) gel electrophoresis with loading dye (95% formamide, 0.25% bromophenol blue and 0.25% xylene cynol) on 2% (w/v) agarose gel in 1 x TAE buffer using a 100bp ladder as marker for confirmation of the length of the PCR products. Gels were stained with ethidium bromide.
The PCR product was resolved by SSCP analysis. Several factors were tested for each fragment in order to optimize factors such as the amount of PCR product, denaturing solution, acrylamide concentration, percentage cross linking, glycerol, voltage, running time and temperature. Each PCR product was diluted in a denaturing solution (95% formamide, 10 mM NaOH, 0.05% xylene cynol and 0.05% bromophenol blue, 20 mM EDTA), denatured at 85 °C for 13 min, chilled on ice and resolved on polyacrylamide gel. The electrophoresis was carried out in a Bio Rad Protean II xi vertical electrophoresis unit using 1X TBE buffer for SSCP analysis of all the fragments. Gel was silver-stained (Sambrook & Russell, 2001) and dried on cellophane using a Bio Rad 583 gel dryer.
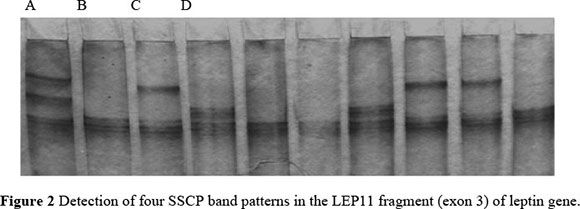
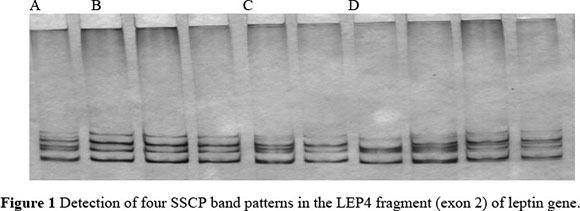
Thirteen leptin fragments (LEP1-LEP13) that covered almost the entire length of the bovine leptin gene, were amplified. The length of PCR products of the leptin gene was confirmed by 100bp ladder in agarose gel electrophoresis. No SSCP band patterns were detected in the LEP6, LPT9 and LEP12 regions under various electrophoretic conditions. However, analysis of the LEP1-LEP5, LEP8, LEP10, LEP11 and LEP13 fragments did reveal the polymorphisms. Twenty-eight band patterns were detected over the entire leptin gene in PCR-SSCP analysis and their respective percentages are shown in Table 3. These variants can further be sequenced to develop SNP markers. These SNP markers would be helpful to breeders for future association studies, selecting superior germplasm and conservation strategies. Single Strand Conformation Polymorphism offers a simple, inexpensive and sensitive method for detecting whether or not DNA fragments are identical in sequence, and so can greatly reduce the amount of sequencing necessary (Orita et al., 1989a; b; Hayashi, 1991; 1992; Hayashi & Yandell, 1993). Several authors have pointed out that PCR-SSCP analysis is an appropriate, reliable and reproducible tool for detection of structural gene polymorphism that is primarily due to point mutations (Neibergs et al., 1993; Sheffield et al., 1993; Barroso et al., 1998; 1999; Kumar et al., 2006). In fact, Neibergs et al. (1993) have stated that PCR-SSCP analysis is the technique of choice when screening point mutations and minor deletion within a given fragment, although it is necessary to optimize conditions for each case. In the present study several factors were tested for each fragment in order to optimize factors such as the amount of PCR product, denaturing solution, acrylamide concentration, percentage cross linking, glycerol, voltage, running time and temperature. Our results provided the first evidence of genetic variability of the leptin gene within the Indian Sahiwal cattle breed. These obtained gel variants from the current study may further be sequenced to identify the SNPs and these SNPs may be useful for establishing a possible association with productive parameters.

Acknowledgments
The authors would like to thank the following people for help in blood sample collection and providing critical information: Sudhakar Yogi, Dean College of Veterinary and Animal Husbandry, Durg, Chhattisgarh; R.J. Sharma, Dean, College of Veterinary and Animal Science, Pantnagar, Uttarakhand; Director, Animal Husbandry, Government of Uttar Pradesh and Director Animal Husbandry, Government of Punjab. We are also thankful to Shushil Kumar, Director, National Dairy Research Institute (NDRI), S.P.S. Ahlawat, Director, National Bureau of Animal Genetic Resources (NBAGR) for providing facilities. The National Fellowship to PPD for support of this work and financial support of the Indian Council of Agricultural Research, Ministry of Agriculture, Government of India is thankfully acknowledged. The critical comments and suggestions of anonymous reviewers and editors are thankfully acknowledged.
References
Aninnsworth, P., Surh, L. & Coulter-Mackie, M., 1991. Diagnostic single strand conformation polymorphism, (SSCP): A simplified non-radioisotopic method as applied to Tay-Sachs B1 variant. Nucleic Acids Res. 19, 405-406. [ Links ]
Barroso, A., Dunner, S. & Canon, J., 1998. Technical note: detection of bovine kappa-casein variants A, B, C and E by means of Polymerase Chain Reaction-Single Strand Conformation Polymorphism (PCR-SSCP). J. Anim. Sci. 76, 1535-1538. [ Links ]
Barroso, A., Dunner, S. & Canon, J., 1999. Technical note: Use of PCR-single strand conformation polymorphism analysis for detection of bovine b-casein variants A1, A2, A3 and B. J. Anim. Sci. 77, 2629-2632. [ Links ]
FAO, 1998. Secondary guidelines for development of national farm animal genetic resources management plans. Measurement of Domestic Animal Diversity: Food and Agricultural Organization of United Nation original working group. http://dad.fao.org/en/refer/library/guideline/workgrp.pdf [ Links ]
Fisher, S. & Lerman, L., 1980. Separation of random fragments of DNA according to properties of their sequences. Proc. Natl. Acad. Sci. USA. 77, pp. 4420-4424. [ Links ]
Gelderman, H., 1997. Investigations on inheritance of quantitative characters in animals by gene markers. 1. Methods. Theor. Appl. Gen. 46, 319-330. [ Links ]
Hayashi, K., 1991. PCR-SSCP: A simple and sensitive method for detection of mutations in the genomic DNA. PCR Methods Appl. 1, pp. 34-38. [ Links ]
Hayashi, K., 1992. PCR-SSCP- A method for detection of mutations. GATA. 9, 73-79. [ Links ]
Hayashi, K. & Yandell, D.W., 1993. How sensitive is PCR- SSCP? Human Mutation 2, 338-346. [ Links ]
Kumar, Dinesh, Gupta, Neelam, Ahlawat, S.P.S., Satyanarayana, R., Sunder, Shyam & Gupta, S.C., 2006. Single Strand Confirmation Polymorphism (SSCP) detection in exon l of the α-lactalbumin gene of Indian Jamunapari milk goats (Capra hircus). Genet. Mol. Biol. 29 (2), 287-289. [ Links ]
Liefers, S.C., Veerkamp, R.F., Pas, M.F.W., Delavaud, C., Chilliard, Y. & Van der Lende, T., 2002. Association between leptin gene polymorphisms and production, live weight, energy balance, feed intake, and fertility in Holstein heifers. J. Dairy Sci. 85, 1633-1638. [ Links ]
Liefers, S.C., Veerkamp, R.F., Pas, M.F.W., Delavaud, C., Chilliard, Y. & Van der Lende, T., 2003a. Leptin concentrations in relation to energy balance, milk yield, intake, live weight, and estrus in dairy cows. J. Dairy Sci. 86, 799-807. [ Links ]
Liefers, S.C., Pas, M.F.W., Virkamp, R.F., Delavaud, C., Chilliard, Y. & Van der Lende, T., 2003b. Association of leptin gene polymorphism with serum leptin concentration in dairy cows. Mamm. Genome 14, 657-663. [ Links ]
Neibergs, H., Dietz, A. & Womack, J., 1993. Single-Strand Conformation Polymorphisms (SSCPs) detected in five bovine genes. Anim. Genet. 24, 81-84. [ Links ]
Nivsarkar, A.E., Vij, P.K. & Tantia, M.S., 2000. Animal genetic resources of India: Cattle and Buffalo. ICAR, India. pp. 382. [ Links ]
Orita, M., Iwahana, H., Kanazawa, H., Hayashi, K. & Sekiya, T., 1989a. Detection of polymorphism of human DNA by gel electrophoresis as single strand confirmation polymorphisms. Proc. National Academy of Science, USA, 86, 2766-2770. [ Links ]
Orita, M., Suzuki, Y., Sekiya, T. & Hayashi, K., 1989b. Rapid and sensitive detection of point mutations and DNA polymorphisms using polymerase chain reaction. Genomics 5, 874-879. [ Links ]
Patino-Garcia, A., Sotillo-Pineiro, E., Modesto, C. & Sierrasesumaga, L., 1999. Screening of the human tumor necrosis factor-alpha (TNF-a) gene promoter polymorphisms by PCR-DGGE analysis. Mutation Research/Mutation Research Genomics 406, 2-4, 121-125. [ Links ]
Pfister-Genskow, M., Hayes, H., Eggen A., & Bishop, M.D., 1996. Chromosomal localization of the bovine obesity (OBS) gene. Mamm. Genome 7, 398-399. [ Links ]
Prakash, B., Singh, Satbir, Singh, P.K., Pundir, R.K., Mukesh, M., Sodhi, M. & Ahlawat, S.P.S., 2005. Cattle genetic resources of India: Sahiwal- The champion dairy breed. National Bureau of Animal Genetic Resources. pp. 80. [ Links ]
Ramasay, T.G. & Cronwell, P.D., 1999. A review- Leptin: A regulation of feed intake and physiology in swine. Proc. seventh Biennial Conf. Aust. Pig Science Association (APSA), 28 Nov. - 1 Dec., Adelaide, Australia. pp. 157-170. [ Links ]
Riesner, D., Steger, G., Zimmat, R., Owens, R., Wagenhofer, M., Hillen, W., Vollach, S. & Henco, K., 1989. Temperature-gradient gel electrophoresis of nucleic acids: Analysis of conformational transitions, sequence variations, and protein-nucleic acid interactions. Electrophoresis 10, 377-389. [ Links ]
Sambrook, J. & Russell, D.W., 2001. Molecular Cloning- A Laboratory Manual. 3rd ed. Cold Spring Harbor, New York. pp. 5.61-5.90. [ Links ]
Sheffield, V., Beck, J., Kwitek, A., Sandstrom, D. & Stone, E., 1993. The sensitivity of single-strand conformation polymorphism analysis for the detection of single base substitutions. Genomics 16, 325-332. [ Links ]
Tellam, R.L., 2004. Direct Submission accession number BTU50365. CSIRO Division of tropical Animal Production, PMB 3, Indooroopilly, QLD 4068, Australia. http://www.ncbi.nlm.nih.gov [ Links ]
Zhang, Y., Proenca R., Maffei M., Barone M., Leopold L. & Friedman, J.M., 1994. Positional cloning of the mouse obese gene and its human homologue. Nature 372, 425-432. [ Links ]
Zwierzchowski, L., Krzyzewski, J., Strzalkowska, N., Siadkowska, E. & Ryniewicz, Z., 2002. Effects of polymorphism of growth hormone (GH), Pit-1 and leptin (LEP) genes, cow's age, lactation stage and somatic cell count on milk yield and composition of Polish black and white cows. Animal Science Papers and Reports 20 (4), 213-227. [ Links ]
 Correspondence:
Correspondence:
Email: dineshkumarbhu@gmail.com














