Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
SAMJ: South African Medical Journal
On-line version ISSN 2078-5135
Print version ISSN 0256-9574
SAMJ, S. Afr. med. j. vol.105 n.5 Pretoria May. 2015
http://dx.doi.org/10.7196/SAMJ.9647
CONTINUING MEDICAL EDUCATION
ARTICLE
Diagnosis of bacterial infection
T H BoylesI; S WassermanII
IMA, BM BCh, MRCP, MD, DTM&H, Cert ID (SA); Division of Infectious Diseases and HIV Medicine, Department of Medicine, Faculty of Health Sciences, Groote Schuur Hospital and University of Cape Town, South Africa
IIMB ChB, MMed, FCP (SA), Cert ID (SA); Division of Infectious Diseases and HIV Medicine, Department of Medicine, Faculty of Health Sciences, Groote Schuur Hospital and University of Cape Town, South Africa
ABSTRACT
Accurate diagnosis of bacterial infection is crucial to avoid unnecessary antibiotic use and to focus appropriate therapy. Bacterial infection is the combination of the presence of bacteria and inflammation or systemic dysfunction; therefore, more than one diagnostic modality is usually required for confirmation. History and examination to determine if a patient fits a clinical case definition is sometimes adequate to confirm or exclude a diagnosis. The second stage is bedside tests - some are used widely, such as urine dipstick tests, but others, such as skin scrapings of petechial rashes, are underutilised. The third stage is laboratory tests - indirect non-culture-based tests, including C-reactive protein and procalcitonin tests, when negative, can be used to prevent the unnecessary use of antibiotics. Direct non-culture-based tests detect antigens or specific antibodies, e.g. group A streptococcal antigen testing can be employed to reduce antibiotic use. Culture-based tests are often considered the reference standard in modern microbiology. Because of slow turnaround times, these tests are frequently used to focus or stop antibiotic therapy after empiric initiation. Nucleic acid amplification tests raise the possibility of detecting organisms with high sensitivity, specificity and reduced turnaround time, and novel diagnostic modalities relying on nanotechnology and mass spectrometry may dramatically alter the practice of microbiology in future.
The Longitude Prize 2014 is a challenge with a £10 million (~ZAR180 million) prize fund to help solve one of the greatest issues of our time.[1] The chosen issue is antimicrobial resistance (AMR) and the challenge is to create a cost-effective, accurate, rapid and easy-to-use test for bacterial infections. Clearly, this is a very important problem that has yet to be fully solved.
Overuse of antibiotics, either because of unnecessary initiation or excessive duration, is a key driver of AMR. The rational use of existing diagnostic tests and development of new technologies to help clinicians confirm or exclude bacterial infection have an important role in improving antibiotic prescribing practices, ultimately improving patient outcomes and reducing AMR.
The first ever bacteriological diagnosis was made in 1676, when Anton van Leeuwenhoek observed bacteria using a single-lens microscope and Robert Koch later advanced the science by focusing on pure culture of organisms. Since then, there has been great progress in these techniques and modern laboratories now also include state-of-the-art molecular diagnostics. This article describes the breadth of diagnostics currently available for bacterial infection and introduces some newer technologies that are on the horizon.
Bacterial infection
Bacterial infection is primarily a clinical concept that may require the use of supportive bedside or laboratory tests to confirm or exclude. There are two broad factors that are always necessary to confirm the diagnosis:
- inflammation or systemic dysfunction; and
- direct or indirect evidence of a compatible bacterial pathogen.
Inflammation may be localised or result in a systemic inflammatory response syndrome (SIRS) (Box 1). Sepsis refers to the presence of a SIRS response caused by presumed or confirmed infection. Severe sepsis occurs if there is accompanying organ dysfunction and septic shock in the case of hypotension requiring ionotropic support.
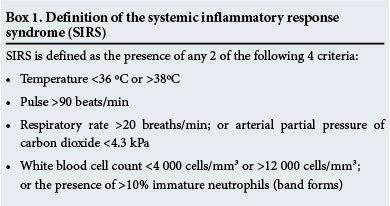
However, these definitions have recently been criticised; e.g. in one study of >100 000 patients with confirmed infection and organ failure, 12% did not meet the criteria for SIRS.[2]
Signs of inflammation are nonspecific, i.e. not limited to bacterial infection, and may be a consequence of non-bacterial infections, trauma, autoimmune diseases or drug reactions. Examples of symptoms associated with local inflammation include dysuria in urinary tract infection and skin redness in cellulitis, but both of these symptoms could also be due to non-infectious causes. Pancreatitis is the classic example of a SIRS response that mimics sepsis, but does not require antibiotics. At the opposite end of the spectrum, bacterial organisms that are recovered from non-sterile sites or other sites without evidence of local inflammation do not necessarily imply infection. For example, a superficial skin swab may culture Staphylococcus aureus, a highly pathogenic and lethal organism, from up to 30% of otherwise healthy people,[3] but does not require antibiotic therapy. Similarly, the finding of nitrites on a urine disptick in the absence of symptoms or urine leucocytes may suggest asymptomatic bacteriuria or bacterial contamination, but not infection. Certain pathogens, however, are considered to be pathological when detected from any site (Mycobacterium tuberculosis is the most important example).
The definition of bacterial infection, and assessing the need for antibiotic therapy, therefore requires clinicians to combine symptoms and signs of inflammation with diagnostic tests for direct or indirect evidence of a pathogen as a cause of the inflammation.
Diagnostic tests
The diagnostic process begins with a full clinical evaluation (history and examination), followed by bedside or laboratory-based investigations. Each has advantages and disadvantages, but they are rarely employed in isolation. The typical process begins with clinical tests and uses this information to guide bedside tests, followed by laboratory tests - with the probability of the diagnosis shifting with each piece of added information. Bacterial infection can occur in any part of the body with multiple different organisms and therefore the number of available tests is vast. This review discusses broad categories of tests, with specific examples to illustrate the process.
Clinical diagnostics
In general, single clinical parameters are neither sensitive nor specific for bacterial infection. For example, fever is common in bacterial infection, but also often absent, and may be caused by a multitude of other illnesses, e.g. as part of the SIRS response. Therefore, it is usual to group together features from the history and examination to create useful case definitions. Usually case definitions are designed to have high sensitivity but low specificity so that the condition is 'ruled out' in patients not fitting the definition. Further testing is then usually required to 'rule in' the diagnosis. An example is the case definition of acute meningitis: <7 days of at least two of the four cardinal features (fever, confusion, headache and neck stiffness). This is useful to exclude patients who are very unlikely to have acute meningitis, but further testing with a lumbar puncture is required to rule in the condition.
Not all case definitions have low specificity and are sometimes the sole basis for confirming a diagnosis. For example, the case definition for community-acquired pneumonia, which includes a chest radiograph, is fever and/or breathlessness or tachypnoea and/ or tachycardia as well as new or progressive infiltrate on a chest radiograph. Patients fulfilling this case definition are considered to have pneumonia, and the condition is excluded in patients to whom the case definition does not apply. Further bedside or laboratory testing is not required to confirm the diagnosis, but may be useful for monitoring purposes.
Bedside tests
The urine dipstick test is a good example of a bedside test that can assist with diagnosing bacterial infection. The absence of nitrites or leucocytes makes urinary tract infection very unlikely, although pyuria in particular may have many alternative causes. Another routinely used bedside test is wet prep microscopy for the diagnosis of sexually transmitted infections in women. A bedside Gram stain can be extremely helpful for identifying Gram-negative diplococci in a joint aspirate to confirm gonoccocal arthritis,[4] or similarly for making a rapid diagnosis of meningococcaemia from a skin scraping of a petechial rash.[5] Unfortunately, these office tests are not often performed, even though they require minimal training and are cheap.
Radiographs usually provide nonspecific evidence of infection, such as infiltrates on a chest radiograph or focal bony lysis suggesting osteomyelitis. More sophisticated radiological techniques such as computed tomography scanning or magnetic resonance imaging are sometimes required, but even these often lack the specificity to confidently rule in bacterial infection. Under these circumstances, it is usually necessary to use a laboratory and possibly a culture-based test to confirm the diagnosis.
Laboratory tests
There are a vast number of relevant laboratory tests that can be broadly divided into indirect and direct non-culture-based and culture-based tests, and nucleic acid amplification tests (NAATs). Indirect tests such as peripheral white cell count (WCC) and C-reactive protein (CRP) confirm the presence of inflammation when positive, but lack the specificity to differentiate bacterial from non-bacterial causes. However, high neutrophil count with left-shift and CRP >100 mg/mL suggest bacterial infection.[6] Negative results generally exclude inflammation and therefore bacterial infection. There has been great interest in developing more sophisticated point-of-care (POC) tests that can accurately exclude bacterial infection. A number of useful tests are now available, including CRP, which have been shown in clinical trials to reduce antibiotic prescription for respiratory tract infections.[7]
Procalcitonin (PCT) is 10 times more expensive than CRP or WCC tests, but more specific for bacterial infection. There is good evidence that antibiotics can safely be withheld, based on low PCT in acute meningitis, acute exacerbations of chronic obstructive pulmonary disease and upper respiratory tract infections.[8] PCT has no value in differentiating bacterial infection from tuberculosis[9] and has very limited value in sepsis.[10] Serial measurements can be used to decide when to discontinue antibiotics, particularly in the intensive care unit setting,[11] although use for this indication may be limited by cost. Current research is examining the complete host-proteome response to bacterial and viral infection to find signatures that can reliably differentiate the two. A recent study showed that the combination of tumour necrosis factor-related apoptosis-inducing ligand (TRAIL), interferon gamma-induced protein-10, and CRP outperformed any individual proteins and routinely used clinical parameters and their combinations.[12]
Direct non-culture-based tests generally depend on identifying an antigen from an organism or a serological reaction to that antigen. These tests vary in their performance characteristics, but in general are specific for a particular bacterial infection in the correct clinical setting. An office-based rapid antigen test is available for the detection of group A Streptoccocus in children with sore throat; a positive test rules in the diagnosis with a high degree of certainty and can be used as a basis to confidently prescribe antibiotics.[13] Similarly, stool antigen tests accurately detect the presence of Helicobacter pylori infection,[14] facilitating the use of eradication therapy without the need for endoscopy. Antigen testing is sometimes preferred over culture for diagnosing bacterial infection due to fastidious or unusual organisms - Legionella pneumonia being a good example. A drawback of many antigen tests is reduced sensitivity compared with culture, and in severe or life-threatening illness this cannot reliably exclude infection. An example is bacterial meningitis, where a positive meningococcal[15] or pneumococcal antigen[16] test on cerebrospinal fluid is excellent at confirming those infections, but a negative test is insufficient for discontinuing empiric therapy. Another drawback of antigen tests is their inability to provide antimicrobial susceptibility data; therefore, even if pneumococcal meningitis is diagnosed on the basis of an antigen test, ceftriaxone will have to be continued until minimum inhibitory concentrations of penicillin can be determined from a cultured isolate.
Serological tests are sometimes used to diagnose infections caused by bacteria that are difficult to culture. Certain classic tests are unreliable and should not be used, such as the Widal reaction for typhoid. Problems with serology include cross-reactivity with other antigens, particularly when measuring IgM, which reduces specificity, and false-negatives in early infection, which reduce sensitivity. Examples of serological tests used in clinical practice to guide treatment are those for Rickettsia in suspected tick bite fever and amoebic serology in liver abscess. For some infections the diagnosis is only confirmed when comparing serological titres from a sample taken at the time of disease with a convalescent sample taken after the patient has recovered. This information is too late to change patient management, but can be useful epidemiologically.
Culture-based tests are the basis of modern microbiology and are usually considered the reference standard with which other diagnostic tests are compared. However, cultures are seldom used in clinical practice to make an initial diagnosis of bacterial infection and rarely influence decisions about whether to initiate empiric antibiotic therapy; these are clinical decisions, influenced by findings on examination and supported by nonspecific bedside and laboratory tests. Culture results, which usually take at least 48 - 72 hours to become available, are then generally used to focus or discontinue antibiotics. Exceptions to this rule are patients who are clinically stable enough for antibiotic therapy to be delayed, and conditions such as pyrexia of unknown origin or suspected subacute bacterial endocarditis, where culturing of organisms is usually required before antibiotics can be initiated.
The performance of cultures is influenced by the type and site of infection, e.g. the sensitivity of blood cultures for detecting an organism in uncomplicated cellulitis is extremely poor because of the small risk of bacteraemia in this condition.[17] In septic shock, however, the probability of a positive blood culture is about 65%. Although cultures are usually considered to have a high specificity, inappropriate indications or poor sampling technique may diminish this greatly.[18] Examples of unreliable culture results include superficial skin swabs or urine collected from long-term indwelling catheters.
Standard culture techniques generally require 48 - 72 hours to provide final results, whereas NAATs have the potential to reduce this window to a few hours. The most widely available is polymerase chain reaction (PCR), which relies on amplification and identification of nuclear material. NAATs provide the greatest advantage in terms of turnaround time when they can be performed directly on clinical specimens, such as a stool sample for Clostridium difficile. [19] Other PCRs require the organism to first be cultured from a clinical specimen, which necessarily increases the time to a result. Detailed information about the infection can be obtained from PCR, e.g. the FilmArray blood culture identification panel tests for 24 organisms and multiple resistance genes.[20] The degree of complexity associated with performing the test also varies, e.g. the GeneXpert system (Cepheid, Sunnyvale, CA, USA) is almost fully automated, requiring samples to be added to sealed cartridges that contain all the reagents for the reaction. More complex systems require highly skilled staff working in high-level laboratories, which increase costs and turnaround time.
Peptide nucleic acid fluorescent in situ hybridisation (PNA FISH) (AdvanDx, Woburn, Mass., USA) uses fluorescence-labelled synthetic oligonucleotide probes that bind to species-specific ribosomes and can be detected using a fluorescence microscope. Modern platforms can perform tests on blood cultures as soon as they become positive, giving a result within 30 minutes so that Gram stain and species identification results can be released simultaneously.
New technologies
New technologies that are likely to emerge as important diagnostic tools in future include nanoparticle probe technology and matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry (MALDI-TOFMS). Nanosphere's Verigene Gram-positive (BC-GP) blood culture assay is performed directly on positive blood cultures using nucleic acid extraction and PCR amplification. Target DNA is then hybridised to oligonucleotides on a microarray with automated qualitative analysis. The test was approved by the US Food and Drug Administration in 2014 and currently has the ability to identify 10 Gram-positive and 8 Gram-negative organisms along with multiple resistance genes. Currently, two MALDI-TOFMS platforms are available in the USA, MALDI Biotyper (Bruker Corporation, Billerica, Mass.) and VitekMS System (bio-Merieux, Durham, NC). MALDI-TOF performs MS on target molecules following ionisation and disintegration; these patterns are compared with known organism fingerprints. It is capable of analysing thousands of samples from specimens per day, including blood, sputum and urine. Another promising infection testing platform that uses PCR followed by electrospray ionisation MS (PCR/ESI-MS) technology is able to rapidly detect >800 bacteria, including unculturable organisms and three classes of antibiotic resistance markers, directly from clinical specimens. In a recent study of 331 blood samples it was able to detect twice as many organisms as culture.[21]
Conclusion
Clinicians must use a combination of clinical signs, bedside tests and laboratory investigations to diagnose bacterial infection. Knowledge of the types of tests available and how to use them appropriately are important tools in improving patient outcomes and rational antibiotic prescribing.
References
1. The Longitude Prize 2014. https://longitudeprize.org/ (accessed 6 February 2015). [ Links ]
2. Kaukonen KM, Bailey M, Pilcher D, Cooper DJ, Bellomo R. Systemic inflammatory response syndrome criteria in defining severe sepsis. N Engl J Med 17 March 2015. [Epub ahead of print] [ Links ]
3. Kluytmans J, van Belkum A, Verbrugh H. Nasal carriage of Staphylococcus aureus: Epidemiology, underlying mechanisms, and associated risks. Clin Microbiol Rev 1997;10(3):505-520. [ Links ]
4. Bardin T. Gonococcal arthritis. Best Pract Res Clin Rheumatol 2003;17(2):201-208. [ Links ]
5. Arend SM, Lavrijsen AP, Kuijken I, van der Plas RN, Kuijper EJ. Prospective controlled study of the diagnostic value of skin biopsy in patients with presumed meningococcal disease. Eur J Clin Microbiol Infect Dis 2006;25(10):643-649. [ Links ]
6. Povoa P. C-reactive protein: A valuable marker of sepsis. Intensive Care Med 2002;28(3):235-243. [ Links ]
7. Huang Y, Chen R, Wu T, Wei X, Guo A. Association between point-of-care CRP testing and antibiotic prescribing in respiratory tract infections: A systematic review and meta-analysis of primary care studies. Br J Gen Pract 2013;63(616):e787-e794. [http://dx.doi.org/10.3399/bjgp13X674477] [ Links ]
8. Schuetz P, Chiappa V, Briel M, Greenwald JL. Procalcitonin algorithms for antibiotic therapy decisions: A systematic review of randomized controlled trials and recommendations for clinical algorithms. Arch Intern Med 2011;171(15):1322-1331. [http://dx.doi.org/10.1001/archinternmed.201L318] [ Links ]
9. Huang SL, Lee HC, Yu CW, et al. Value of procalcitonin in differentiating pulmonary tuberculosis from other pulmonary infections: A meta-analysis. Int J Tuberc Lung Dis 2014;18(4):470-477. [http://dx.doi.org/10.5588/ijtld.13.0449] [ Links ]
10. Wacker C, Prkno A, Brunkhorst FM, Schlattmann P. Procalcitonin as a diagnostic marker for sepsis: A systematic review and meta-analysis. Lancet Infect Dis 2013;13(5):426-435. [http://dx.doi.org/10.1016/S1473-3099(12)70323-7] [ Links ]
11. Assink-de Jong E, de Lange DW, van Oers JA, Nijsten MW, Twisk JW, Beishuizen A. Stop antibiotics on guidance of procalcitonin study (SAPS): A randomised prospective multicenter investigator-initiated trial to analyse whether daily measurements of procalcitonin versus a standard-of-care approach can safely shorten antibiotic duration in intensive care unit patients - calculated sample size: 1816 patients. BMC Infect Dis 2013;13:178. [http://dx.doi.org/10.1186/1471-2334-13-178] [ Links ]
12. Oved K, Cohen A, Boico O, et al. A novel host-proteome signature for distinguishing between acute bacterial and viral infections. PLoS One 2015;10(3):e0120012. [http://dx.doi.org/10.1371/joumaLpone.0120012] [ Links ]
13. Bisno AL, Gerber MA, Gwaltney JM, Jr, Kaplan EL, Schwartz RH, Infectious Diseases Society of America. Practice guidelines for the diagnosis and management of group A streptococcal pharyngitis. Clin Infect Dis 2002;35(2):113-125. [ Links ]
14. Lee YC, Chiu HM, Chiang TH, et al. Accuracy of faecal occult blood test and Helicobacter pylori stool antigen test for detection of upper gastrointestinal lesions. BMJ Open 2013;3(10):e003989. [http://dx.doi.org/10.1136/bmjopen-2013-003989] [ Links ]
15. Sobanski MA, Barnes RA, Coakley WT. Detection of meningococcal antigen by latex agglutination. Methods Mol Med 2001;67:41-59. [ Links ]
16. Samra Z, Shmuely H, Nahum E, Paghis D, Ben-Ari J. Use of the NOW Streptococcus pneumoniae urinary antigen test in cerebrospinal fluid for rapid diagnosis of pneumococcal meningitis. Diagn Microbiol Infect Dis 2003;45(4):237-240. [ Links ]
17. Perl B, Gottehrer NP, Raveh D, Schlesinger Y, Rudensky B, Yinnon AM. Cost-effectiveness of blood cultures for adult patients with cellulitis. Clin Infect Dis 1999;29(6):1483-1488. [ Links ]
18. Aronson MD, Bor DH. Blood cultures. Ann Intern Med 1987;106(2):246-253. [ Links ]
19. Pancholi P, Kelly C, Raczkowski M, Balada-Llasat JM. Detection of toxigenic Clostridium difficile: Comparison of the cell culture neutralization, Xpert C. difficile, Xpert C. difficile/Epi, and Illumigene C. difficile assays. J Clin Microbiol 2012;50(4):1331-1335. [http://dx.doi.org/10.1128/JCM.06597-11] [ Links ]
20. Bauer KA, Perez KK, Forrest GN, Goff DA. Review of rapid diagnostic tests used by antimicrobial stewardship programs. Clin Infect Dis 2014;59(Suppl 3):S134-S145. [http://dx.doi.org/10.1093/cid/ciu547] [ Links ]
21. Bacconi A, Richmond GS, Baroldi MA, et al. Improved sensitivity for molecular detection of bacterial and Candida infections in blood. J Clin Microbiol 2014;52(9):3164-3174. [http://dx.doi.org/10.1128/JCM.00801-14] [ Links ]
 Correspondence:
Correspondence:
T H Boyles
tomboyles@yahoo.com














