Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
SAMJ: South African Medical Journal
On-line version ISSN 2078-5135
Print version ISSN 0256-9574
SAMJ, S. Afr. med. j. vol.100 n.12 Pretoria Dec. 2010
ORIGINAL ARTICLES
Bacteria isolated from bloodstream infections at a tertiary hospital in Dar es Salaam, Tanzania - antimicrobial resistance of isolates
S MoyoI; S AboudI; M KasubiIII; S Y MaselleII
IMD, MMed (Microbiology and Immunology). Department of Microbiology and Immunology, Muhimbili University of Health and Allied Sciences, Dar es Salaam, Tanzania
IIMD, PhD. Department of Microbiology and Immunology, Muhimbili University of Health and Allied Sciences, Dar es Salaam, Tanzania
IIIMD, MMed (Microbiology and Immunology), PhD. Department of Laboratory Services, Microbiology Section, Central Pathology Laboratory Muhimbili National Hospital, Dar es Salaam
ABSTRACT
OBJECTIVES: A bloodstream infection (BSI) is a life-threatening condition. We studied the causative agents of BSIs and antimicrobial susceptibility patterns of bacterial isolates at Muhimbili National Hospital (MNH) in Dar es Salaam, Tanzania.
METHODS: A retrospective analysis of blood culture results obtained at MNH from January 2005 to December 2009 was done. Blood culture isolates judged to be clinically significant and antimicrobial susceptibility results of the bacteria were included. The frequencies and proportions of bacteria isolated and antimicrobial susceptibility results were analysed and compared using Pearson's chi-square test and Fisher's exact test where applicable, or the Mann-Whitney U-test.
RESULTS: A total of 13 833 blood cultures were performed. Bacterial pathogens were detected in 1 855 (13.4%), Gram-positive bacteria (1 523; 82.1%) being significantly more prevalent than Gramnegative bacteria (332; 17.9%) (p=0.008). The most common bacterial pathogens isolated were coagulase-negative staphylococci (CoNS) (1 250; 67.4%), S. aureus (245; 13.2%), Escherichia coli (131; 7%) and Klebsiella spp. (130; 7.0%). All bacteria isolated showed high resistance to penicillin G (70.6%), tetracycline (63.8%), cefotaxime (62.5%) and ampicillin (62.3%). Moderate to high resistance was seen against chloramphenicol (45.2%), erythromycin (35.0%), ciprofloxacin (29.3%), co-trimoxazole (25.0%) and gentamicin (23.5%). Of S. aureus isolates, 23.3% were resistant to methicillin.
CONCLUSION: CoNS accounted for two-thirds of the bacterial pathogens isolated. High-level resistance was seen to first-line and inexpensive antimicrobial agents. Routine screening for extendedspectrum beta-lactamase production and methicillin resistance among Gram-negative rods and S. aureus from blood cultures should be instituted to monitor spread of multidrug-resistant isolates.
A bloodstream infection (BSI) is a life-threatening condition that may be complicated by septic shock and death. BSI due to bacteria is an important cause of morbidity and mortality.1 Gram-negative bacteria have been associated with more deaths than Gram-positive bacteria.1 Mortality due to septic shock can be as high as 60% despite treatment. A better understanding of the spectrum of pathogens causing BSI is crucial for prompt management of patients, as antimicrobial therapy greatly influences the outcome of patients with BSI.1
Over the past 20 - 30 years there have been significant changes in the causation and epidemiology of BSI. While Staphylococcus aureus and Escherichia coli continue to be the most common causative agents, there has been a substantial increase in BSI due to coagulasenegative staphylococci (CoNS).
Recent studies have reported an increasing frequency of antimicrobial resistance among microbial pathogens causing BSI in Tanzania and elsewhere,2,3 and several classes of antimicrobial agents have become less effective as a result of the emergence of antimicrobial resistance, often caused by the selective pressure of antimicrobial usage. The important emerging problems are resistance to extended-spectrum cephalosporins and fluoroquinolones in the Enterobacteriaceae, oxacillin resistance in staphylococci, penicillin resistance in streptococci, and vancomycin resistance in enterococci (and eventually staphylococci).2,4 The changing spectrum of microbial pathogens together with antimicrobial resistance trends and clinical significance necessitate continuous monitoring of antimicrobial resistance. Surveillance of antimicrobial resistance is critical in guiding physicians towards appropriate choice of antimicrobial agents to treat BSI. We aimed to describe the cauative agents and antimicrobial susceptibility patterns of bacterial isolates from patients with BSI.
Methods
The study was a retrospective analysis of blood cultures performed during the past 5 years (January 2005 -December 2009) at the Microbiology Section, Central Pathology Laboratory (CPL), Muhimbili National Hospital (MNH), Dar es Salaam, Tanzania. MNH is a tertiary health care facility handling patients with severe disease and those who require specialised management from three municipal hospitals in Dar es Salaam. All blood culture isolates that were detected and judged to be clinically significant, and antimicrobial drug susceptibility results of the bacteria isolated, were reviewed and included. Data including date of blood culture specimen collection, admitting hospital department (intensive care unit (ICU), paediatrics), blood culture and antimicrobial susceptibility results were recorded using a study data collection form.
Bacterial isolation, identification and antimicrobial susceptibility testing
Micro-organisms were isolated and identified according to a standard operating procedure.5 Blood culture bottles received at the CPL were incubated aerobically at 37ºC for 7 days. The bottles were examined daily for evidence of bacterial growth, haemolysis, gas production or turbidity above the red cell line. Subcultures using sterile syringes were done on blood agar, chocolate agar and MacConkey agar media daily for 7 days before blood cultures were reported as negative.6 For antimicrobial susceptibility, disc diffusion tests were performed for all the isolates according to the method recommended by the Clinical and Laboratory Standard Institute.7 Antibiotic disc (Oxoid) was applied to each plate, and after incubation at 35ºC for 24 hours zone sizes were measured and interpreted accordingly.
Statistical analysis
Data were entered in Microsoft Excel 2000 (Microsoft Corp., Redmond, WA). The frequencies and proportions of bacteria isolated and antimicrobial susceptibility results were analysed. The data were also subjected to normality and homogeneity of variance tests. Where assumptions for parametric tests were not met, data were subjected to non-parametric tests and analysed using GraphPad InstantTM 1990 -1993 software. A p-value of <0.05 was regarded as statistically significant.
Results
During the 5-year period, a total of 13 833 blood cultures were performed, 8 682 (62.8%) from children and 5 151 (37.2%) from adults. Bacterial pathogens were detected in 1 855 (13.4%) of the cultures; of these 48.8% (906) were from children and 51.2% (949) from adults. Positive blood cultures were detected in only 33 blood specimens from the ICU (1.8%) compared with 1 822 (98.2%) from other wards. Gram-positive bacteria (1 523; 82.1%) were significantly more prevalent (p=0.008) than Gram-negative bacteria (332; 17.9%) (Fig. 1).
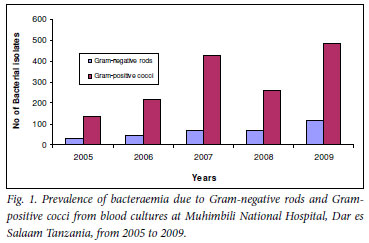
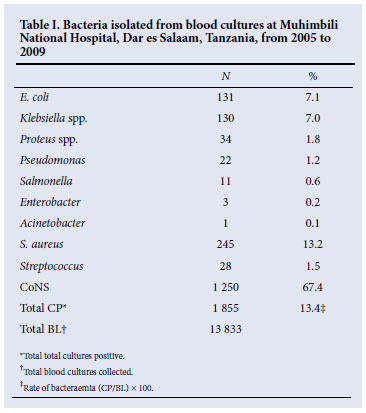
The most common bacterial pathogens isolated were CoNS (1 250; 67.4%), S. aureus (245; 13.2%), E. coli (131; 7%) and Klebsiella spp. (130; 7.0%). Other bacterial pathogens detected were Proteus spp. (34; 1.8%), Pseudomonas spp. (22; 1.2%), Streptococcus spp. (28; 1.5%), Salmonella spp. (11; 0.6%), Enterobacter spp. (3; 0.2%) and Acinetobacter spp. 1 (0.1%) (Table I). Rates of detection of E. coli and Proteus spp. have been increasing over the years, while those of Klebsiella spp. and S. aureus have fluctuated in the past 5 years.
All bacteria isolated in our study showed a high level of resistance to the commonly used antimicrobial agents, including penicillin G (70.6%), tetracycline (63.8%), cefotaxime (62.5%) and ampicillin (62.3%). There were moderate to high levels of resistance to chloramphenicol (45.2%), erythromycin (35.0%), ciprofloxacin (29.3%), co-trimoxazole (25.0%) and gentamicin (23.5%). Grampositive bacteria showed a very low level of resistance to vancomycin (1.5%), whereas S. aureus showed moderate level (23.3%) of resistance to methicillin (Table II).
Discussion
Clinically significant BSI is associated with high morbidity and mortality and considerably impacts on health care costs, especially when caused by antimicrobial-resistant bacteria. In our study 13.4% of all blood cultures yielded positive results. The low rate may be explained by the fact that most patients probably received clinical care, including antimicrobial agents, at the municipal hospitals before admission to MNH, the referral hospital. Information on patients' clinical history before referral, including the use of antimicrobial agents, was not available.
Our study showed that Gram-positive bacteria (1 523; 82.1%) were significantly more prevalent than Gram-negative bacteria (332; 17.9%). These findings concur with those reported in previous studies in Tanzania and other African countries.8,9 The current shift from Gram-negative to Gram-positive bacteria in causing BSI has been explained by others.6 It has been suggested that this change is due to evolution of medical care, Gram-positive cocci having begun to re-emerge as predominant pathogens in BSI in the early 1980s.10
The most commonly isolated bacteria in this study were CoNS, followed by S. aureus, E. coli and Klebsiella spp. Our findings are consistent with the results of other surveillance studies.9,11 CoNS are one of the most common causes of nosocomial BSI and also the most common blood culture contaminant.11 Because only one blood culture was obtained for each of our study patients, it was not possible to determine whether patients who had CoNS isolated had a true bacteraemia or the finding was due to skin contamination. It is also possible that the recovery of CoNS could have resulted from other factors such as prolonged use of invasive intravascular devices, prolonged hospital stay, or other underlying co-morbidities. However, other previous studies identified CoNS (37%) as the leading pathogens in BSI.10 The increased isolation of CoNS in blood cultures may also reflect a change from regarding these organisms as normal skin flora to viewing them as clinically significant. This is supported by the tremendous increase in the isolation of CoNS from 49 isolates in the year 2005 to 476 isolates in the year 2009. We therefore recommend that in our tertiary hospital guidelines should be provided to ensure that two sets of blood cultures are obtained to distinguish true CoNS bactaeremia from contamination. Hospital staff responsible for drawing blood for cultures should also receive training on aseptic technique.12 S. aureus was the most common pathogen detected, accounting for 13.2% of all positive blood cultures.
Paediatric wards accounted for 62.8% of blood cultures in this study, but few strains of S. pneumoniae were recovered and there was no recovery of H. influenzae from children. These pathogens are common causes of invasive bacterial infections in an unvaccinated paediatric population.13-15 The World Health Organization (WHO) estimates that H. influenzae is the leading cause of bacterial meningitis in children under 5 years of age and the second leading cause of deaths due to bacterial pneumonia in resource-limited areas where vaccination is not performed.16,17 However, H. influenzae type B vaccination was introduced in 2009 and could not have influenced the current findings. It is possible that laboratory-prepared culture media made with human blood might inhibit or not support the growth of strains of H. influenzae. A laboratory surveillance system for S. pneumoniae and H. influenzae is needed as an important first step to determine the prevalence and incidence of invasive pneumococcal and H. influenzae infections, and to measure the impact of the introduction of vaccination. As noted, measures to build laboratory capacity will ultimately improve diagnostic and surveillance capabilities.
Mortality and morbidity from BSI infections are greater when caused by antimicrobial-resistant bacteria, and proper treatment becomes a challenge.18 We observed resistance to several antimicrobial agents used for first-line and inexpensive treatment of BSI, such as ampicillin, co-trimoxazole, penicillin and gentamicin. This concurs with findings of Aboud et al., who reported resistance to first-line antibiotics among bacterial isolates from BSI in the same hospital.8 We also documented high methicillin resistance among S. aureus; 23.3% of our bloodstream isolates were resistant to methicillin. The methicillin-resistant S. aureus (MRSA) rate found in this study is higher than that reported 10 years ago from the same hospital (0.4%), and relatively higher than reported 5 years ago (12%).18,19 In Bugando referral hospital in northern Tanzania a prevalence of MRSA of 16.2% was reported,20 implying that there has been an increasing trend in MRSA spread in the country, as is the case worldwide.
Although not investigated in our study, bacteraemia due to MRSA is associated with high mortality compared with methicillinsensitive S. aureus bacteraemia.9 The Enterobacteriaceae in this study demonstrated a high level of resistance (62.5%) towards third-generation cephalosporins. Third-generation cephalosporin resistance is usually caused by expression of extended-spectrum betalactamase (ESBL) enzymes, and this was probably the case among our strains, although specific confirmation of ESBL production was not performed. ESBL strains have been reported previously in septicaemic and ICU patients from the same hospital.2,3
Lack of enforcement of mechanisms regulating antibiotic use as a part of infection control programmes could have influenced the pattern of resistance results to a considerable degree. In addition, there are no guidelines to isolate hospitalised patients found to carry MRSA or ESBL producing strains. This situation in part explains the increased spreading of MRSA and ESBL strains observed in this study.
Conclusions
CoNS accounted for two-thirds of the bacterial pathogens isolated in our study. A high level of resistance to first-line and inexpensive antimicrobial agents was seen. Routine screening for ESBL production and methicillin resistance among Gram-negative rods and S. aureus from blood cultures should be instituted to monitor spread of multidrug-resistant isolates.
Competing interests. The authors have no competing interests to declare.
We would like to thank all the technical staff in the Microbiology Section, Central Pathology Laboratory, Muhimbili National Hospital, for performing the blood cultures and antibiotic susceptibility testing, and Donald Chigamba for retrieval and recording the data for analysis.
References
1. Weinstein MP, Towns ML, Quartey SM, et al. The clinical significance of positive blood cultures in the 1990s: a prospective comprehensive evaluation of microbiology, epidemiology, and outcome of bacteremia and fungemia in adults. Clin Infect Dis 1997; 24: 584-602. [ Links ]
2. Ndugulile F, Jureen R, Harthug S, Urassa W, Langeland N. Extended spectrum β- lactamses among Gram-negative bacteria of nosocomial origin from an intensive care unit of a tertiary health facility in Tanzania. BMC Infect Dis 2005; 5: 85-91. [ Links ]
3. Blomberg B, Jureen R, Manji KP, et al. High rate of fatal cases of pediatric septicemia caused by Gram-negative bacteria with extended-spectrum beta-lactamases in Dar es Salaam, Tanzania. J Clin Microbiol 2005; 43:745-749. [ Links ]
4. Shah AA, Hasan F, Ahmed S, Hameed A. Characteristics, epidemiology and clinical importance of emerging strains of Gram-negative bacilli producing extended-spectrum β-lactamases. Res Microbiol 2004; 155: 409-421. [ Links ]
5. Baron EJ, Fineglod SM, Peterson LR. Methods for identification of etiologic agents of infectious diseases. In: Forbes BA, Sahm DF, Weissfeld AS, eds. Bailey & Scott's Diagnostic Microbiology. 10th ed. St Louis, Mo.: Mosby, 1996: 327-529. [ Links ]
6. Cheesbrough M. District Laboratory Practice in Tropical Countries. Part II. Cambridge: Cambridge University Press, 2006: 63. [ Links ]
7. Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing: 16th Informational Supplement (CLSI document M100-S16). Wayne, Penn.: CLSI, 2006. [ Links ]
8. Aboud S, Sabaya S, Msangi V. Bacteria isolated from blood culture specimens at Muhimbili National Hospital in Dar es Salaam, Tanzania from 1995-2004. Tanzania Medical Journal 2005; 20(2): 22-26. [ Links ]
9. Ahmed SH, Daef EA, Badary MS, Mahmoud MA, Abd-Elsayed AA. Nosocomial blood stream infection in intensive care units at Assiut University Hospitals (Upper Egypt) with special reference to extended spectrum β-lactamase producing organisms. BMC Research Notes 2009; 2: 76. [ Links ]
10. Warran D, Zack J, Elward A, Cox M, Fraser V. Nosocomial primary bloodstream infections in intensive care unit patients in a non teaching community medical center: a 21-month prospective study. Clin Infect Dis 2001; 33: 1329-1335. [ Links ]
11. Newton O, English M. Young infant sepsis: aetiology, antibiotic susceptibility and clinical signs. Trans R Soc Trop Med Hyg 2007; 101: 959-966. [ Links ]
12. Levine OS, O'Brien KL, Knoll M, Adegbola RA, Black S, Cherian T. Pneumococcal vaccination in developing countries. Lancet 2006; 367: 1880-1882. [ Links ]
13. World Health Organization. Global program for vaccines and immunization. The WHO position paper on pneumococcal vaccines. Wkly Epidemiol Rec 1999; 74: 177-184. [ Links ]
14. Consortium. Serious community-acquired paediatric infections in rural Asia (Bohol Island, Philippines): bacterial meningitis in children less than 5 years of age. Scand J Infect Dis 2007; 39: 983-989. [ Links ]
15. Zaidi AK, Huskins WC, Thaver D, Bhutta ZA, Abbas Z, Goldmann DA. Hospital-acquired neonatal infections in developing countries. Lancet 2005; 365: 1175-1188. [ Links ]
16. Weinstein MP. Blood culture contamination: persisting problems and partial progress. J Clin Microbiol 2003; 41: 2275-2278. [ Links ]
17. Allegranzi B, Pittet D. Healthcare-associated infection in developing countries: simple solutions to meet complex challenges. Infection Control and Hospital Epidemiology 2007; 28: 1323-1327. [ Links ]
18. Bloomberg B, Manji KP, Urassa WK, et al. Antimicrobial resistance predicts death in Tanzanian children with bloodstream infections: a prospective cohort study. BMC Infect Dis 2007; 7: 43. [ Links ]
19. Urassa WK, Haule EA, Kagoma C, et al. Antimicrobial susceptibility of Staphylococcus aureus strains at Muhimbili Medical Centre, Tanzania. East Afr Med J 1999; 76: 693-695. [ Links ]
20. Mshana SE, Kamugisha E, Mirambo M, et al. Prevalence of clindamycin inducible resistance among methicillin-resistant Staphylococcus aureus at Bugando Medical Centre, Mwanza, Tanzania. Tanzania Journal of Health Research 2009; 11: 59-64. [ Links ]
Accepted 30 June 2010.
Corresponding author: S Moyo (sabmoyo26@yahoo.com)














