Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Science
On-line version ISSN 1996-7489
Print version ISSN 0038-2353
S. Afr. j. sci. vol.110 n.9-10 Pretoria Oct. 2014
http://dx.doi.org/10.1590/sajs.2014/20130041
RESEARCH ARTICLE
Characterisation of Mycosphaerella species associated with pink spot on guava in South Africa
Adriaana JacobsI; Mariette TruterI; Maritha H. SchoemanII
IBiosystematics Programme, Plant Protection Research Institute, Agricultural Research Council, Pretoria, South Africa
IICrop Protection Programme, Institute for Tropical and Subtropical Crops, Agricultural Research Council, Nelspruit, South Africa
ABSTRACT
Pink spot symptoms on guava fruit in the Lowveld region were in the past attributed to Colletotrichum gloeosporioides, but recently Mycosphaerella species were suggested to be part of a disease complex, including pink spot symptoms. During routine surveys of guava diseases in the Lowveld area of the Mpumalanga Province in South Africa, Mycosphaerella species were consistently isolated from guava fruit. Colletotrichum gloeosporioides was also retrieved, especially from older, bigger lesions. The Mycosphaerella isolates were compared based on their growth characteristics in culture and on DNA sequences of the internal transcribed spacer region, large subunit of the ribosomal DNA as well as the β-tubulin and translation elongation factor 1α gene regions. The phylogenetic analyses indicate that the isolates from the present study represent at least three species not previously reported on guavas. This report is therefore the first report of Mycosphaerella species associated with Psidium guajava in South Africa.
Keywords: Capnodiales; Psidium guajava; multi-gene phylogeny; internal transcribed spacer region.
Introduction
Guava (Psidium guajava L.) is a fruit-bearing tree in the Myrtaceae plant family. Closely related genera include Eucalyptus L'Heritier and Syzygium R.Br. ex Gaertn. The tree is native to Mexico, the Caribbean and Central America, where it is an economically important crop.1,2 Leading producers of guava fruit are Brazil, India and Mexico.2 In South Africa, approximately 41 000 tonnes of guava are harvested per annum for fresh sales and processing.3
Several diseases affecting the health and productivity of guava plants and fruit have been reported internationally. Of these, anthracnose, caused by Colletotrichum gloeosporioides (Penz.) Penz. & Sacc., is the most common fruit disease in guava-growing countries globally.2 In Puerto Rico, up to 50% of the guava crop (mainly from wild trees) is threatened by anthracnose, which mummifies and blackens immature fruits and rots mature fruits. Similarly, dry rot, caused by Lasiodiplodia theobromae (Pat.) Griffon & Maubl. (Diplodia natalensis Pole-Evans), has been reported to affect 40% of the crop on some trees in South India.1
Fruit diseases occurring on guava in South Africa include anthracnose, blossom end rot (caused by Phomopsis psidii Nag Raj & Ponnappa) and pink spot.4,5 Pink spot symptoms were in the past attributed to C. gloeosporioides6 - the same fungus causing anthracnose - but later the possibility of a Guignardia sp. as the causal agent was investigated (Schoeman, unpublished data). Recently, it has been suggested that Mycosphaerella-like species could be associated with the small pink and red/purple specks of the disease called pink spot. These specks have been seen on fruit in all guava production areas in South Africa, but are most severe in the Mpumalanga Province.5
Mycosphaerella Johanson is one of the largest genera of plant pathogenic fungi, many of which cause economically important diseases in temperate and tropical crops.7-10Mycosphaerella species occur on all aboveground plant parts including the leaves and stems of several hundred different host plants.11Mycosphaerella can spread to healthy hosts either as waterborne conidia or as airborne ascospores. Once in contact with compatible host tissue, the spores can germinate and penetrate the plant through the leaf stomata.8
Identification of Mycosphaerella species based on morphology alone is impossible. The production of very small fruiting structures with highly conserved morphology, host-specificity and poor growth in culture contribute to a lack of informative morphological characteristics. Although ascospore germination patterns812 and morphology of the asexual state13 greatly facilitate species identification, co-inhabitancy14 makes it difficult to link the asexual state from the isolated culture to their correct sexual state as observed on the host tissue15 using only morphological approaches.
Comparisons of DNA-based methods such as RAPDs, species-specific primers, microsatellites and DNA sequence data have in recent years been employed to distinguish between Mycosphaerella species, resulting in substantial changes to genus and species concepts in this group.12·13·15-17 The majority of studies employing DNA sequence data for species identification has relied on sequence data from the internal transcribed spacer (ITS) region, large subunit of the ribosomal DNA (LSU) as well as translation elongation factor 1 α (TEF) gene regions.18-20
The objective of this study was to identify possible Mycosphaerella species associated with pink spot on FI guajava fruit in the Lowveld area of the Mpumalanga Province, South Africa. Identification was performed using DNA sequence data of the ITS, LSU, TEF as well as the β-tubulin (BT) gene regions.
Materials and methods
Symptoms and isolations
Symptomatic guava fruit were collected from various orchards in the Mpumalanga Province of South Africa during the 2007-2009 production seasons and field observations were recorded. Fruit were surface disinfected by spraying twice with 75% ethanol, followed by air drying. Lesions of different sizes were randomly selected, aseptically removed and small (1-2 mm in diameter) segments plated on potato dextrose agar (PDA) (Biolab, Merck, Darmstadt, Germany) in Petri dishes. Plates were incubated at 25 °C, under 24-h cool white fluorescent light for 3-4 weeks. The emerging fungal colonies were examined and selected colonies were purified for further identification.
Isolates were sub-cultured onto carnation leaf agar, malt extract agar (MEA) (Biolab, Merck, Darmstadt, Germany) and oatmeal agar (OA) for morphological grouping and determination of growth characteristics.21 Petri dishes were incubated at 25 °C under continuous near-ultraviolet light, and examined weekly for 2 months. Mounts were prepared in lactic acid and examined using phase and bright-field phase contrast microscopy at x400 and x1000 magnification. Colony colour was assigned based on Rayner's colour chart.22 Cultures are maintained in the South African National Collections of Fungi (PPRI collection) (Table 1).
DNA sequence comparisons
DNA extraction and amplification
All isolates obtained were grown on PDA at 25 °C for 21 days. DNA was isolated using the DNAeasy® Plant Mini Kit (Qiagen, Valencia, CA, USA) according to the manufacturer's specifications. Extracted DNA was used as a template in polymerase chain reactions (PCR) to amplify a part of the ITS region using the primer set ITS 1 (5'-TCCGTAGGTGAACCTGCGG-3') and ITS 4 (5'-TCCTCCGCTTATTGATATGC-3')23, LSU region (including domains D1-D3) using the primer set LR0R (5'-ACC CGC TGA ACT TAA GC-3')24 and LR7 (5'-TAC TAC CAC CAA GAT CT-3')25, BT region using the primer set TUB2Fd (5'-GTB CAC CTY CAR ACC GGY CAR TG-3') and TUB4Rd (5'-CCR GAY TGR CCR AAR ACR AAG TTG TC-3')26 and TEF region using primer set EF1-728F (5'-CAT CGA GAA GTT cGa GAAGG-3') and EF2-986R (5'-TAC TTG AAG GAA CCC TTACC-3')27.
The PCR reactions consisted of 1x Roche Taq reaction buffer with MgCl2, dNTPs (250 µM each), primers (0.2 µM each), template DNA (25 ng) and Roche Taq polymerase (0.5 U) (Roche Pharmaceuticals, Berlin, Germany). The PCR reaction conditions for the amplification of the ITS gene region were an initial denaturation at 94 °C for 2 min, followed by 35 cycles of denaturation at 94 °C for 1 min, annealing at 52 °C for 1 min and elongation at 72 °C for 1 min, with a final elongation step of 72 °C for 5 min. PCR cycling conditions for the LSU, BT and TEF gene regions differed from the above-mentioned protocol only in the annealing temperatures (62 °C for LSU and 52 °C for BT and TEF). The resulting amplicons were purified using a QIAquick PCR Purification Kit (Qiagen, Hilden, Germany).
DNA sequencing and sequence comparisons
DNA sequences were determined from PCR amplicons using the ABI PRISM™ Dye Terminator Cycle Sequencing Ready Reaction Kit with AmpliTaq® DNA Polymerase, (Applied Biosystems, Paisley, UK) using both forward and reverse primers for each gene region. Sequences generated in this study have been deposited in GenBank (Table 1).
BLAST comparisons on the Mycosphaerellaceae database (MycoBank, www.mycobank.org) were done for the ITS, TEF, LSU and BT sequences generated. These comparisons were performed to enable the construction of data sets for phylogenetic analyses and comparisons with closely related species. Multiple sequence alignments for all gene regions were generated for our data using MAFFT version 5.28 Gaps were treated as missing data in the subsequent analysis. Phylogenetic analysis was based on parsimony using PAUP 4.0* (Phylogenetic Analysis Using Parsimony *and Other Methods version 4).29 Heuristic searches were done with random addition of sequences (100 replicates), tree bisection-reconnection branch swapping, MULPAR effective and MaxTrees set to auto-increase. The combinability of the data sets was determined by the partition homogenicity test.30 Species previously isolated from Myrtaceae plant species were included in the final phylogenetic data set. The consistency and retention indices were determined for the data set. The phylogenetic tree was rooted with Uwebraunia commune (Crous & Mansilla) Crous as monophyletic sister outgroup to the rest of the taxa. Bootstrap analyses were performed to determine branching point confidence intervals (1000 replicates) for the most parsimonious trees generated for the respective data sets.
Results
Symptoms and isolations
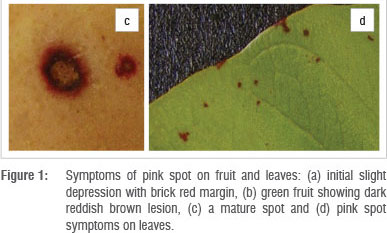
Spots and lesions were observed on fruit from February to September, depending on the time of pruning, and on leaves from December to August during the 3-year inspection period. Symptoms on fruit developed before harvesting and did not increase significantly in size after harvest, but became more obvious as the fruit changed colour. Single spots originated from an initial slight depression (1-2 mm in diameter) with a brick-red margin (Figure 1a). On green fruit, lesions had a dark reddish-brown colour (Figure 1b) and as the fruit coloured, the centres became light brown and the brick red margins showed up more clearly. As a spot aged, it enlarged to about 3-4 mm in diameter with the tissue in the centre turning lighter brown, surrounded by a dark brown to purple-red margin which faded to a brick red colour (Figure 1c). Not all lesions enlarged with age, giving rise to various-sized lesions on the fruit at harvest. No fruiting structures were observed in the fruit lesions at the time of harvest. Fruit incubated in moist conditions developed excessive sporulation of a Colletotrichum species and no fruiting structures of fungi in the Mycosphaerellaceae could be detected.
Symptoms on leaves documented during field inspections were small (<2 mm in diameter) and visible on both sides of the leaf. These lesions were roundish and had a red-brown colour (Figure 1d). Lesions did not enlarge with age. Symptoms on leaves were most severe in July and August - at times, large areas of the leaves were covered with spots. In December to June, only a few spots per leaf were found. No further investigations were performed on these lesions.
Big, mature lesions on fruit yielded mostly C. gloeosporioides-like fungi (90%), identified based on iTs gene sequences, with only a few isolates resembling the Mycosphaerellaceae. Fewer Mycosphaerellaceae species were retrieved when lesion segments larger than 2 mm in diameter were used for isolations; however, with small (1-2 mm in diameter) lesions, 80-90% of lesion segments yielded isolates resembling fungi in the Mycosphaerellaceae.
Mycosphaerella-like cultures could be grouped into three morpho-groups based on colony morphology and colour. Cultures were, however, sterile, making it impossible to confirm their identity based on morphology. Colonies of Mycosphaerella group A cultures were characterised by thin, white, smooth margins (1 mm wide), sparse to moderate aerial mycelium and grey-olivaceous (21'''''i) surfaces on OA; whereas on MEA, the colony margins were evenly edged, aerial mycelium moderate,surfaces were olivaceous-grey (23'''''i) and the reverse was greenish black (33'''''k).
Colonies of Mycosphaerella group B cultures on OA were characterised by smooth margins, moderate aerial mycelium and grey-olivaceous (21""i) surfaces; whereas on MEA, the colony margins were irregular but smooth with moderate aerial mycelium and olivaceous-grey (23'''''i) surfaces with olivaceous-black (27'''''k) reverse sides.
Colonies of Mycosphaerella group C isolates on OA displayed thin, white, smooth margins (1-2 mm wide), sparse aerial mycelium and pale olivaceous-grey (21'''''d) surfaces. On MEA, the margins were smooth, aerial mycelium sparse and variable in colour, and surfaces were predominantly pale olivaceous-grey (23'''''d) with patches of olivaceous-grey (21'''''i) and smoke grey (19""d), and the reverse was dark olivaceous-grey (21'''''i).
DNA sequence comparisons
DNA sequencing and sequence comparisons
BLAST results of sequences for the ITS, LSU, BT and TEF gene regions generated for Mycosphaerella-like isolates from guava fruit in this study suggested that all our isolates represented species of Mycosphaerella (Table 1). Parsimony analyses of the sequences for the ITS and TEF gene regions were done to determine the phylogenetic placement of the Mycosphaerella-like isolates from guava amongst other Mycosphaerella species associated with Myrtaceae plants (Table 2) - the plant family to which Psidium guajava belongs. Alignment by inserting gaps resulted in a total of 1153 characters used in the combined data set for the I TS and TEF gene sequences. All parsimony-uninformative and constant characters were excluded, resulting in 394 parsimony-informative characters for the combined data set. Heuristic searches on the data set generated 100 most parsimonious trees and partition homogeneity tests indicated that the two data sets could be combined (Figure 2). The LSU and BT data sets were excluded from the combined data set because the LSU data set did not resolve the relationships amongst the species (data not shown) and the BT data available to include for reference strains are limited.
Two South African isolates from guava (group A) grouped with Mycosphaerella heimii Bouriquet ex Crous, supported by a 90% bootstrap value (Figure 2), while three isolates (group B) grouped with M. keniensis Crous & TA. Cout., although this grouping was only supported by a 74% bootstrap value. An additional five South African isolates (group C) grouped with M. acaciigena Crous & M.J. Wingf, supported by a 100% bootstrap value. PPRi 8914 clustered basal to the clade of M. ellipsoidea Crous & M.J. Wingf, M. africana Crous & M.J. Wingf, M. aurantia A. Maxwell and M. keniensis, while PPRI 8913 grouped basal to the M. acaciigena cluster, both supported by a 100% bootstrap value. Both of these isolates may represent new species. All these groupings support the MycoBank BLAST results.
Discussion
Pink spot of guava fruit in South Africa has in the past been attributed to C. gloeosporioides.6Mycosphaerella species were only recently suspected to be associated with the pink spot observed on guava fruit and leaves in the Lowveld of Mpumalanga, South Africa. In the current study we identified isolates from pink spot symptoms by means of phylogenetic comparisons, as opposed to morphology only. The analyses indicate that the Mycosphaerella isolates from fruit lesions represent at least three species not previously reported on guavas. These species are M. heimii, M. keniensis and M. acaciigena. Mycosphaerella heimii have previously been reported on Eucalyptus species and M. acaciigena on Acacia mangium Willd.20,31-33 A number of Mycosphaerella species are associated with Myrtaceae species other than Eucalyptus, including Mycosphaerella syzygii Crous, Mycosphaerella aequatoriensis Petr., Mycosphaerella eugeniae Rehm and Mycosphaerella metrosideri F. Stevens & PA. Young.34 None of these were isolated from the infected guava trees.
In this study, we applied DNA sequences of the ITS, LSU and TEF regions. Although ITS has been shown to be insufficient for the Mycosphaerella species in anamorph genera such as Cercospora and Septoria, it appears to be useful for distinguishing species with Pseudocercospora Speg., Ramularia Unger and most other Mycosphaerella anamorph genera.13 Our data of the ITS and TEF regions enabled us to distinguish between all the Mycosphaerella species included in this study, except for two isolates that grouped basal to existing clades. These isolates may represent new species. The BLAST results did, however, cluster them with individual species. The LSU data failed to resolve the phylogenetic relatedness amongst the species.
Species of Mycosphaerella are usually assumed to be host-specific and most species have a narrow host range.13,14 But, in some cases, Mycosphaerella species have been reported from other plants, albeit in low numbers.8 Crous and Groenewald13 suggested that in the absence of their natural host, Mycosphaerella species can infect and reproduce on non-host plants or catch crops in an attempt to find the appropriate host species. They referred to the phenomenon as the 'pogo stick hypothesis'. It is possible that guava serves as only a catch crop for the associated Mycosphaerella species identified in the present study. Another hypothesis explaining the presence of Mycosphaerella species on non-host plants is adaptation to a new host plant species.13
Most of the South African Mycosphaerella isolates associated with pink spot clustered with Mycosphaerella species from unresolved clades, as previously indicated by Crous et al.35 Further work is required to resolve the taxa associated with pink spot on guava in South Africa. In addition, the pathogenicity of Mycosphaerella isolates on guava needs to be resolved before conclusions on the host range of M. acaciigena, M. heimii and M. keniensis can be made. Characterisation of fungal isolates associated with leaf lesions observed during the 3-year period, also warrants further investigation. This report is the first of Mycosphaerella species associated with pink spot of guava in South Africa.
Authors' contributions
M.H.S. was the project leader. A.J. and M.T were responsible for experimental and project design and performed most of the experiments. All the authors contributed to the writing of the manuscript.
References
1. Morton JF. Fruits of warm climates. Miami, FL: Florida Flair Books; 1987. [ Links ]
2. Lim TK, Manicom BQ. Diseases of guava. In: Ploetz RC, editor. Diseases of tropical fruit crops. Wallingford: CABI Publications; 2003. p. 275-289. http://dx.doi.org/10.1079/9780851993904.0275 [ Links ]
3. Anon. Increased demand for guavas creates vital job opportunities. South African Vegetables and Fruit. 2009 Jun/Jul;19. [ Links ]
4. Schoeman MH. Diseases. In: De Villiers EA, Joubert PH, editors. The cultivation of guava. Nelspruit: ARC-Institute for Tropical and Subtropical Crops; 2004. p. 91-97. [ Links ]
5. Schoeman MH. Koejawelverwelksiekte [Guava wilt]. Afgriland. 2007;51:68-69. Afrikaans. [ Links ]
6. Matthee FN. Siektebestryding by vrugte 49: Koejawelverwelksiekte: Beter siektebeheer kan nuwe markte open [Disease eradication in fruit 49: Guava wilt: Better disease management will open new markets]. Landbouweekblad. 1981;1534:38-43. Afrikaans. [ Links ]
7. Barr ME. Preliminary studies on the Dothideales in temperate North America. Contributions of the University of Michigan Herbarium. Volume 9. Ann Arbor, MI: University of Michigan Herbarium; 1972. p. 523-638. [ Links ]
8. Crous PW. Mycosphaerella spp. and their anamorphs associated with leaf spot diseases of Eucalyptus. Mycologia Memoir. 1998;21:1-170. http://dx.doi.org/10.5598/imafungus.2011.02.01.08 [ Links ]
9. Crous PW, Tanaka K, Summerell BA, Groenewald Z. Additions to the Mycosphaerella complex. IMA Fungus. 2011;2:49-64. http://dx.doi.org/10.3114/sim.2009.64.02 [ Links ]
10. Crous PW, Schoch CL, Hyde KD, Wood AR, Gueidan C, De Hoog GS, et al. Phylogenetic lineages in the Capnodiales. Stud Mycol. 2009;64:17-47. [ Links ]
11. Park RF, Keane PJ. Leaf diseases of Eucalyptus associated with Mycosphaerella species. T Brit Mycol Soc. 1982;79:101-105. http://dx.doi.org/10.1016/S0007-1536(82)80195-2 [ Links ]
12. Crous PW, Wingfield MJ, Mansilla JP Alfenas AC, Groenewald JZ. Phylogenetic reassessment of Mycosphaerella spp. and their anamorphs occurring on Eucalyptus. II. Stud Mycol. 2006;55:99-131. http://dx.doi.org/10.3114/sim.55.1.99 [ Links ]
13. Crous PW, Groenewald JZ. Hosts, species and genotypes: Opinions versus data. Australas Plant Path. 2005;34:463-470. http://dx.doi.org/10.1071/AP05082 [ Links ]
14. Crous PW. Adhering to good cultural practice (GCP). Mycol Res. 2002;106:1378-1379. http://dx.doi.org/10.1017/S0953756202227136 [ Links ]
15. Carnegie AJ, Ades PK, Ford R. The use of RAPD-PCR analysis for the differentiation of Mycosphaerella species from Eucalyptus in Australia. Mycol Res. 2001;105:1313-1320. http://dx.doi.org/10.1017/S0953756201004762 [ Links ]
16. Maxwell A, Jackson SL, Dell B, Hardy E St J. PCR-identification of Mycosphaerella species associated with leaf diseases of Eucalyptus. Mycol Res. 2005;109:992-1004. http://dx.doi.org/10.1017/S0953756205003539 [ Links ]
17. Crous PW, Hong L, Wingfield MJ, Wingfield BD, Kang J. Uwebraunia and Dissoconium, two morphologically similar anamorph genera with distinct teleomorph affinity. Sydowia. 1999;51:155-166. [ Links ]
18. Crous PW, Kang JC, Braun U. A phylogenetic redefinition of anamorph genera in Mycosphaerella based on ITS rDNA sequence and morphology. Mycologia. 2001;93:1081-1101. http://dx.doi.org/10.2307/3761670 [ Links ]
19. Crous PW, Groenewald JZ, Mansilla JP Hunter GC, Wingfield MJ. Phylogenetic analysis of Mycosphaerella spp. and their anamorphs occurring on Eucalyptus. Stud Mycol. 2004;50:195-214. [ Links ]
20. Hunter GC, Wingfield BD, Crous PW, Wingfield MJ. A multigene phylogeny for species of Mycosphaerella occurring on Eucalyptus leaves. Stud Mycol. 2006;55:147-161. http://dx.doi.org/10.3114/sim.55.1.147 [ Links ]
21. Dhingra OD, Sinclair JB. Basic plant pathology methods. London: CRC Press; 1995. [ Links ]
22. Rayner RW. A mycological colour chart. Kew: CMI and British Mycological Society; 1970. [ Links ]
23. White TJ, Bruns T, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis AM, Gelfand DH, Snisky JJ, White JW, editors. A guide to molecular methods and applications. New York: Academic Press; 1990. p. 315-322. [ Links ]
24. Moncalvo JM, Wamg HH, Hseu RS. Phylogenetic relationships in Ganoderma inferred from the internal transcribed spacers and 25S ribosomal DNA sequences. Mycologia. 1995;87:223-238. http://dx.doi.org/10.2307/3760908 [ Links ]
25. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol. 1990;172:4238-4246. [ Links ]
26. Aveskamp MM, Verkley GJM, De Gruyter J, Murace MA, Perelló A, Woudenberg JHC, et al. DNA phylogeny reveals polyphyly of Phoma section Peyronellaea and multiple taxonomic novelties. Mycologia. 2009;101:363-382. http://dx.doi.org/10.3852/08-199 [ Links ]
27. Carbone I, Kohn LM. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia. 1999;91:553-556. http://dx.doi.org/10.2307/3761358 [ Links ]
28. Katoh K, Kuma K, Toh H, Miyata T. MAF6FT version 5: Improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 2005;33:511-518. http://dx.doi.org/10.1093/nar/gki198 [ Links ]
29. Swofford DL. PAUP*: Phylogenetic analysis using parsimony (*and other methods). Version 4.0b10. Sunderland, MA: Sinauer Associates; 2002. [ Links ]
30. Farris JS, Kallersj M, Kluge AG, Bult C. Testing significance of congruence. Cladistics. 1994;10:315-320. http://dx.doi.org/10.1111/j.1096-0031.1994.tb00181.x [ Links ]
31. Crous PW, Groenewald JZ, Pongpanich K, Himaman W, Arzanlou M, Wingfield MJ. Cryptic speciation and host specificity among Mycosphaerella spp. occurring on Australian Acacia species grown as exotics in the tropics. Stud Mycol. 2004;50:457-469. [ Links ]
32. Maxwell A, Dell B, Neumeister-Kemp HG, Hardy GE St J. Mycosphaerella species associated with Eucalyptus in south-western Australia: New species, new records and a key. Mycol Res. 2003;107:351-359. http://dx.doi.org/10.1017/S0953756203007354 [ Links ]
33. Crous PW, Swart WJ. Foliicolous fungi of Eucalyptus spp. from eastern Madagascar: Implications for South Africa. S Afr For J. 1995;172:1-5. http://dx.doi.org/10.1017/S0953756298007692 [ Links ]
34. Crous PW. Species of Mycosphaerella and related anamorphs occurring on Myrtaceae (excluding Eucalyptus). Mycol Res. 1999;103:607-621. [ Links ]
35. Crous PW, Summerell BA, Carnegie AJ, Wingfield MJ, Hunter GC, Burgess TI, et al. Unravelling Mycosphaerella: Do you believe in genera? Persoonia. 2009;23:99-118. http://dx.doi.org/10.3767/003158509X479487 [ Links ]
 Correspondence:
Correspondence:
Adriaana Jacobs
Biosystematics Programme, Plant Protection Research Institute
Agricultural Research Council
Private Bag X134, Queenswood 0121, South Africa
EMAIL: JacobsR@arc.agric.za
Received: 18 Feb. 2013
Revised: 04 Feb. 2014
Accepted: 07 Feb. 2014