Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Science
On-line version ISSN 1996-7489
Print version ISSN 0038-2353
S. Afr. j. sci. vol.109 n.11-12 Pretoria Jan. 2013
RESEARCH ARATICLES
Molecular characterisation of Mycoplasma gallisepticum genotypes from chickens in Zimbabwe and South Africa
Serena A. Moretti; Charlotte E. Boucher; Robert R. Bragg
Department of Microbial, Biochemical and Food Biotechnology, University of the Free State, Bloemfontein, South Africa
ABSTRACT
Mycoplasma gallisepticum is an economically important pathogen of poultry worldwide. Yet the characterisation of M. gallisepticum field strains present in southern Africa has not previously been reported. We characterised various M. gallisepticum genotypes within the region and highlight the unique differences between two genotypes found in South Africa and Zimbabwe. PCR targeting a partial region of the mgc2 gene was used to screen various poultry farms in South Africa and Zimbabwe for M. gallisepticum. Samples were characterised using multilocus gene-targeted sequencing. Portions of the surface protein encoding pvpA, gapA and mgc2 genes and the uncharacterised surface lipoprotein gene designated MGA_0319 were sequenced and analysed. Nucleotide sequences were compared to vaccine and reference strains as well as to strains from different countries. The South African genotype contained unique mgc2 and pvpA gene regions, while the Zimbabwean genotype proved to be even more distinct with unique gapA, mgc2 and pvpA gene regions. In addition, BLAST results showed high similarities in the partial mgc2 gene region between the South African and Zimbabwean genotypes and the 'atypical' Israeli RV-2 strain, suggesting a link in its epidemiology. These results also allow for improved control strategies for southern Africa, and the use of more effective vaccine strains.
Keywords: Mycoplasma; poultry; genotypes; southern Africa
Introduction
Mycoplasma gallisepticum (MG) causes chronic respiratory disease in chickens and infectious sinusitis in turkeys.1 In breeders and layers, the disease causes a drop in egg production and an increase in embryo mortality2 with symptoms resulting in considerable economic losses. MG is transmitted vertically through infected eggs and horizontally by inhalation of contaminated dust, airborne droplets and feathers.3 In recent years, a re-emergence of MG infection has been observed in poultry, possibly as a result of the concentration of large poultry populations in small geographic areas and under poor biosecurity.4 The Southern African Poultry Association5 announced that the gross poultry farm income for 2010 was R22 940 billion - the largest segment of South African agriculture at 23% of all agriculture production. With an already large and continually expanding poultry industry in southern Africa, efficient methods of biosecurity are required for the control of possible diseases, like chronic respiratory disease, caused by MG.
Control of MG has largely been based on the eradication of the organism from breeder flocks and their progeny, with confirmation of Mycoplasma-free status of the flocks by periodic serological monitoring, following culture and/or PCR monitoring.6 In some countries where complete eradication is implausible, live vaccines have been utilised as an alternative control strategy.7,8 Because of the increased use of vaccination, methods with greater discriminatory power are needed to differentiate vaccine strains from circulating field isolates. The most widely used method for the differentiation of strains and to track epidemiology-related isolates in the field has been random amplified polymorphic DNA (RAPD). However, this technique has intrinsic problems with reproducibility and has not allowed for inter-laboratory comparisons.9 In addition, RAPD analysis is only feasible for laboratories which specialise in Mycoplasma research and diagnosis because the assay relies on the fastidious cultivation of sample strains together with that of a wide array of control isolates.
More recently, a multi-locus sequence typing method referred to as gene-targeted sequencing (GTS) has been used to identify and differentiate MG strains.10 GTS is believed to be a better method than RAPD because of its improved reproducibility and because it allows rapid global comparisons among laboratories. In this study, GTS was used to target three surface protein genes - gapA, mgc2 and pvpA - and a predicted surface lipoprotein designated MGA_0319. A total of 67 MG field genotypes from the USA, Israel and Australia, and 10 reference strains were characterised in the study.10 The analysis of these four surface-protein genes combined showed a better discriminatory power than RAPD analysis, thus providing an improved typing method for MG strains available to a wider range of laboratories.
Although MG has been well characterised in some countries, very little information has been documented on the southern African genotypes. This deficiency in information has led to poor biosecurity in the poultry industry. Based on the characterisation of the various MG genotypes, we aim to highlight the differences between the genotypes found in South Africa and Zimbabwe. This report is believed to be the first on the use of GTS on strains from South Africa and Zimbabwe.
Materials and methods
Samples were taken during July 2010 to October 2011 from chicken farms in South Africa and Zimbabwe where MG infection was suspected based on serology and clinical signs. Swabs were taken from the choanal cleft, oropharynx, oesophagus or trachea of the chickens and were analysed individually (i.e. not pooled). Samples collected from Zimbabwe were taken from live chickens, while those collected from South Africa were isolated from layers post mortem. The Zimbabwean flock had been formerly vaccinated with an inactive vaccine produced from the MG Rlow strain (AE015450.2). Despite vaccination, the flock succumbed to clinical symptoms of MG infection shortly after and was treated with the antibiotics tylosin and tiamulin (Denagard®) a week prior to sampling. The South African flock showed symptoms of head shaking and cyanosis before death.
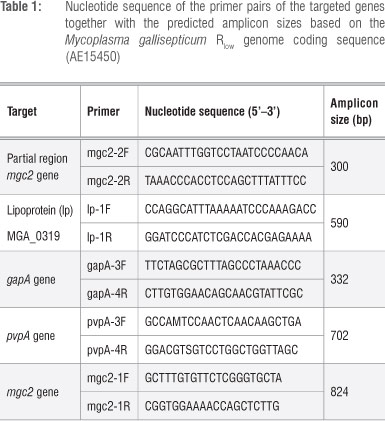
Genomic DNA was extracted from individual swabs using the QIAamp DNA Mini Kit (QIAGEN, Düsseldorf, Germany) according to the manufacturer's recommendations. Samples were screened for MG using PCR targeting a partial region (~300 bp) of the mgc2 gene11 previously described to be highly sensitive and specific.12 Each PCR reaction consisted of 5 µL. DNA template, 10 mM dNTPs, 0.5 µM of each appropriate primer pair (as listed in Table 1), 1.5 units of Taq polymerase (New England Biolabs, Ipswich, MA, USA), 1x Thermopol buffer (New England Biolabs) and sterile nanopure water to the volume of 50 µL. All amplifications were performed in an Applied Biosystems Thermal Cycler 2720 (Life Technologies Corporation, Carlsbad, CA, USA). Amplification products were separated by electrophoresis at 4.5 V/cm on a 1% (w/v) agarose (Lonza, Basel, Switzerland) gel in TAE buffer with 0.2 µg/mL ethidium bromide (Sigma-Aldrich chemie GmbH, Steinheim, Germany) and visualised under UV light.
MG-positive samples were further characterised by targeting the gapA, lipoprotein (MGA_0319), mgc2 and pvpA partial gene sequences as previously described.10 PCR products were excised with a sterile scalpel and cleaned by means of the illustra GFX™ PCR DNA and Gel Band Purification Kit (GE Healthcare, Buckinghamshire, UK). Each purified amplification product was sequenced in both directions using the BigDye Terminator V3.1 Cycle Sequencing Kit (Applied Biosystems). Sequencing was performed with the capillary sequencer 3130xl ABI Genetic Analyser (Applied Biosystems) at the University of the Free State, Department of Microbial, Biochemical and Food Biotechnology. Complete overlapping of complementary sequences, editing and consensus construction was performed using Geneious Pro v5.4.4.13 All consensus sequences were trimmed so as to start and end at an equivalent coding sequence position as determined by sequences previously submitted.10
A total of 39 MG genotypes was used for comparative analysis; all of the RAPD types previously described were included.10 The total number of genotypes included 23 from the USA, 8 from Israel, 4 from Australia, 3 vaccine strains (6/85, ts-11 and F-CK58) and the reference R genotype. All sequence data was taken from GenBank.10 Alignments of individual gene sequences were constructed by the CLUSTAL V method with a gap penalty of 10 using the MEGALIGN program (in LASERGENE; DNASTAR).
From this sequence, identity was reported. So as not to exclude any similar strains not included in the comparative analysis, sequence similarities from the Zimbabwean and South African genotypes were also detected using nucleotide-nucleotide BLAST.14
Sequences of the two genotypes of interest (those from South Africa and Zimbabwe) were submitted to GenBank under the following accession numbers: MGA_0319, KC130902, KC130906, gapA, KC130901, KC130905, mgc2, KC130903, KC130907, pvpA, KC130904 and KC130908.
Results
Poultry flocks were screened for MG by using DNA extracted directly from individual swab samples for PCR of mgc2. Visualisation of 237-303-bp amplicons on an agarose gel confirmed the presence of MG. Of the 30 swabs taken from the Zimbabwean poultry farm, 11 were positive for MG, while 5 samples taken from the South African poultry farm were positive for MG. The samples were further genetically characterised by targeting and sequencing a larger region of the above-mentioned mgc2 gene together with the gapA, lipoprotein (MGA_0319) and pvpA genes. CLUSTAL V alignments of these individual genes were carried out against reference genotypes and genotypes from various countries, representing all the RAPD groups previously described.10 The results are tabulated in Table 2.
The discriminatory power hierarchy, from lowest to highest, for GTS analysis of the individual genes is ranked as: gapA, lipoprotein MGA_0319, mgc2 and pvpA.10 This hierarchy is evident in the matched identity of the gapA sequence of the South African genotype to numerous MG genotypes, including the ts-11 and 6/85 vaccine strains and some among those from the USA and Australia. On the contrary, the South African lipoprotein portion (MGA_0319) diverged from the previous vaccine strains and showed sequence identity to those among the genotypes from the USA and Australia. The mgc2 gene region further discriminated the South African genotype, showing closest identity at only 96% to an Australian MG strain.
The sequence data for the South African and the Zimbabwean genotypes differed from each other, as already seen with the partial mgc2 gene region in Figure 1. The pvpA gene region is known to be the most discriminatory of the targeted genes and showed 95% identity between these two genotypes. Nonetheless, in terms of the mgc2 gene region (ranked second with highest discriminatory power), the Zimbabwean genotype remained unique and showed the highest identity percentage (96%) to the 6/85 vaccine strain. In addition, the gapA gene was unique for the Zimbabwean genotype. This finding was unexpected because this gene is known to show the least sequence variation of the targeted genes. On the contrary, the lipoprotein (MGA_0319) region where the Zimbabwean genotype showed 100% identity to both the 6/85 and K5111ATK01 US strains, belonged to the RAPD type group M.
Other genotypes showing high sequence similarity using nucleotide-nucleotide BLAST, but not included in the analysis, were the RV-2 and TLS-2 Israeli strains. Their exclusion from this study was based on inadequate sequence data in GenBank for the various target genes, and also because their discovery succeeded the RAPD type classification.10 The partial region of the mgc2 gene of both the South African and Zimbabwean genotypes showed 100% BLAST identity to that of the atypical Israeli RV-2 genotype (EU939449.1); however, the Zimbabwean genotype included an additional in-frame insert of 18 nucleotide base pairs. The partial mgc2 gene regions of the genotypes were aligned using CLUSTAL15 and the reference MG R (AE015450.2) sequence was included for comparison (Figure 1). The Zimbabwean genotype continued to show high similarity between the available sequence data of the MGA_319 lipoprotein and gapA gene regions. In addition, both South African and Zimbabwean genotypes showed high sequence similarities in the pvpA gene region, with the exception of a unique and significant nucleotide region deletion present within the RV-2 strain. Thus, the pvpA gene regions of the Zimbabwean and South African genotypes remained unique.
Discussion
Few studies have reported on the characterisation of MG field strains present in southern Africa. Poultry farms from two geographical regions - Zimbabwe and South Africa - suspected of MG infection were screened for novel MG genotypes using molecular techniques. MG-positive samples were genetically characterised according to analysis of the gapA, lipoprotein MGA_0319, mgc2 and pvpA gene regions. Of the 39 MG genotypes included in this study, spanning all of the RAPD types previously described,10 the South African genotype contained unique mgc2 and pvpA gene regions. The Zimbabwean genotype was shown to be even more divergent with unique gapA, mgcA and pvpA regions representing a distinct genotype.
The Israeli RV-2 strain was not included in this comparative study but showed high similarity to the partial regions of the mgc2 gene of the South African strain. The RV-2 strain was originally isolated from two outbreaks in broiler breeders in 2007 in Israel.16 It was noted as being distinct from the 'Israeli'-type strain previously described10 and suggested to be a new MG strain introduced to Israel. In addition, the strain also showed susceptibility to enrofloxacin (0.05 µg/µL), unlike other strains isolated in Israel at the time.17 The mgc2 gene region of the Zimbabwean genotype also showed high similarity to the RV-2 strain; however, the Zimbabwean genotype contained an additional insert of 18 nucleotide bases. Nevertheless, a link between the atypical strain found in Israel and those prevalent in southern Africa has been established.
It is also important to note the possibility of multiple MG genotypes or species causing infection in the same poultry flock or even the same bird.18 This possibility could pose a foreseeable predicament for GTS when analysing various target regions of a sample. Although cloning of the PCR fragments would eliminate mixed sequencing data, the various targeted regions would not be able to be correlated to a specific genotype, especially when dealing with novel genotypes. With this in mind, pooling of samples from the same poultry farms should be avoided as a precautionary measure. Fortunately, no mixed sequencing data was seen for the gapA, lipoprotein MGA_0319, mgc2 and pvpA gene regions for the two genotypes of interest. In addition, different samples collected from the respective poultry farms also showed identical sequence data for the various regions.
The molecular characterisation of MG field genotypes in southern Africa has allowed for a better understanding of the diversity and epidemiology of the poultry pathogen worldwide. The use of GTS has also allowed the characterisation of MG genotypes without the need for cultivation and a wide array of control isolates otherwise found in more Mycoplasma-specialised laboratories. In addition, the novel genotypes may help explain the failed vaccinations reported throughout the southern African poultry industry. Thus, the identification of novel genotypes present in the different geographical areas has implications for improved control strategies of the disease. Future work would therefore include investigations into whether certain nucleotide changes present in southern African MG field genotypes have resulted in the altering of the antigenic profile of MG which could result in the effective avoidance of immune recognition and antibodies produced as a result of vaccination.
Acknowledgements
We acknowledge the financial assistance of the National Research Foundation (NRF); opinions expressed and conclusions arrived at are those of the authors and are not necessarily to be attributed to the NRF. We also thank Natalie Armour from the Poultry Diagnostic and Research Center (University of Georgia) for her help with analysis of the mgc2, lp and gapA gene regions and for access to their MG sequence database.
Authors' contributions
S.A.M. performed the experiments and wrote the manuscript under the supervision of C.E.B. and R.R.B.
References
1. Ley DH, Yoder HW. Mycoplasma gallisepticum infection. In: Calnek BW, editor. Diseases of poultry. Ames, iA: Iowa State University Press; 1997. p. 194-207. [ Links ]
2. Ley DH. Mycoplasma gallisepticum infection. In: Saif YM, Barnes HJ, Glisson JR, Fadly AM, McDougald LR, Swayne DE. Diseases of poultry. 11th ed. Ames, IA: Iowa State University Press; 2003. p. 722-744. [ Links ]
3. McMartin DA, Khan MI, Christie G. Delineation of the lateral spread of Mycoplasma gallisepticum infection in chickens. Avian Dis. 1987;31(4):814-819. http://dx.doi.org/10.2307/1591037 [ Links ]
4. Liu T, Garcia M, Levisohn S, Yogev D, Kleven SH. Molecular variability of the adhesin-encoding gene pvpA among Mycoplasma gallisepticum strains and its application in diagnosis. J Clin Microbiol. 2001;39(5):1882-1888. http://dx.doi.org/10.1128/JCM.39.5.1882-1888.2001 [ Links ]
5. Southern African Poultry Association. Annual statistical reports - SAPA industry profile 2010: Animal health disease [homepage on the Internet]. No date [cited 2011 Aug 21]. Available from: http://www.sapoultry.co.za/industry_profile.php] [ Links ]
6. Levisohn S, Kleven SH. Avian mycoplasmosis (Mycoplasma gallisepticum). Rev Sci Tech Off Int Epizoot. 2000;19:425-442. [ Links ]
7. Kleven SH. Changing expectations in the control of Mycoplasma gallisepticum. Acta Vet Hung. 1997;45(3):299-305. [ Links ]
8. Whithear KG. Control of avian mycoplasmoses by vaccination. Rev Sci Tech. 1996;15(4):1527-1553. [ Links ]
9. Tyler KD, Wang G, Tyler SD, Johnson WM. Factors affecting the reliability and reproducibility of amplification based DNA fingerprinting of representative bacterial pathogens. J Clin Microbiol. 1997;35(2):339-346. [ Links ]
10. Ferguson NM, Hepp D, Sun S, Ikuta N, Levisohn S, Kleven SH, et al. Use of molecular diversity of Mycoplasma gallisepicum by gene-targeted sequencing (GTS) and random amplified polymorphic DNA (RAPD) analysis for epidemiological studies. Microbiology. 2005;151(6):1883-1893. http://dx.doi.org/10.1099/mic.0.27642-0 [ Links ]
11. Hnatow LL, Keeler CL, Tessmer LL, Czymmek K, Dohms JE. Characterization of MGC2, a Mycoplasma gallisepticum cytadhesin with homology to the Mycoplsma pneumonia 30-kilodalton protein P30 and Mycoplasma genitalium P32. Infect Immun. 1998/66(7):3426-3442. [ Links ]
12. Garcia M, Ikuta N, Levisohn S, Kleven SH. Evaluation and comparison of various PCR methods for detection of Mycoplasma gallisepticum infection in chickens. Avian Dis. 2005;49:125-132. http://dx.doi.org/10.1637/7261-0812204R1 [ Links ]
13. Drummond AJ, Ashton B, Buxton S, Cheung M, Cooper A, Duran C, et al. Geneious v5.4. Auckland: Geneious; 2011. Available from: http://www.geneious.com [ Links ]
14. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403-410. [ Links ]
15. Larkin MA, Blackshields G, Brown NP Chenna R, McGettigan PA, McWilliam H, et al. ClustalW and ClustalX version 2.0. Bioinformatics. 2007;23(21):2947-2948. http://dx.doi.org/10.1093/bioinformatics/btm404 [ Links ]
16. Lysnyansky I, Gerchman I, Perk S, Levisohn S. Molecular characterization and typing of enrofloxacin-resistant clinical isolates of Mycoplasma gallisepticum. Avian Dis. 2008;52(4):685-689. http://dx.doi.org/10.1637/8386-063008-RESNOTE.1 [ Links ]
17. Gerchman I, Lysnyansky I, Perk S, Levisohn S. In vitro susceptibilities to fluoroquinolones in current and archieved Mycoplasma gallisepticum and Mycoplasma synoviae isolates from meat-type turkeys. Vet Microbiol. 2008;131(3-4):266-276. http://dx.doi.org/10.1016/j.vetmic.2008.04.006 [ Links ]
18. Mallinson ET, Rosenstein M. Clinical, cultural, and serological observations of avian mycoplasmosis in two chicken breeder flocks. Avian Dis. 1976;20:211-215. http://dx.doi.org/10.2307/1589495 [ Links ]
 Correspondence:
Correspondence:
Robert Bragg
Department of Microbial, Biochemical and Food Biotechnology
University of the Free State
PO Box 339
Bloemfontein 9300, South Africa
Email: braggRR@ufs.ac.za
Received: 23 Apr. 2013
Revised: 11 Jun. 2013
Accepted: 25 Jun. 2013














