Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Science
On-line version ISSN 1996-7489
Print version ISSN 0038-2353
S. Afr. j. sci. vol.107 n.9-10 Pretoria Oct. 2011
RESEARCH ARTICLE
Caricain: a basis for enzyme therapy for coeliac disease
Hugh J. CornellI; Teodor StelmasiakII
ISchool of Applied Science, RMIT University, Melbourne, Australia
IIGlutagen Pty Ltd, Maribyrnong, Victoria, Australia
Gliadin, a glycoprotein present in wheat and other grass cereals, is a causative agent in coeliac disease. It is therefore important to find methods for the detoxification of gliadin. Lysosomal integrity is lost in patients with active coeliac disease but restored when gliadin is removed from the diet. We employed a rat liver lysosome assay to monitor the extent of detoxification of a gliadin digest by caricain, a protein enzyme found in papaya. Pre-incubating the gliadin digest for different durations with caricain allowed the kinetics of the detoxification process to be studied. A significant degree of protection (80%) of the lysosomes was achieved with 1.7% w/w of caricain on substrate after incubation for 2 h at 37ºC. The detoxification followed first-order kinetics with a rate constant of 1.7 x 10-4/s. The enzyme was strongly inhibited by imidazole, but weakly by phenylmethyl sulphonyl fluoride, as was also a caricain-enriched fraction from ion-exchange chromatography of papaya oleo-resin. The value of caricain in the detoxification of gliadin was confirmed in the present studies and this enzyme shows promise for enzyme therapy in coeliac disease.
Introduction
The use of enzyme therapy in coeliac disease depends upon choosing the most appropriate enzyme for detoxification of gluten-type proteins and understanding the way in which this enzyme can be evaluated.1 We took into account the pathogenic mechanisms operating in predisposed individuals and suggest that more complete digestion of gliadin peptides is essential in order to limit the concentration of specific immunogenic peptides that can trigger damage to tissue. This work began with the use of peptic-tryptic-pancreatic digests of gliadin, representing a source of the most toxic peptides in wheat,2 and then focused on the toxicity of specific peptides in A-gliadin. This focus was made possible by the use of synthetic peptides within the A-gliadin sequence3 and their evaluation using the foetal chick assay.4
The potential value of enzyme therapy was shown in a clinical trial5 with 21 biopsy-proven volunteers with coeliac disease. The volunteers' coeliac disease was in remission and they were on a gluten-free diet.5 In this double-blind crossover trial using a porcine intestinal extract, it was demonstrated that in those patients who developed symptoms as a result of a 2-week gluten challenge, the symptom scores during enzyme therapy were significantly lower than those for the period on placebo (p < 0.02). Furthermore, the need for enzyme therapy was seen in five out of six patients who had some small bowel abnormalities shown by histology at the start of the trial. There was evidence of improvement in villous architecture after enzyme therapy in three of these patients, even though they received a gluten challenge.5
More recently, screening of plant enzymes was carried out using a rat liver lysosome assay as a means of evaluating the detoxification process.6 This work culminated in the use of crude papaya latex as a source of proteolytic enzymes that could be considered for enzyme therapy. Crude papaya latex contains several other thiol hydrolases besides papain, such as chymopapapain, glycyl endopeptidase and caricain,7 as well as glutamine cyclotransferase.8 With the use of ionexchange chromatography, size exclusion high-performance liquid chromatography and mass fragmentation analysis, it was indicated that caricain was the most effective detoxifying enzyme present in crude papaya latex.6 In the present work, we used a rat liver lysosomal assay to test the detoxifying activity of highly purified caricain.9
The lysosomal assay gave us the ability to measure the detoxification of gliadin by the corresponding reduction in the degree of disruption of rat liver lysosomes that occurred after incubation of the gliadin digest with the caricain. Riecken et al.10 are credited with being the first group to study the effects of gluten on lysosomes in the small intestine. They showed that these organelles were disrupted in patients with acute coeliac disease, but that they regained their integrity after the patients were placed on a gluten-free diet. We found that the same disruption occurred in vitro with rat liver lysosomes when treated with peptic-tryptic-pancreatic digests of gliadin. This finding became the basis for testing the detoxifying effects of enzyme extracts.6 As a turbidimetric assay, the rat liver lysosome assay is simple and rapid and the degree of disruption of the lysosomes was similar to that observed by Riecken at al.10 Disruption can be seen under the microscope. Furthermore the results of this assay correlated well with those of the foetal chick assay in studies of coeliac disease.4 The effects of enzyme concentration and time of incubation were able to be studied, giving an insight into the kinetics of the reaction under physiological conditions. This ability is important for indicating the effectiveness of enzyme therapy using a tablet of concentrated enzyme to be administered before a meal containing gluten. In enzymatic reactions, the enzyme-substrate (ES) complex is formed in a fast step, followed by the decomposition of the complex to give the product in a rate-controlled step. That is:


The rate of the forward reaction = k2 [ES]. In other words the rate is proportional to the concentration of the complex and would be expected to be first order.11
In these experiments, we attempted to determine if the reaction was first order by following the disappearance of the toxic gliadin peptides over time using the rat liver lysosome assay at a fixed level of enzyme.12 It was also necessary to check the percentage protection (P) by using different levels of enzyme over a standard time (2 h) which approximates physiological residence time in the small intestine. It was important to determine if highly purified caricain offered high protection as this could be relevant to its use in therapy for coeliac disease.
Materials and methods
Protection of lysosomes by enzymes
Toxic peptides, present in high concentration at the start of the reaction with the enzyme, cause disruption of lysosomes on incubation, whereas non-toxic peptides formed after preincubation with the active enzyme do not. A corresponding reduction in absorbance (measured at 410 nm) after incubation with the lysosomes is thereby observed.
The more effective the enzyme, the higher will be the absorbance at 410 nm (Abs410) after a given pre-incubation period with the gliadin digest followed by the 1.5 h incubation with the lysosomes. This amount of protection (P) offered by the enzyme is able to be used to follow the disappearance of the toxic peptides and is calculated by:

where R o equals the reduction in absorbance (%) without the enzyme and RE equals the reduction in absorption (%) with the enzyme. The values of P can be used to measure the extent of the reaction: a low P indicates ineffective digestion and a high P indicates effective digestion of toxic products.6
Enzyme assays
Evaluation of enzyme activity was carried out using crude papaya latex (Enzyme Solutions Pty Ltd, Melbourne, Australia) and purified caricain, generously supplied by Professor Yvan Looze (Free University of Brussels, Brussels, Belgium). Solutions of the crude papaya latex in phosphate buffered saline (PBS) were prepared at concentrations of 2.5 mg/mL, 5 mg/mL, 10 mg/mL and 20 mg/mL. Duplicate 0.20 mL samples were pre-incubated with 0.10 mL of a peptictryptic-pancreatic digest of wheat gliadin13 (50 mg/mL in 0.15 mol/L NaCl) for 2 h at 37 ºC as described previously.6 A suspension of rat liver lysosomes in PBS (0.10 mL) was then added and the mixtures incubated again at 37 ºC for 1.5 h. The concentration of the lysosome suspension was adjusted so that 0.1 mL diluted to 3.0 mL with PBS gave an absorbance value of 0.9 - 1.0 at 410 nm. After this time, PBS (3.0 mL) was then added and the absorbance at 410 nm measured in a spectrophotometer. Blank experiments without the incubation (zero-time controls) were also performed, as were controls without any inclusion of enzyme with the gliadin digest during reaction with the lysosomes. P values were calculated for the crude papaya latex at each concentration of the enzyme. An additional two determinations were carried out at 10 mg/mL of crude enzyme to enable standard errors to be calculated. Specific activity was obtained by dividing the P value by the mass of the sample (mg) in the assay tube.
The purified caricain sample was supplied as a solution of S-methyl thio derivative in 2 mol/L ammonium sulphate and required dialysis against water and PBS, followed by activation with dithiothreitol (2.5 mmol/L) in the presence of ethylenediaminetetraacetic acid (EDTA, 5 mmol/L) for 1 h at room temperature (Set 1). In another experiment, activation was carried out with 5 mmol/L dithiothreitol in the presence of 5 mmol/L EDTA for 2 h (Set 2). In Set 1, dilution of 0.6 mL of the original sample (1.8 mg/mL) to 3.0 mL with PBS resulted in a concentration of 0.36 mg/mL in the activated test sample and this solution was further diluted to one-tenth and one-hundredth of the solution for preliminary tests in the lysosome assay.
In Set 2, 0.6 mL of the original sample was diluted to 2.5 mL, giving 0.430 mg/mL and this solution was diluted to a half, one-fifth and one-tenth for further tests. The percentage protection (P) was calculated for each concentration and specific activity obtained, as before, by dividing the P value by the mass (in mg) in the assay tube.
Enzyme kinetic studies
The effect of incubation time was used to determine if the reaction was first or second order.11 First-order reactions yield a straight line from plots of  against time (t), substituting P for x and making a0 equal 100 (initial concentration of toxic peptides). Tests for second-order reactions were likewise carried out by plotting
against time (t), substituting P for x and making a0 equal 100 (initial concentration of toxic peptides). Tests for second-order reactions were likewise carried out by plotting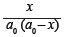 against t, where again a0 equalled 100.
against t, where again a0 equalled 100.
Incubation times of up to 120 min were employed. Rate constants (k) for the reactions were calculated for linear portions of the graphs by measurement of the gradients as follows: k = 2.303 x gradient, calculated in /s.
Experiments were carried out using the highly purified caricain, activated as in Set 2, at a concentration of 0.215 mg/mL. As supplies of the purified caricain were limited, statistical treatment of the results was not possible. However, reference to Table 1 shows that standard errors of the means for the assay, when crude enzyme and fractions thereof were used, represent only about 1% - 2% of the mean result.
Enzyme inhibition studies
Inhibition studies using active fractions from CM Sephadex chromatography of papaya latex was seen as another way of characterising the major enzymes in these fractions. The lysosomal assay was used as a way of obtaining information which might indicate whether glutamine cyclotransferase, inhibited by 1 mmol/L imidazole,14 or serine proteases, often inhibited by phenylmethyl sulphonyl fluoride (PMSF) were present. Lysosome assays6 were carried out in the normal way with 2.0 mmol/L and 6.0 mmol/L concentrations of the inhibitors and the results compared against controls without any inhibitor. The inhibition was calculated from the protection index (P) as follows:
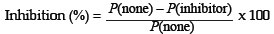
Tests were carried out on the highly purified caricain, activated as in Set 2, at a concentration of 0.215 mg/mL. Additional determinations were carried out on Fractions 3 and 4 from previous work6 in order to verify previous findings and to estimate standard errors. There was insufficient purified caricain available for estimations of standard error, so the crude fractions were used for this purpose.
Statistical analysis
Means ± s.e.m. (n = 4) were calculated where applicable.
Results and discussion
Protection offered by enzymes
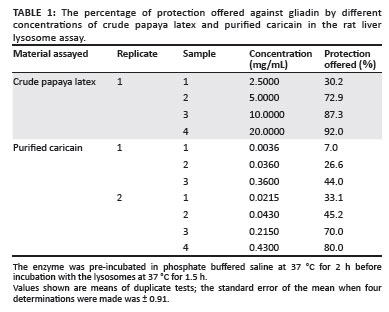
The results of protection offered by different concentrations of papaya latex and purified caricain in the rat liver lysosome assay are shown in Table 1. The protection offered by crude papaya latex was not useful at the 2.5 mg/mL concentration of the enzyme, but at a concentration of 10 mg/mL and above, high levels of protection were obtained after 2-h incubations with the gliadin digest. The reactivation of purified caricain did not appear to be complete in the Set 1 results and a longer time of reaction with dithiothreitol at 5 mmol/L (as in Set 2), produced a more active enzyme. The caricain in Set 2 gave a high degree of protection (80%) at a concentration of only 0.43 mg/mL (1.7% w/w on gliadin substrate). At a slightly lower concentration of the enzyme, P was 44% for the Set 1 result. Specific activity of the purified caricain at 0.215 mg/mL was 7674, compared with the specific activity of 60.4 for crude papaya latex at a similar P value. This indicates a 127fold increase in purification from crude papaya to purified caricain.
It was evident that the protective activity of caricain in the rat liver lysosome assay was detectable at a concentration of 3.6 µg/mL. There was some degree of linearity for a plot of P versus enzyme concentration at low enzyme concentrations (Table 1). Comparing a P value of 70.0% at 0.215 mg/mL for caricain with that of 72.9% at 5 mg/mL for crude papaya latex, indicated that the crude papaya extract contained about 5% - 10% caricain.
Enzyme kinetic studies
We can assume that the first-order kinetics applies to the decomposition of the ES complex because the gliadin digest (substrate) is the material influencing the absorbance figures measured in the assay. The lysosome assay is thus a valuable means of monitoring the disappearance of toxic gliadin peptides during the incubation period, making it useful for further kinetic studies. The results in Table 2 and Figure 1 confirm that the reaction followed first-order kinetics as the degree of linearity of the plot of  against t was high (r2 = 0.9975), with a gradient of 4.410-3 /min, yielding a rate constant of 1.7 x 10-4 /s for a concentration of 0.86% enzyme on substrate. The results are seen as being typical of an enzymatic reaction.
against t was high (r2 = 0.9975), with a gradient of 4.410-3 /min, yielding a rate constant of 1.7 x 10-4 /s for a concentration of 0.86% enzyme on substrate. The results are seen as being typical of an enzymatic reaction.


Second-order plots were markedly curvilinear, which was expected based on other work in the area of enzymecatalysed hydrolysis of proteins.10 Correlation was markedly less (r2 = 0.9604) than for first-order calculations.
The significance of our results is that they indicate that protection by caricain depends upon an enzymatic reaction which hydrolyses gliadin to smaller, less toxic peptides.
Enzyme inhibition studies
Table 3 summarises the results of inhibition of caricain by imidazole and PMSF. These results show that imidazole inhibits the enzyme strongly at a concentration of 6 mmol/L, whereas PMSF has little inhibitory action. Table 3 also shows the results obtained using fractions prepared in previous work.6 In accordance with those studies, Fraction 4, the most active fraction obtained by CM Sephadex chromatography of crude papaya latex, was shown to depend mainly upon caricain for its activity. Hence it was of interest to observe that Fraction 4 was likewise strongly inhibited by imidazole, but not to the same extent as Fraction 3 was inhibited by PMSF. However, Fraction 4 may contain some glutamine cyclotransferase, as this enzyme is inhibited by imidazole.14 Fraction 3, prepared by the same technique, gave contrasting results indicating the presence of other proteases, such as prolyl endopeptidase (PEP). We found 35% inhibition of PEP from Flavobacterium at a concentration of 6 mmol/L PMSF. Studies at a concentration of 2 mmol/L of these inhibitors gave values which were in nearly all cases between one-third and two-thirds of the percentage inhibition at 6 mmol/L.

Mechanisms of detoxification of gliadin in coeliac disease
There is strong evidence for an enzyme deficiency in coeliac disease. The effects of this deficiency, that is allowing T-cell mediated immunological mechanisms to operate, causes damage to tissues.1,5,13 Considerable knowledge of the structures of the peptides that make up the protein A-gliadin now exists. Examples of those peptides implicated in toxicity in coeliac disease serve to illustrate how caricain can be an effective treatment.
Motifs associated with toxicity have been pointed out by De Ritis.15 Examples occur in peptide 11 - 19 (QNPSQQQPQ), containing the PSQQ and QQQP motifs in overlapping sequence which have been shown to be toxic in vitro.3 Enzymatic attack on the N-terminal side of proline residues is a possible way of disrupting or exposing the motifs and thus detoxifying the peptide.16 Similarly, peptide 75 - 86 (RPQQPYPQPQPQ), also shown to be toxic in vitro, has five proline residues. The most likely sequences associated with toxicity are PQQPY and QQPYP, the ones common to wheat, rye and barley, the most toxic cereals in coeliac disease. The PQQP motif in w-gliadins and homologous proteins from rye and barley was found to play an important role in the mucosal immunopathology of gluten sensitivity.17 Peptide 75 - 86 is open to attack at proline 79, resulting in disruption of these motifs. This approach has been supported by mucosal digestion studies which showed that digestion of these toxic peptides by small intestinal mucosa from coeliac patients in remission was incomplete and still left all these motifs intact, whereas digestion with normal mucosa disrupted the motifs and resulted in smaller peptides being produced.18 Another synthetic gliadin preparation is peptide 31 - 49 of A-gliadin (LGQQQPFPPQQPYPQPQPF), which has been shown to be toxic in vivo.19 This peptide contains the QQQP motif reported by De Ritis, which is part of the QQQPFP motif common to all three coeliac-toxic cereals. Furthermore, it contains PQQPY and QQPYP, mentioned previously as being other motifs common to the toxic cereals.20 Disruption of all these motifs probably occurs at proline residues, and evidence18 suggests it occurs at the N-terminal peptide bond of this amino acid residue rather than on the C-terminal side.
Prolyl endopeptidase, which attacks on the C-terminal side,21 has been investigated as an enzyme which could detoxify gliadin, but it was found to be only partially effective.22 This partial efficacy is what could be expected from its ability to disrupt the PSQQ motif, but not the QQQP motif. No significant role of PEP was found from our previous work,6 nor has PEP been found to be deficient in the mucosa of people whose coeliac disease is in remission.23
Papain catalyses the hydrolysis of esters and amides and is commonly assayed using benzoylarginine ethyl ester as a substrate.24 Papain is not regarded as being useful for detoxification of gliadin and the point has been made that only crude papain, and not pure papain, is of value in this way.25 Glutaminyl cyclase, which the authors (Messer et al.25) suggested as helping detoxification, has a limited number of N-glutaminyl peptides as its substrate, but these are of course generated as detoxification proceeds with the aid of other proteases present in the small intestine. In the context of the present study, the detoxification appears to be mainly as a result of the presence of caricain in crude papain preparations. Further clinical trials with other types of enzymes (e.g. fungal endoproteases26) will determine whether enzyme therapy can counteract the damaging effects of gluten on the small intestine in vivo. We think that these effects can be caused by direct toxic action and by invoking immunological reactions. Earlier diagnosis of the disease, as well as a more positive prognosis for those with coeliac disease, will emerge as better patient management ensues.27
Conclusions
A high degree of protection against the toxic action of gliadin on rat liver lysosomes has been shown to be offered by the enzyme caricain, and this effect appears to be relevant to coeliac disease.
The purified caricain tested was of high specific activity, requiring only 86 µg in the assay (1.7% on substrate) for 80% protection against gliadin digest, with a detectable activity down to 3.6 µg/mL in the assay. The reaction was of first order and was inhibited strongly by imidazole but weakly by PMSF. The results provide further evidence that caricain could have great significance for clinical studies of enzyme therapy in coeliac disease.
Acknowledgements
We acknowledge the generous gift of purified caricain from Professor Yvan Looze which enabled this work to be carried out and his helpful advice on the subject of caricain. We also thank RMIT University for providing the facilities to enable this research to be carried out and Associate Professor Barry Meehan of RMIT University for his expert guidance with the enzyme kinetics.
References
1. Cornell HJ, Stelmasiak T. A unified hypothesis of coeliac disease with implications for management of patients. Amino Acids. 2007;33:43-49. doi:10.1007/s00726-006-0420-0, PMid:17013762 [ Links ]
2. Bronstein HD, Haeffner LJ, Kowlessar OD. Enzymatic digestion of gliadin: The effect of the resultant peptides in adult coeliac disease. Clin Chim Acta. 1966;14:141-155. doi:10.1016/0009-8981(66)90080-5 [ Links ]
3. Kocna P, Mothes T, Krchnak V, Fric P. Relationship between gliadin peptide structure and their effect on the fetal chick duodenum. Zeitschrift fur Lebensmitteluntersuchung und-Forschung. 1991;192:116-119. doi:10.1007/BF01202623 [ Links ]
4. Cornell HJ, Mothes T. The activity of wheat gliadin peptides in in vitro assays for coeliac disease. Biochim Biophys Acta. 1993;1181:169-173. PMid:8481406 [ Links ]
5. Cornell HJ, Macrae FA, Melny J, et al. Enzyme therapy for management of coeliac disease. Scand J Gastroent. 2005;40:1304-1312. doi:10.1080/00365520510023855, PMid:16243716 [ Links ]
6. Cornell HJ, Doherty W, Stelmasiak T. Papaya latex enzymes capable of detoxification of gliadin. Amino Acids. 2010;38:155-165. doi:10.1007/s00726-008-0223-6, PMid:19156482 [ Links ]
7. Dubey VK, Pande M, Singh BK, Jagannadham MV. Papain-like proteases: Applications of their inhibitors. Afr J Biotechnol. 2007;6:1077-1086. [ Links ]
8. Zerhouni S, Amrani A, Nijs M, et al. Purification and characterization of papaya glutamine cyclotransferase, a plant enzyme highly resistant to chemical, acid and thermal denaturation. Biochim Biophys Acta. 1998;1387:275-290. doi:10.1016/S0167-4838(98)00140-X [ Links ]
9. Azarkan M, El Moussaoui A, Van Wuytswinkel D, Dehon G, Looze Y. Fractionation and purification of the enzymes stored in the latex of Carica papaya. J Chromatogr B Analyt Technol Biomed Life Sci. 2003;790:229-238. doi:10.1016/S1570-0232(03)00084-9 [ Links ]
10. Riecken EO, Stewart JS, Booth CC, Pearse AGE. A histochemical study of the role of lysosomal enzymes in idiopathic steatorrhoea before and during a gluten-free diet. Gut. 1966;7:317-332. doi:10.1136/gut.7.4.317, PMid:5917418, PMCid:1552434 [ Links ]
11. Williams VR, Williams HB. Basic physical chemistry for the life sciences. San Francisco: WH Freeman and Company, 1973; p. 277-313. [ Links ]
12. Laidler KJ. Physical chemistry with biological applications. Menlo Park, CA: Benjamin Cummings, 1978; p. 427-451. [ Links ]
13. Cornell HJ, Townley RRW. Investigating possible intestinal peptidase deficiency in coeliac disease. Clin Chim Acta. 1973;43:113-125. doi:10.1016/0009-8981(73)90126-5 [ Links ]
14. Terrell DA. The isolation of native glutaminyl cyclase from Saccharomycescerevisiae. Honours thesis, San Marcos, Texas State University, 2006. [ Links ]
15. De Ritis G, Auricchio S, Jones HW, Lew EJ-L, Bernardin JE, Kasarda DD. In vitro (organ culture) studies of the toxicity of specific A-gliadin peptides in coeliac disease. Gastroenterology. 1988;94:41-47. PMid:3335296 [ Links ]
16. Cornell HJ. The aetiology of coeliac disease and its significance for therapy. Curr Topics Peptide Protein Res. 2005;7:17-22. [ Links ]
17. Ensari A, Marsh MN, Moriarty KJ, Moore CM, Fido RJ, Tatham AS. Studies in vivo of w-gliadins in gluten sensitivity (coeliac sprue disease). Clin Sci. 1998;95:419-424. doi:10.1042/CS19980129, PMid:9748417 [ Links ]
18. Cornell HJ, Rivett DE. In vitro mucosal digestion of synthetic gliadinderived peptides in coeliac disease. J Protein Chem. 1995;14:335-339. doi:10.1007/BF01886790, PMid:8590601 [ Links ]
19. Sturgess R, Day P, Ellis HJ, et al. Wheat peptide challenge in coeliac disease. Lancet. 1994;343:758-761. doi:10.1016/S0140-6736(94)91837-6 [ Links ]
20. McLachlan A, Cullis PG, Cornell HJ. The use of extended motifs for focussing on toxic peptides in coeliac disease. J Biochem Mol Biol Biophys. 2002;6:319-324. doi:10.1080/1025814021000003238, PMid:12385967 [ Links ]
21. Szwajcer-Dey E, Rasmussen J, Meldal M, Breddam K. Proline-specific endopeptidases from microbial sources: Isolation of an enzyme from Xanthomonas sp. J Bacteriol. 1992;174:2454-2459. PMid:1556065, PMCid:205881 [ Links ]
22. Matysiak-Budnik T, Candalh C, Cellier C, et al. Limited efficiency of prolylendopeptidase in the detoxification of gliadin peptides in coeliac disease. Gastroenterology. 2005;129:786-796. doi:10.1053/j.gastro.2005.06.016, PMid:16143118 [ Links ]
23. Donlon J, Stevens FM. No lack of prolyl oligopeptidase (POP) in the coeliac mucosa. Proceedings of the 11th International Symposium on Coeliac Disease; 2004 April 28 - May 1; Belfast, Ireland. Belfast: The Coeliac Society of Ireland; 2004. p. 19. [ Links ]
24. Arnon R. The cysteine proteases: Papain. Methods Enzymol. 1970;19:226-244. doi:10.1016/0076-6879(70)19017-3 [ Links ]
25. Messer M, Anderson CM, Hubbard L. Studies on the mechanism of destruction of the toxic action of wheat gluten in coeliac disease by crude papain. Gut. 1964;5:295-303. doi:10.1136/gut.5.4.295, PMid:14209911, PMCid:1413471 [ Links ]
26. Stepniak D, Spaenij-Dekking L, Mitea C, et al. Highly efficient gluten degradation with a newly identified prolyl endopeptidase: Implications for coeliac disease. Am J Physiol Gastrointest Liver Physiol. 2006;291:G621- G629. doi:10.1152/ajpgi.00034.2006, PMid:16690904 [ Links ]
27. Cornell HJ, Stelmasiak T. Commentary - Strategies for improved outcomes for those with coeliac disease. In: Edwards MA, editor. Coeliac disease - Etiology, diagnosis and treatment. New York: Nova Science, 2009; p. 207-211. [ Links ]
 Correspondence to:
Correspondence to:
Hugh Cornell
Postal address: 124 LaTrobe Street
Melbourne 3000, Australia
Email: hugh.cornell@rmit.edu.au
Received: 23 Nov. 2010
Accepted: 21 May 2011
Published: 21 Sept. 2011
© 2011. The Authors. Licensee: AOSIS OpenJournals. This work is licensed under the Creative Commons Attribution License.














