Servicios Personalizados
Articulo
Indicadores
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares en Google
Similares en Google
Compartir
South African Journal of Science
versión On-line ISSN 1996-7489
versión impresa ISSN 0038-2353
S. Afr. j. sci. vol.107 no.5-6 Pretoria may./jun. 2011
http://dx.doi.org/10.4102/sajs.v107i5/6.514
REVIEW ARTICLE
A review of marine phylogeography in southern Africa
Peter R. TeskeI,II; Sophie von der HeydenIII; Christopher D. McQuaidII; Nigel P. BarkerI
IMolecular Ecology and Systematics Group, Botany Department, Rhodes University, Grahamstown, South Africa
IIDepartment of Zoology and Entomology, Rhodes University, Grahamstown, South Africa
IIIEvolutionary Genomics Group, Department of Botany and Zoology, Stellenbosch University, South Africa
ABSTRACT
The southern African marine realm is located at the transition zone between the Atlantic and Indo-Pacific biomes. Its biodiversity is particularly rich and comprises faunal and floral elements from the two major oceanic regions, as well as a large number of endemics. Within this realm, strikingly different biota occur in close geographic proximity to each other, and many of the species with distributions spanning two or more of the region's marine biogeographic provinces are divided into evolutionary units that can often only be distinguished on the basis of genetic data. In this review, we describe the state of marine phylogeography in southern Africa, that is, the study of evolutionary relationships at the species level, or amongst closely related species, in relation to the region's marine environment. We focus particularly on coastal phylogeography, where much progress has recently been made in identifying phylogeographic breaks and explaining how they originated and are maintained. We also highlight numerous shortcomings that should be addressed in the near future. These include: the limited data available for commercially important organisms, particularly offshore species; the paucity of oceanographic data for nearshore areas; a dearth of studies based on multilocus data; and the fact that studying the role of diversifying selection in speciation has been limited to physiological approaches to the exclusion of genetics. It is becoming apparent that the southern African marine realm is one of the world's most interesting environments in which to study the evolutionary processes that shape not only regional, but also global patterns of marine biodiversity.
Introduction
Phylogeography is the study of the historical and phylogenetic components of the spatial distribution of gene lineages within and amongst closely related species.1,2 Many phylogeographic studies have focused on species of conservation concern,3 whilst others have used the approach to investigate species complexes4 or address questions in invasion biology.5 In many instances, results have been interpreted in a somewhat narrative manner, linking genetic disjunctions with past climates or physical barriers and limited dispersal. More rigorous interpretations can be obtained when phylogeography is used in a comparative context; if the genetic structure of co-distributed but evolutionarily independent populations is congruent, then this reveals common processes that have driven genetic divergence.6,7 If focused on multiple taxa, phylogeographic studies can thus be a very powerful tool in the identification of locations and processes central to the origin and maintenance of biological diversity.8,9
In a recent review of phylogeographic studies, Beheregaray10 highlighted challenges for the Southern Hemisphere, noting that 77% of all studies were on boreal taxa, whilst biodiversity-rich developing nations are lagging in their use of this powerful method. South Africa was listed as 21st out of the 100 most productive countries in terms of publishing phylogeographic studies, with a total of 68 papers at that time. South Africa was also the 4th most productive country in the Southern Hemisphere, after Australia, Brazil and New Zealand. In light of the imbalance between the North and South, it is timely and appropriate that an assessment of the discipline in southern Africa is undertaken. Here we present a synthesis of key findings and a candid look ahead for phylogeographic research on marine organisms, which we hope can be used to identify research gaps, motivate for new studies and drive new directions, not only in regional, but also global marine biological research.
Southern Africa has a long and diverse coastline, comprising rocky and sandy shores, kelp forests, estuaries and coral reefs, yet marine phylogeography lags behind phylogeographic research on terrestrial biota. Although papers that could be considered to have a phylogeographic component were sporadically published during the 1980s11,12 and 1990s,13,14 a concerted effort to study the region's marine biota began less than a decade ago.15,16 In recent years, marine phylogeography has primarily been driven by three South African research groups, based at Rhodes University, Stellenbosch University and the University of Pretoria. Each group has its own focus: the group at Rhodes University focuses primarily on coastal invertebrates, the group at Stellenbosch University on coastal and deep-water fishes, commercially exploited crustaceans and other coastal invertebrates, and that at the University of Pretoria focuses exclusively on commercially important fish species. As southern Africa is of great interest from a biogeographic point of view because of its location at the transition zone between the Atlantic Ocean and Indian Ocean biomes, it has also featured prominently in a number of key phylogeographic studies with a global focus.17,18 In this review, we highlight several areas of research where southern African marine organisms have featured prominently, identify significant gaps in terms of both sampling design and technical aspects, and discuss how these shortcomings can be addressed in the near future.
Coastal phylogeography
Of the different fields of marine phylogeography that are being studied in southern Africa, coastal phylogeography can be considered the one about which we know most. Since 2000, 23 papers dealing with the phylogeography of coastal taxa have been published and several more are either in press or in preparation. Papers authored by South African researchers understandably dominate the literature; access to coastal sites is easy, sampling is relatively simple and cheap, and usually many samples can be obtained in a short period of time. A recent review paper examined the phylogeographic patterning of southern African coastal taxa19 and some of the general trends identified are briefly discussed below.
Location of coastal phylogeographic breaks
Most coastal species are divided into regionally confined genetic lineages whose distributions in many cases are linked with southern Africa's marine biogeographic provinces.20,21 Phylogeographic breaks separating such lineages have been identified in three regions (Figure 1).19
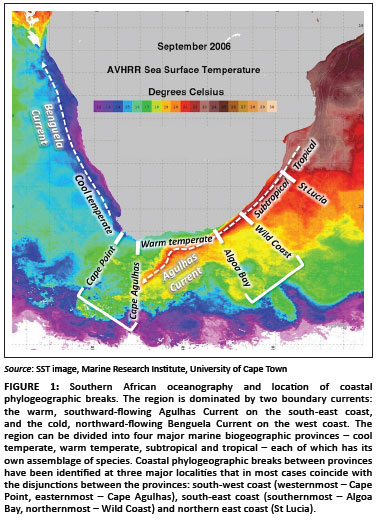
On the south-west coast, phylogeographic breaks that coincide with the biogeographic disjunction between cool-temperate and warm-temperate biota22 have been reported near Cape Point23,24 and Cape Agulhas.25,26 The region between these sites is sometimes considered a transition zone,27 and several species have phylogeographic breaks at both sites, with distinct lineages that are endemic to this transition zone.20,23
Phylogeographic breaks on the south-east coast, at the disjunction between warm-temperate and subtropical biota,22 have been difficult to define because their exact locations differ considerably for different species, and, in some, there is considerable overlap of genetic lineages.23,28 The continental shelf in this region gradually widens from north to south, deflecting the warm Agulhas Current away from the coast, limiting its influence on coastal biota (Figure 1).29 The northernmost breaks in this region have been identified on the Central Wild Coast (Transkei region)20,30 and the southernmost breaks were reported near Algoa Bay.24,31
The third area where phylogeographic breaks have been identified coincides approximately with the transition zone between subtropical and tropical biotas on the east coast;32 some species have phylogeographic breaks in north-eastern South Africa near St Lucia21,33 and others have breaks farther north in Mozambique.34,35
An important finding is that not all species that occur in more than one marine biogeographic province exhibit genetic structure, and those that do need not have phylogeographic breaks at the same localities. Some species are not genetically structured across one or more biogeographic disjunctions,20 and several taxa show no genetic structuring along their entire ranges.36,37 In addition, several species with low capacity for dispersal exhibit phylogeographic breaks that do not coincide with present-day marine biogeographic disjunctions,20,23 suggesting that in these, historical patterns are retained by limited gene flow.38 Also, although planktonic dispersers usually do not have any phylogeographic breaks within marine biogeographic provinces, this does not necessarily imply that all are panmictic within provinces. Whilst panmixia has been identified in a highly philopatric coastal fish that disperses primarily by means of planktonic larvae,37 significant genetic structure was found in the brown mussel, Perna perna.39 Populations of this species residing in different bays in the warm-temperate province were not only genetically distinct from each other on the basis of differences in haplotype frequencies, but they were also distinct from populations on the open coast.
Maintenance of coastal phylogeographic breaks
Even though many of southern Africa's coastal species have high dispersal potential because of well-developed locomotory abilities and/or extended planktonic dispersal phases, phylogeographic breaks are often surprisingly abrupt. By linking oceanography with life history, it should be possible to establish the relative importance of the interacting factors that contribute to population genetic structuring and population connectivity.40 Hypotheses explaining how distinct genetic patterns are maintained fall into two major categories, (1) genetic lineages are separated by barriers that limit dispersal and (2) regional genetic lineages are adapted to the environmental conditions characteristic of their marine biogeographic province and in many cases are unable to establish themselves in adjacent provinces.
Oceanic dispersal barriers
Proposed dispersal barriers that limit mixing of adjacent genetic lineages include upwelling cells,17 river discharge,14 coastal currents or eddies30,41 and even a coastal dunefield.20
Cold-water upwelling: Numerous studies on marine species have indicated that cold-water upwelling can represent a strong dispersal barrier.17,42 On the South African west coast, some coastal species have gaps in their distribution across a region with strong, persistent upwelling that may extend over hundreds of kilometres (e.g. the mussel Perna perna)28 and dispersal of marine organisms from the Indian Ocean into the Atlantic Ocean is limited.18,43 Some marine species show high levels of differentiation on the west coast (unpublished data), whilst in other studies, the same genetic lineages were identified on either side of the cold-water barrier,28,44 suggesting that in these, divergence was either very recent or that populations on either side are connected by ongoing gene flow. Whether genetic disjunctions on the west coast are solely linked to upwelling cells or whether local oceanographic features such as eddies retain larvae in their natal environments has yet to be examined.
Freshwater discharge: On the south-east coast, freshwater discharge from the Mbashe River has been invoked as a dispersal barrier that prevents mixing of subtropical and temperate biota.14 However, in many invertebrate species, the phylogeographic breaks in this region are not located near this river.4,30 It remains to be tested whether larger rivers, such as the Tugela or the Gariep, represent dispersal barriers that limit dispersal of marine organisms along the coast.
Currents: Currents may represent dispersal barriers when water and larvae are mostly displaced offshore, away from suitable habitat in which to settle.45 The trajectories of drifters released on the South African south and east coasts showed remarkably little overlap.41 None of the drifters released on the south coast moved close to the east coast, and drifters released on the east coast eventually became entrained in the Agulhas Current and were moved hundreds of kilometres offshore. This suggests that large-scale regional hydrodynamics significantly reduce mixing between the temperate and subtropical biotas. However, drifters were released several kilometres offshore, so it is likely that wind-driven inshore currents facilitate some northward dispersal on the south-east coast, which would explain the presence of the temperate lineages of some coastal invertebrates as far north as the central Wild Coast (Figure 1).26,30 Indeed, in an experiment using plastic drift cards, it was found that twice as many cards were retained in this region compared to cards released from two sites on the east coast, the majority of which were caught in the Agulhas Current.46
Genetic methods of analysing the strength and directionality of gene flow represent a useful additional tool for studying the role of currents in dispersal. They estimate long-term trends and only incorporate information from individuals that have dispersed and recruited successfully.
Broad-scale patterns from the different biogeographic areas show the influence of the major current systems on dispersal, and the evidence points to the importance of the interplay of the duration of larval dispersal with local current regimes. Four major gene-flow scenarios have been identified, (1) strong northward flow on the west coast with the Benguela Current, (2) strong southward flow on the east coast with the Agulhas Current, (3) some bidirectional gene flow inshore of the Agulhas Current on the south-east coast and (4) bidirectional gene flow on the south coast. These are discussed in more detail below.
The west coast, which is dominated by the northward-flowing Benguela Current, shows the strongest signal of asymmetrical or unidirectional gene flow patterns.24 Very little information on gene flow is available from the east coast, but the limited data there are support the idea that long-distance dispersal is mostly facilitated by the southward-flowing Agulhas Current.23
On the south and south-east coasts, migration is not as clear. In the barehead goby, Caffrogobius caffer, gene flow was shown to be predominantly with the Agulhas Current,36 but in another rocky shore fish, the bluntnose klipfish (Clinus cottoides), most gene flow occurred in the opposite direction.24 In invertebrates, bidirectional gene flow was identified on the south coast, with more eastward than westward dispersal,23 and gene flow on the south-east coast was also bidirectional, indicating that much dispersal takes place by means of nearshore currents.30 Together, these somewhat contradictory examples illustrate why life history plays an important role in determining population genetic structuring. For example, like many other gobioid fishes, C. caffer probably has a long larval dispersal phase47 whereas adult fishes are confined to high-shore rock pools and probably do not disperse at all.48 Dispersal is therefore only by larvae that make use of the Agulhas Current. In contrast, clinid fishes have extremely limited larval dispersal and it is highly unlikely that young fish are able to disperse by means of the offshore Agulhas Current, but likely rather use the inshore Agulhas counter-current for dispersal.24 Further evidence for counter-current driven dispersal comes from the eastward range expansion of the invasive Mediterranean mussel, Mytilus galloprovincialis.49 It is also likely that strengthening of the current during the winter months facilitates the annual sardine run in South Africa.50
Dunefields: Dunefields seem to be an unlikely dispersal barrier for marine species, but their importance has also been documented elsewhere.51 A possible explanation is that, in addition to coastal dunefields representing long stretches of unsuitable habitat for rocky shore and estuarine species, regions where these are located are characterised by strong, persistent onshore winds, which may limit long-shore dispersal of plankton in the surface water.
Province-specific adaptations
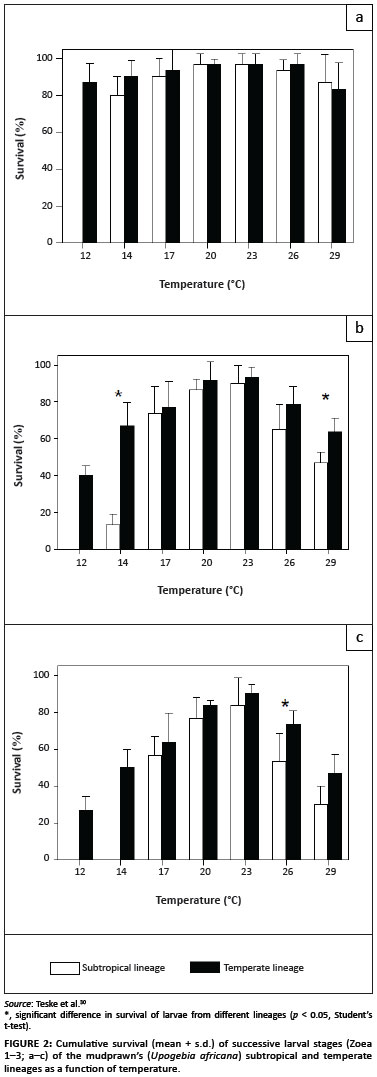
Although there is little doubt that dispersal barriers limit gene flow between provinces, many can be considered to be incomplete. For example, many upwelling cells affect the surface waters for only short periods of time,52 and many wind patterns (such as the shoreward south-easterly) are seasonal.53 Maintenance of genetic structure in the absence of strong dispersal barriers is possible only when levels of selection are high.54 An alternative hypothesis explaining the maintenance of coastal phylogeographic breaks suggests that although many species can reach adjacent provinces, they do not establish themselves permanently, either because they are ill-adapted to local environmental conditions or because they are outcompeted by their sister taxa. Adaptation of genetic lineages to environmental conditions that differ between provinces has been documented in several recent studies. The larvae of the subtropical lineage of the mudprawn, Upogebia africana, cannot survive the colder water temperatures that are typical of the temperate province during winter (Figure 2).30 This observation suggests that, even though they can potentially settle outside their own province during summer,23 they are unable to establish themselves in the temperate provinces. Differences in osmoregulatory abilities of warm-temperate and subtropical lineages of the estuarine sandprawn, Callianassa kraussi, may reflect adaptation to differences in the salinities of the estuaries of each region, and therefore limit dispersal of each genetic lineage into adjacent provinces.21 Lastly, the fact that the temperate lineage of the brown mussel, Perna perna, is less tolerant of sand inundation and high temperatures than its subtropical sister lineage may partly explain its absence from the east coast.41
Origin of coastal phylogeographic breaks
Most studies on southern African coastal taxa that describe phylogeographic breaks limit themselves to suggesting factors that are likely to maintain them. Explaining how such patterns have arisen is proving more challenging. Molecular dating indicates that coastal phylogeographic breaks are the result of historical processes that precede the beginning of the present interglacial period.20,55 However, such estimates are mostly based on few loci, which limits accuracy, and the markers used may not provide sufficient resolution to detect very recent divergence events (discussed below). The ages of congruent genetic disjunctions may differ considerably for different species, and genetic differentiation between sister lineages in adjacent provinces may range from differences in haplotype frequencies in recently diverged lineages24,34 to lineages being so distinct that each can be considered to be a distinct species.21,33 For example, divergence time estimates that were based on more than one locus indicated that in the brachyuran crown crab species complex, Hymenosoma orbiculare, a split into temperate and subtropical lineages occurred at least 16 million years ago,4 whereas congruent regional genetic units of the clinid fish C. cottoides diverged as recently as 60 000 years ago.24
In contrast to south-eastern Australia56 or Indonesia,57 there are no geological features in southern Africa that could have acted as land bridges during episodes of low sea level and that could have completely isolated populations of coastal taxa. There is consequently no compelling evidence for any geological vicariance events along the coast that could have driven simultaneous divergence in multiple species. The region's coastal morphology nonetheless changed considerably as a result of climate oscillations during the Pleistocene. For example, during the Last Glacial Maximum (26 500 - 19 000 years ago),58 when the sea level was about 120 m lower than it is today,59 large areas of continental shelf were exposed, particularly south of Cape Agulhas.60 Also, the region's sea surface temperatures cooled as a result of intensified upwelling on the west coast61 and a reduced influence of the Agulhas Current.62 How these changes may have affected habitat availability and the amount of gene flow along the coast is poorly understood, but the role of oceanic dispersal barriers (discussed in the previous section) in driving the evolution of regional lineages needs to be assessed in this context. The exposure of the Agulhas Bank during the Last Glacial Maximum resulted in the southern tip of Africa being about 200 km south of where it is today, and, in combination with colder water temperatures in the region during that time, this may have presented a cold-water dispersal barrier similar to that on the west coast. The Agulhas Current weakened during glacial phases and may have ceased to flow during winter,62 suggesting that advection of larvae away from the coast would have been considerably reduced, with stronger bidirectional longshore dispersal by means of nearshore currents. This possibility suggests that the role of the Agulhas Current in limiting mixing of regional biotas may never have been substantially more important than it is today. The fact that species from the east coast can temporarily establish themselves in the eastern portion of the temperate province during the summer months63 indicates that, even today, it represents a highly permeable barrier. The same can be said of upwelling cells and the freshwater plumes of large rivers. Also, some dispersal barriers have formed more recently than the genetic lineages they separate,20 suggesting that they only contribute towards maintaining genetic structure that was already present. Population genetic theory suggests that even a small amount of migration between populations will prevent genetic divergence by drift,64 which indicates that southern Africa's historical oceanic dispersal barriers may be insufficient to explain the origin of marine phylogeographic breaks.
The association of genetic lineages with marine biogeographic provinces could point to ecological factors driving genetic divergence. Two recent studies have indicated that divergence could be driven by climate oscillations. In the first study, a range expansion from the south-east coast to the south-west coast that occurred during the previous interglacial period (~120 000 years ago) was identified in the coastal snail Nassarius kraussianus by means of coalescent-based molecular dating of mitochondrial DNA (mtDNA) sequence data (Figure 3).31 Westward range expansions of warm-water molluscs during this period are well documented in the fossil record.65 However, in contrast to other coastal molluscs, subsequent climatic cooling did not result in a range contraction in N. kraussianus, whose shells were used as ornaments by humans living on the south-west coast during the last glacial period.66 This lack of range contraction suggests that the species' western populations adapted to cooler water. The species is today represented by a younger temperate lineage and a more ancient lineage that occurs in the subtropical and tropical provinces, with a phylogeographic break located near Algoa Bay (Figure 1). Congruent but much older divergence events that could be linked to range expansions during warm climatic phases, followed by adaptation and speciation during cooler phases, have also been reported in the Hymenosoma orbiculare species complex using multilocus DNA sequence data.4
Ecological divergence scenarios linked to climate oscillations may explain why divergence times differ considerably amongst species with congruent phylogeographic breaks. Although most species may undergo range expansions in response to shifting boundaries between marine biogeographic provinces as a result of climate oscillations,65 adaptations to unfavourable environmental conditions during a particular range shift may only have arisen in a small fraction of the species affected, whilst the ranges of most others would have contracted.
Offshore marine phylogeography
Genetic studies of offshore populations are rare compared to coastal species. This difference can be ascribed to a number of factors, including the expense of obtaining samples, the lack of taxonomic expertise for some groups, and the lack of inclusion in multinational, large-scale research programmes. Research initiatives such as the Western Indian Ocean Marine Science Association, the Agulhas and Somali Current Large Ecosystem Project and the African Coelacanth Ecosystem Programme should have boosted such research, but, to date, little research from such initiatives has been published. There is considerable room for scientists working in southern Africa to expand into offshore phylogeographic studies, and perhaps some national strategy in this regard should be developed.
Recent offshore marine phylogeographic studies in southern Africa have primarily focused on commercially exploited species, with special emphasis on the Cape hakes that comprise the most important demersal fisheries resource in South Africa and Namibia. The major focus regarding offshore stocks is on transboundary management between Namibian and South African fish stocks. Interestingly, there appears to be genetic structure in the deep-water hake, Merluccius paradoxus,55,67 between Namibia and South Africa, as well as between individuals within South Africa. However, as M. paradoxus was shown to only have population genetic structuring with adult fishes and no structure for juvenile non-spawning fish,67 the focus to date has been on understanding the structuring of adult fishes. One study has examined the distribution of the eggs and larvae of M. paradoxus and those of its shallow-water sister taxon M. capensis (which in the early stages are morphologically identical) and found that spawning depth differs signficantly, but that most larvae of a certain size are found on the continental shelf at depths of about 200 m.68 Several other studies have used molecular techniques on offshore marine species in southern Africa; for example, the lanternfish, Lampanyctodes hectoris, showed no significant genetic differentiation between South African and Namibian stocks.69 There also appears to be no differentiation in the squid Loligo vulgaris between disparate spawning areas in South Africa.70 In contrast, horse mackerels, Trachurus capensis, show slight differentiation between Namibia and South Africa based on allozyme loci.11 Preliminary research using allozymes also suggested slight stock differentiation of orange roughy, Hoplostethus atlanticus, in Namibian waters.71
There have also been a number of studies on commercially important crustaceans, in particular rock lobsters of the genera Jasus and Palinurus. These have primarily focused on understanding population genetic structuring and demographic changes of lobster species in the region. Even though lobsters have extended planktonic dispersal phases, some genetic structuring between sampling areas was recovered for the Tristan rock lobster, Jasus tristani,72 and for the Natal deep-sea lobster, Palinurus delagoae, which exhibits shallow but significant structuring between Mozambican and South African populations.34 In the most important commercially exploited crustacean in South Africa, the west-coast rock lobster, Jasus lalandii, genetic diversity is highest on the south-west coast and decreases towards the edge of the species' distribution.73 In contrast, the south-coast rock lobster, Palinurus gilchristi, shows no population genetic structuring along its range.74
Southern Africa and beyond
Indo-West Pacific phylogeography
Many marine organisms with high dispersal potential have long been considered to have distributions incorporating the entire Indian Ocean whereas the presence of temperate low-dispersal species in southern Africa and Australasia has traditionally been attributed to morphological stasis in Gondwanan relics that diverged as a result of the break-up of the ancient supercontinent. Both paradigms have been challenged by genetic studies. Large-scale phylogeographic studies have indicated that the populations of many marine organisms can be divided into lineages that are confined to the western Indian Ocean and lineages that are associated with the eastern Indian Ocean and/or the West Pacific.75,76 A recent study on fish species that occur both in southern Africa and in Australia indicated that, although there is little genetic structure in pelagic species, many inshore species are highly divergent between the regions. This study suggests that a third of the nearly 1000 fish species that occur in both regions may include cryptic species.77 Molecular dating further indicates that many of the low-dispersal species thought to be Gondwanan relics diverged long after the break-up of the supercontinent, and post-Gondwanan transoceanic dispersal is considered to be a more appropriate hypothesis explaining the observed sister-taxon relationships.78 Colonisation of Australia from southern Africa via the west wind drift has been proposed for low dispersal species,78 but colonisation patterns of highly dispersed taxa are not yet fully understood.79
Coastal phylogeography in the western Indian Ocean
Despite the importance of the western Indian Ocean as a biodiversity hotspot and several major research initiatives, the region's phylogeography remains poorly explored. The majority of phylogeographic studies dealing with marine species from the western Indian Ocean have included samples from South Africa only.28,80 Studies that included samples from other western Indian Ocean countries have mostly compared large-scale genetic structure between the western and eastern Indian Ocean, or throughout the Indo-West Pacific.75,81 Very few studies have focused on genetic structure and gene flow along the East African coast, or between the African mainland and the region's islands. In those that have, low sample sizes and the inclusion of just a handful of sampling sites have made inferences about the location of phylogeographic breaks and levels of gene flow throughout the region problematic,82,83 a notable exception being a recent study of the fiddler crab, Uca annulipes.84 An earlier attempt at summarising what few data there are suggested that genetic structure in the western Indian Ocean exists mostly at tropical locations, whereas south-eastern African marine populations lacked genetic structure,85 a trend that was rejected by several more recent studies.34,35 To date, most of the phylogeographic studies that have employed a fine-scale sampling approach, and that have not dealt exclusively with South African fauna, involved extensive sampling in South Africa plus some additional Mozambican sites.33,35 Even these studies have suffered from the problem of large gaps between the South African and Mozambican sites. To study the phylogeography of the western Indian Ocean more comprehensively requires that the level of sampling that has proven so useful to detect genetic structure in temperate and subtropical South Africa be extended to the tropical regions to the north-east. However, the size of this region, and the logistical difficulties involved in reaching sampling sites, will require not only substantial funding, but also a strengthening of collaborations amongst researchers from different western Indian Ocean countries.
Antitropical distributions in the eastern Atlantic Ocean
Several temperate southern African marine animals have sister-taxon relationships with species in the temperate north-eastern Atlantic. Examples include hake (Merluccius spp.),86 anchovies (Engraulis spp.),87 krill (Nyctiphanes spp.),88 Octopus vulgaris,89 spiny lobsters (Palinurus spp.)90 and intertidal ascidians (Pyura herdmani).91 Although it is possible that some of these disjunct distributions are the result of recent human-mediated transport from one region to the other, molecular dating indicates that most divergence events considerably predate the historical period. This suggests that migrants must have crossed the highly significant dispersal barrier represented by warm equatorial waters to establish themselves successfully. The findings to date indicate that the dispersal direction may differ amongst taxa. Whereas dispersal in Nyctiphanes spp. and P. herdmani was most likely from southern Africa to the north-eastern Atlantic, Merluccius spp., Engraulis spp. and Palinurus spp. most likely originated in the Northern Hemisphere.
Genetic markers used in marine phylogeography
Mirroring a global trend in phylogeographic research,10 the majority of studies on southern African marine organisms have used mtDNA sequence data, with the cytochrome oxidase c subunit I gene being particularly popular. The reason for this is obvious - the primers for this marker are 'universal' and can be used for a wide variety of taxa. However, there are numerous disadvantages to using mtDNA exclusively, including that with a few exceptions, it is only inherited in the female line and is thus unsuitable for the study of hybridisation or reproductive isolation amongst different genetic lineages, and that molecular dating based on a single marker is less accurate than dating based on multilocus data.92
Several recent studies have used nuclear sequence data such as nuclear genes, introns or ribosomal RNAs in conjunction with mtDNA, and congruent genetic patterns were recovered for the two types of genetic markers.24,33 These studies have so far rejected the notion that in species with low dispersal potential, haphazard genetic structure can readily arise in the absence of any underlying environmental factors.93 To researchers who have exclusively used mtDNA sequence data until now, introns are likely to become the nuclear marker of choice. Not only are similar skills required in terms of data generation and interpretation, but the information content of introns is similar to that of mtDNA. Recent software developments for phasing the two sequences superimposed onto each other in trace files generated from heterozygous individuals94,95 have rendered tedious cloning unnecessary, and a number of universal96 and taxon-specific33,97,98 primer sets have been developed. In non-model organisms for which no suitable primers are available for amplifying introns, the development of anonymous nuclear markers99 may be a suitable alternative.
Whilst DNA sequence data from mtDNA or nuclear markers have proven suitable for detecting phylogeographic breaks and identifying cryptic speciation, they are of limited use in the study of very recently evolved genetic patterns, such as those that formed during or after the Last Glacial Maximum, or those that formed during historical times. Microsatellites (also known as short tandem repeats) are excellent markers for the study of such recent evolutionary events because of their high mutation rate. Even though a number of microsatellite libraries have been developed specifically for South African marine organisms, particularly for teleosts,100,101,102,103 we are aware of only four research papers that have actually used these markers to study marine phylogeography in southern Africa.25,35,37,104
Other types of genetic markers with considerable potential for the elucidation of marine phylogeography have yet to be used in southern Africa. For example, amplified fragment length polymorphism (AFLP) is now firmly established as a genetic marker for terrestrial plants,105 but we are not aware of any studies on southern African algae, seagrasses or mangroves that have used them, and their use in animals is so far limited to aquaculture.106
Where to from here?
Although there are substantial data on some aspects of marine phylogeography in southern Africa, other aspects require further attention. Firstly, despite considerable insight into marine phylogeographic breaks gained during the past decade, two regions have not received sufficient attention. In the tropical north-east, phylogeographic breaks in species with low dispersal potential were identified near St Lucia,21,33 and several planktonic dispersers have phylogeographic breaks in southern Mozambique.34,35 Because of logistical difficulties in accessing sites, there were large gaps between sampling sites in all studies focusing on this region, and it is possible that there is in fact more than one phylogeographic break. Even less research has focused on the west coast, and more intensive sampling, which also includes sites in Namibia and Angola, is needed to better understand genetic structuring in this region.19
In terms of the nearshore biota studied, most research has so far focused on rocky shore or estuarine species, and only two studies have been on sandy shore organisms.13,16 As sandy beaches make up about 42% of the South African shoreline and are a dominant feature particularly on the east coast,107 more research efforts should be concentrated on understanding the genetic structuring of sandy shore organisms.
Surprisingly little phylogeographic research has been conducted on commercially important species.37,67,89,104 Given that commercially exploited coastal teleosts have primarily been used as model taxa to position Marine Protected Areas, more phylogeographic research evaluating the current Marine Protected Areas network is warranted. In addition, the dearth of offshore genetic research demands serious attention. In the light of increased commercial, artisanal and recreational fishing, as well as possible warming of ocean currents in the region,108 it becomes all the more important not only to understand population structuring, but also the likely evolutionary response of offshore marine species to climatic change.55 Phylogeographic studies have also inadvertently uncovered cryptic speciation in marine species.109 With at least 25% of southern African endemic fishes yet to be described,110 it is likely that biodiversity inventories not only of fishes, but of all marine taxa will greatly benefit from phylogeographic research.
In addition to focusing on neglected taxa and obtaining samples from regions where little research has been conducted, considerably more effort needs to be placed on generating, not only multispecies, but also multilocus genetic data sets. In addition to the increased use of nuclear sequence data and AFLPs, the development of microsatellite libraries needs to be a major focus of southern African marine phylogeography in the coming years. As a result of their high mutation rate, microsatellites will allow researchers to study genetic patterns driven by factors such as fishing pressure and climate change, as well as to obtain more reliable information on gene flow. For example, migration rates estimated using coalescent-based methods such as those implemented in MIGRATE-N111 or IMa112 are often interpreted as reflecting contemporary gene flow,23,24 but they may in fact be strongly influenced by historical events,111 particularly when they are based on comparatively slowly evolving markers such as mtDNA or introns. Microsatellites would further allow the identification of cryptic species or stocks that have evolved too recently to be detectable using DNA sequence data. Recent advances in sequencing technology (e.g. 454 pyrosequencing) are likely to make the development of microsatellite libraries and single nucleotide polymorphism libraries more accessible to southern African researchers.
Conclusion
The southern African marine realm is an exceptionally interesting environment in which to study evolutionary processes. Because it is located at the transition zone between the Atlantic Ocean and Indo-Pacific biomes, the region's biodiversity is particularly high. Although South Africa has a very active marine biological community and conventional marine research is of a high standard, research addressing fundamental evolutionary concepts is still poorly developed. In the coming years, marine phylogeographic research needs to move from being mostly descriptive to becoming more analytical. For example, most studies have been limited to interpreting phylogeographic patterns on the basis of oceanographic data, but it would be desirable to explore how marine organisms' evolutionary histories have shaped present-day patterns, which should include testing alternative hypotheses of when and how genetic structure evolved.113,114 Oceanographic research in southern Africa has concentrated on offshore features, with an enormous emphasis on the economically important Benguela upwelling system,115 and to a lesser degree on the Agulhas Current.52 Although recent initiatives have begun to address this,116 we have a relatively poor understanding of the hydrodynamically complex nearshore region that hampers our ability to interpret genetic data from taxa that live in shallow waters and disperse within the nearshore arena. In addition, the fact that there is strong evidence for adaptive differentiation between recently evolved sister lineages in the region's different marine provinces suggests that a greater focus needs to be placed on studying selection pressure. In addition to conducting physiological studies on evolutionary lineages that have been identified using selectively neutral genetic markers, focusing on markers that are under selection would greatly improve our understanding of the relative importance of dispersal barriers and selection gradients in driving the evolution of new species. We believe that it is time to put southern Africa 'on the map' as one of the world's most interesting regions in which to study marine phylogeography, and help afford it a similar status to that presently occupied by the Cape Floristic Kingdom, the African Great Lakes and the terrestrial fauna of Madagascar.
Acknowledgements
This review emanates from a workshop on phylogeography held at Rhodes University in June 2007. This workshop was funded by the National Research Foundation (NRF) of South Africa through the South African Biosystematics Initiative (SABI) and was attended by 28 South African delegates involved in phylogeographic research, including NPB, CDM and PRT. The ideas presented here are a consequence of discussions held during this workshop, augmented by subsequent research findings. We are grateful to J. Field for providing the SST map of southern Africa. PRT was supported by a Postdoctoral Research Fellowship from Rhodes University. SvdH was supported by a Claude Leon Postdoctoral fellowship. NPB acknowledges the NRF and Rhodes University Joint Research Council for financial support. This work is based upon research supported by the South African Research Chairs Initiative of the Department of Science and Technology and the NRF.
References
1. Avise JC. Phylogeography: Retrospect and prospect. J Biogeogr. 2009;36:3-15. doi:10.1111/j.1365-2699.2008.02032.x [ Links ]
2. Hewitt GM. Genetic consequences of climatic oscillations in the Quaternary. Phil Trans Roy Soc Lond B. 2004;359:193-195. doi:10.1098/ rstb.2003.1388, PMid:15101575, PMCid:1693318 [ Links ]
3. Bowen BW, Karl SA. Population genetics and phylogeography of sea turtles. Mol Ecol. 2007;16:4886-4907. doi:10.1111/j.1365-294X.2007.03542.x, PMid:17944856 [ Links ]
4. Teske PR, McLay C, Sandoval-Castillo J, et al. Tri-locus sequence data reject a "Gondwanan origin hypothesis" for the African/South Pacific crab genus Hymenosoma. Mol Phylogenet Evol. 2009;53:23-33. doi:10.1016/j. ympev.2009.05.031, PMid:19501181 [ Links ]
5. Rius M, Pascual M, Turon X. Phylogeography of the widespread marine invader Microcosmus squamiger (Ascidiacea) reveals high genetic diversity of introduced populations and non-independent colonizations. Divers Distrib. 2008;14:818-828. doi:10.1111/j.1472-4642.2008.00485.x [ Links ]
6. Knowles LL. Statistical phylogeography. Annu Rev Ecol Evol S. 2009;40:593-612. doi:10.1146/annurev.ecolsys.38.091206.095702 [ Links ]
7. Hickerson MJ, Carstens BC, Cavender-Bares J, et al. Phylogeography's past, present, and future: 10 years after Avise, 2000. Mol Phylogenet Evol. 2010;54:291-301. doi:10.1016/j.ympev.2009.09.016, PMid:19755165 [ Links ]
8. Bermingham E, Moritz C. Comparative phylogeography: Concepts and applications. Mol Ecol. 1998;7:367-369. doi:10.1046/j.1365- 294x.1998.00424.x [ Links ]
9. Soltis DE, Morris AB, McLachlan JS, Manos PS, Soltis PS. Comparative phylogeography of unglaciated eastern North America. Mol Ecol. 2006;15:4261-4293. doi:10.1111/j.1365-294X.2006.03061.x, PMid:17107465 [ Links ]
10. Beheregaray LB. Twenty years of phylogeography: The state of the field and the challenges for the southern hemisphere. Mol Ecol. 2008;17:3754-3774. PMid:18627447 [ Links ]
11. Zenkin VS, Komarov YA. Genetic differentiation in the Cape horse mackerel (Trachurus trachurus capensis Castelnau) population. Colln scient pap Int Comm SE Atl Fish. 1981;8:291-298. [ Links ]
12. Becker II, Grant WS, Kirby R, Robb FT. Evolutionary divergence between sympatric species of southern hakes, Merluccius capensis and M. paradoxus. II. Restriction enzyme analysis of mitochondrial DNA. Heredity. 1988;61:21-30. doi:10.1038/hdy.1988.87 [ Links ]
13. Grant WS, Da Silva-Tatley FM. Lack of genetically-subdivided population structure in Bullia digitalis, a southern African marine gastropod with lecithotrophic development. Mar Biol. 1997;129:123-137. doi:10.1007/ s002270050153 [ Links ]
14. Ridgway TM, Stewart BA, Branch GM, Hodgson AM. Morphological and genetic differentiation of Patella granularis (Gastropoda: Patellidae) along the coast of South Africa. J Zool Lond. 1998;245:317-333. doi:10.1111/j.1469-7998.1998.tb00107.x [ Links ]
15. Teske PR, Matthee CA, Cherry MI. Population genetics of the endangered Knysna seahorse, Hippocampus capensis. Mol Ecol. 2003;12:1703-1715. doi:10.1046/j.1365-294X.2003.01852.x, PMid:12803625 [ Links ]
16. Laudien J, Flint NS, Van der Bank FH, Brey T. Genetic and morphological variation in four populations of the surf clam Donax serra (Röding) from southern African sandy beaches. Biochem Syst Ecol. 2003;31:751-772. doi:10.1016/S0305-1978(02)00252-1 [ Links ]
17. Lessios HA, Kane J, Robertson DR. Phylogeography of the pantropical sea urchin Tripneustes: Contrasting patterns of population structure between oceans. Evolution. 2003;57:2026-2036. doi:10.1554/02-681, PMid:14575324 [ Links ]
18. Rocha LA, Robertson DR, Rocha CR, Van Tassell JL, Craig MT, Bowen BW. Recent invasion of the tropical Atlantic by an Indo-Pacific coral reef fish. Mol Ecol. 2005;14:3921-3928. doi:10.1111/j.1365-294X.2005.02698.x, PMid:16262848 [ Links ]
19. Von der Heyden S. Why do we need to integrate population genetics into South African marine protected area planning? Afr J Mar Sci. 2009;31:263-269. doi:10.2989/AJMS.2009.31.2.14.886 [ Links ]
20. Teske PR, McQuaid CD, Froneman PW, Barker NP. Impacts of marine biogeographic boundaries on phylogeographic patterns of three South African estuarine crustaceans. Mar Ecol Prog Ser. 2006;314:283-293. doi:10.3354/meps314283 [ Links ]
21. Teske PR, Winker H, McQuaid CD, Barker NP. A tropical/subtropical biogeographic disjunction in southeastern Africa separates two evolutionary significant units of an estuarine prawn. Mar Biol. 2009;156:1265-1275. doi:10.1007/s00227-009-1168-3 [ Links ]
22. Emanuel BP, Bustamante RH, Branch GM, Eekhout S, Odendaal FJ. A zoogeographic and functional approach to the selection of marine reserves on the west coast of South Africa. S Afr J Mar Sci. 1992;12:341-354. doi:10.2989/02577619209504710 [ Links ]
23. Teske PR, Papadopoulos I, Zardi GI, et al. Implications of life history for genetic structure and migration rates of five southern African coastal invertebrates: Planktonic, abbreviated and direct development. Mar Biol. 2007;152:697-711. doi:10.1007/s00227-007-0724-y [ Links ]
24. Von der Heyden S, Prochazka K, Bowie RCK. Significant population structure and asymmetric gene flow patterns amidst expanding populations of Clinus cottoides (Perciformes, Clinidae): Application of molecular data to marine conservation planning in South Africa. Mol Ecol. 2008;17:4812-4826. doi:10.1111/j.1365-294X.2008.03959.x, PMid:19140974 [ Links ]
25. Evans BS, Sweijd NA, Bowie RCK, Cook PA, Elliott NG. Population genetic structure of the perlemoen, Haliotis midae in South Africa: Evidence of range expansion and founder events. Mar Ecol Prog Ser. 2004;270:163-172. doi:10.3354/meps270163 [ Links ]
26. Teske PR, Froneman PW, McQuaid CD, Barker NP. Phylogeographic structure of the caridean shrimp Palaemon peringueyi in South Africa: Further evidence for intraspecific genetic units associated with marine biogeographic provinces. Afr J Mar Sci. 2007;29:253-258. doi:10.2989/ AJMS.2007.29.2.9.192 [ Links ]
27. Bolton JJ, Anderson RJ. Marine vegetation. In: Cowling RM, Richardson DM, Pierce SM, editors. Vegetation of southern Africa. Cambridge: Cambridge University Press, 1997; p. 348-375. [ Links ]
28. Zardi GI, McQuaid CD, Teske PR, Barker NP. Unexpected genetic structure of indigenous (Perna perna) and invasive (Mytilus galloprovincialis) mussel populations in South Africa. Mar Ecol Prog Ser. 2007;337:135-144. doi:10.3354/meps337135 [ Links ]
29. Heydorn AEF, Bang ND, Pearce AF, et al. Ecology of the Agulhas Current region: An assessment of biological responses to environmental parameters in the south-west Indian Ocean. Trans Roy Soc S Afr. 1978;43:151-190. doi:10.1080/00359197809520235 [ Links ]
30. Teske PR, Papadopoulos I, Newman BK, Dworschak PC, McQuaid CD, Barker NP. Oceanic dispersal barriers, adaptation and larval retention: An interdisciplinary assessment of potential factors maintaining a phylogeographic break between sister lineages of an African prawn. BMC Evol Biol. 2008;8:834. doi:10.1186/1471-2148-8-341, PMid:19108720, PMCid:2628387 [ Links ]
31. Teske PR, Papadopoulos I, Newman BK, McQuaid CD, Barker NP. Climate change, genetics or human choice: Why were the shells of mankind's earliest ornament larger in the Pleistocene than in the Holocene? PLoS ONE. 2007;2:e614. [ Links ]
32. Bolton JJ, Leliaert F, de Clerck O, et al. Where is the western limit of the tropical Indian Ocean seaweed flora? An analysis of intertidal seaweed biogeography on the east coast of South Africa. Mar Biol. 2004;144:51-59. doi:10.1007/s00227-003-1182-9 [ Links ]
33. Teske PR, Barker NP, McQuaid CD. Lack of genetic differentiation among four sympatric southeast African intertidal limpets (Siphonariidae): Phenotypic plasticity in a single species? J Molluscan Stud. 2007;73:223-228. doi:10.1093/mollus/eym012 [ Links ]
34. Gopal K, Tolley KA, Groeneveld JC, Matthee CA. Mitochondrial DNA variation in spiny lobster Palinurus delagoae suggests genetically structured populations in the southwest Indian Ocean. Mar Ecol Prog Ser. 2005;319:191-198. doi:10.3354/meps319191 [ Links ]
35. Ridgway T, Riginos C, Davis J, Hoegh-Guldenberg O. Genetic connectivity patterns of Pocillopora verrucosa in southern African Marine Protected Areas. Mar Ecol Prog Ser. 2008;354:161-168. doi:10.3354/meps07245 [ Links ]
36. Neethling M, Matthee CA, Bowie RCK, Von der Heyden S. Evidence for panmixia despite barriers to gene flow in the southern African endemic, Caffrogobius caffer (Teleostei: Gobiidae). BMC Evol Biol. 2008;8:325. doi:10.1186/1471-2148-8-325, PMid:19040766, PMCid:2613416 [ Links ]
37. Teske PR, FRG Forget, Cowley PD, Von der Heyden S, Beheregaray LB. Connectivity between marine reserves and exploited areas in the philopatric reef fish Chrysoblephus laticeps (Teleostei: Sparidae). Mar Biol. 2010;157:2029-2042. doi:10.1007/s00227-010-1471-z [ Links ]
38. Pelc RA, Warner RR, Gaines SD. Geographical patterns of genetic structure in marine species with contrasting life histories. J Biogeogr. 2009;36:1881-1890. doi:10.1111/j.1365-2699.2009.02138.x [ Links ]
39. Nicastro KR, Zardi GI, McQuaid CD, Teske PR, Barker NP. Coastal topography drives genetic structure in marine mussels. Mar Ecol Prog Ser. 2008;368:189-195. doi:10.3354/meps07607 [ Links ]
40. Carson HS, López-Duarte MP, Rasmussen L, Wang D, Levin LA. Reproductive timing alters population connectivity in marine metapopulations. Curr Biol. 2010;20:1926-1931. doi:10.1016/j. cub.2010.09.057, PMid:20951048 [ Links ]
41. Zardi GI, Nicastro KR, McQuaid CD, Hancke L, Helmuth B. The combination of selection and dispersal helps explain genetic structure in intertidal mussels. Oecologia. 2011;165:947-958. doi:10.1007/s00442-010- 1788-9, PMid:20878422 [ Links ]
42. Olivares Banuelos NC, Enriquez-Paredes LM, Ladah LB, De La Rosa-Velez J. Population structure of the purple sea urchin, Strongylocentrotus purpuratus, along the Baja California peninsula. Fisheries Sci. 2008;74:804-812. doi:10.1111/j.1444-2906.2008.01592.x [ Links ]
43. Floeter SR, Rocha LA, Robertson DR, et al. Atlantic reef fish biogeography and evolution. J Biogeogr. 2008;35:22-47. [ Links ]
44. Edkins MT, Teske PR, Griffiths CL, Papadopoulos I. Genetic and morphological analyses suggest that southern African crown crabs, Hymenosoma orbiculare, represent five distinct species. Crustaceana. 2007;80:667-683. doi:10.1163/156854007781360694 [ Links ]
45. Hare MP, Guenther C, Fagan WF. Nonrandom larval dispersal can steepen marine clines. Evolution. 1995;59:2509-2517. doi:10.1554/05- 150.1, PMid:16526499 [ Links ]
46. Steinke TD, Ward CJ. Use of plastic drift cards as indicators of possible dispersal potential of propagules of the mangrove Avicennia marina by ocean currents. Afr J Mar Sci. 2003;25:169-176. doi:10.2989/18142320309504007 [ Links ]
47. Victor BC, Vasquez-Yeomans L, Valdez-Moreno M, et al. The larval, juvenile, and adult stages of the Caribbean reef goby, Coryphopterus kuna (Teleostei: Gobiidae): A reef fish with a pelagic larval duration longer than the post-settlement lifespan. Zootaxa. 2010;2346:53-61. [ Links ]
48. Butler GS. Aspects of the biology of Caffrogobius caffer (Günther) (Pisces: Teleostei: Gobiidae) in the eastern Cape. MSc thesis, Grahamstown, Rhodes University, 1980. [ Links ]
49. McQuaid CD, Phillips TE. Limited wind-driven dispersal of intertidal mussel larvae: In situ evidence from the plankton and the spread of the invasive species Mytilus galloprovincialis in South Africa. Mar Ecol Prog Ser. 2000;201:211-220. doi:10.3354/meps201211 [ Links ]
50. Roberts MJ, Van den Berg M. Currents along the Tsitsikamma coast, South Africa, and potential transport of squid paralarvae and ichthyoplankton. Afr J Mar Sci. 2005;27:375-388. doi:10.2989/18142320509504096 [ Links ]
51. Ayre DJ, Minchinton TE, Perrin C. Does life history predict past and current connectivity for rocky intertidal invertebrates across a marine biogeographic barrier? Mol Ecol. 2009;18:1887-1903. doi:10.1111/j.1365- 294X.2009.04127.x, PMid:19434808 [ Links ]
52. Lutjeharms JRE. The Agulhas Current. Berlin: Springer; 2006. [ Links ]
53. Shillington FA. Oceanography of the southern African region. In: Smith MM, Heemstra PC, editors. Smith's sea fishes. Cape Town: Struik, 1995; p. 22-28. [ Links ]
54. Wares JP, Pringle JM. Drift by drift: Effective population size limited by advection. BMC Evol Biol. 2008;8:235. doi:10.1186/1471-2148-8-235, PMid:18710549, PMCid:2536672 [ Links ]
55. Von der Heyden S, Lipinski MR, Matthee CA. Remarkably low mtDNA control region diversity in an abundant demersal fish. Mol Phylogenet Evol. 2010;55:1183-1188. doi:10.1016/j.ympev.2009.09.018, PMid:19761857 [ Links ]
56. Waters JM. Marine biogeographical disjunction in temperate Australia: Historical landbridge, contemporary currents, or both? Divers Distrib. 2008;14:692-700. doi:10.1111/j.1472-4642.2008.00481.x [ Links ]
57. Teske PR, Hamilton H, Matthee CA, Barker NP. Signatures of seaway closures and founder dispersal in the phylogeny of a circumglobally distributed seahorse lineage. BMC Evol Biol. 2007;7:138. doi:10.1186/1471- 2148-7-138, PMid:17697373, PMCid:1978501 [ Links ]
58. Clark PU, Dyke AS, Shakun JD, et al. The last glacial maximum. Science. 2011;325:710-714. doi:10.1126/science.1172873, PMid:19661421 [ Links ]
59. Fairbanks RG. A 17,000 year glacio-eustatic sea level record: Influence of glacial melting rate on the Younger Dryas event and deep-ocean circulation. Nature. 1989;342:637-642. doi:10.1038/342637a0 [ Links ]
60. Dingle RV, Rogers J. Effects of sea-level changes on the Pleistocene palaeoecology of the Agulhas Bank. Palaeoecol Afr. 1972;6:55-58. [ Links ]
61. Romero O, Mollenhauer G, Schneider RR, Wefer G. Oscillations of the siliceous imprint in the central Benguela Upwelling System from MIS 3 through to the early Holocene: The influence of the Southern Ocean. J Quaternary Sci. 2003;18:733-743. doi:10.1002/jqs.789 [ Links ]
62. Hutson WH. The Agulhas Current during the Late Pleistocene: Analysis of modern faunal analogs. Science. 1980;207:64-66. doi:10.1126/ science.207.4426.64, PMid:17730815 [ Links ]
63. Beckley LE. The ichthyoplankton assemblage of the Algoa Bay nearshore region in relation to coastal zone utilization by juvenile fish. S Afr J Zool. 1985;21:244-252. [ Links ]
64. Slatkin M. Gene flow in natural populations. Ann Rev Ecol Syst. 1985;16:393-430. doi:10.1146/annurev.es.16.110185.002141 [ Links ]
65. Tankard AJ. Thermally anomalous late Pleistocene mollusks from the south-western Cape Province, South Africa. Ann S Afr Mus. 1975;69:17-45. [ Links ]
66. Henshilwood C, d'Errico F, Vanhaeren M, Van Niekerk K, Jacobs Z. Middle Stone Age beads from South Africa. Science. 2004;304:404. doi:10.1126/science.1095905, PMid:15087540 [ Links ]
67. Von der Heyden S, Lipinski MR, Matthee CA. Mitochondrial DNA analyses of the Cape hakes reveal an expanding, panmictic population for Merluccius capensis and population structuring for mature fish in Merluccius paradoxus. Mol Phylogenet Evol. 2007;42:517-527. doi:10.1016/j. ympev.2006.08.004, PMid:16982203 [ Links ]
68. Von der Heyden S, Lipinski MR, Matthee CA. Species specific genetic markers for identification of early life history stages of Cape hakes, Merluccius capensis and M. paradoxus in the southern Benguela Current. J Fish Biol. 2007;70:262-268. doi:10.1111/j.1095-8649.2007.01409.x [ Links ]
69. Florence WK, Hulley RA, Stewart BA, Gibbons MJ. Genetic and morphological variation of the lanternfish Lampanyctodes hectoris (Myctophiformes: Myctophidae) off southern Africa. S Afr J Mar Sci. 2002;24:193-203. doi:10.2989/025776102784528466 [ Links ]
70. Shaw PW, Hendrickson L, McKeown NJ, Stonier T, Naud MJ, Sauer WHH. Discrete spawning aggregations of loliginid squid do not represent genetically distinct populations. Mar Ecol Prog Ser. 2010;408:117-127. doi:10.3354/meps08593 [ Links ]
71. Flint NS, Van der Bank FH, Theron PJ, Staby A. Genetic variation in two populations of the orange roughy (Hoplostethus atlanticus) from Namibia. S Afr J Aqua Sci. 1998/99;24:71-83. [ Links ]
72. Von der Heyden S, Groeneveld JC, Matthee CA. Long current to nowhere? - Genetic connectivity of Jasus tristani populations in the southern Atlantic Ocean. Afr J Mar Sci. 2007;29:491-497. doi:10.2989/AJMS.2007.29.3.15.345 [ Links ]
73. Matthee CA, Cockcroft AC, Gopal K, Von der Heyden S. Mitochondrial DNA variation of the west-coast rock lobster, Jasus lalandii: Marked genetic diversity differences among sampling sites. Mar Freshwater Res. 2007;58:1130-1135. doi:10.1071/MF07138 [ Links ]
74. Tolley KA, Groeneveld J, Gopal K, Matthee CA. Mitochondrial DNA panmixia in spiny lobster Palinurus gilchristi suggests a population expansion. Mar Ecol Prog Ser. 2005;297:225-231. doi:10.3354/meps297225 [ Links ]
75. Benzie JAH, Ballment E, Forbes AT, et al. Mitochondrial DNA variation in Indo-Pacific populations of the giant tiger prawn, Penaeus monodon. Mol Ecol. 2002;11:2553-2569. doi:10.1046/j.1365-294X.2002.01638.x, PMid:12453239 [ Links ]
76. Stow A, Zenger K, Briscoe D, et al. Isolation and genetic diversity of endangered grey nurse shark (Carcharias taurus) populations. Biol Lett. 2006;2:308-311. doi:10.1098/rsbl.2006.0441, PMid:17148390, PMCid:1618890 [ Links ]
77. Zemlak TS, Ward RD, Connell AD, Holmes BH, Hebert PDN. DNA barcoding reveals overlooked marine fishes. Mol Ecol Res. 2009;9:237-242. doi:10.1111/j.1755-0998.2009.02649.x [ Links ]
78. Waters JM, Roy MS. Out of Africa: The slow train to Australasia. Syst Biol. 2004;53:18-24. doi:10.1080/10635150490264671, PMid:14965897 [ Links ]
79. Matthee CA, Fourie F, Oosthuizen WH, Meyer MA, Tolley KA. Mitochondrial sequence data of the Cape fur seal (Arctocephalus pusillus pusillus) suggest that population numbers may be affected by climatic shifts. Mar Biol. 2005;148:900-905. [ Links ]
80. Ridgway T, Hoegh-Guldberg O, Ayre DJ. Panmixia in Pocillopora verrucosa from South Africa. Mar Biol. 2001;139:175-181. doi:10.1007/s002270100573 [ Links ]
81. Teske PR, Hamilton H, Palsbøll PJ, et al. Molecular evidence for long-distance colonization in an Indo-Pacific seahorse lineage. Mar Ecol Prog Ser. 2005;286:249-260. doi:10.3354/meps286249 [ Links ]
82. Fratini S, Vannini M. Genetic differentiation in the mud crab Scylla serrata (Decapoda: Portunidae) within the Indian Ocean. J Exp Mar Biol Ecol. 2002;272:103-116. doi:10.1016/S0022-0981(02)00052-7 [ Links ]
83. Ragionieri L, Cannicci S, Schubart CD, Fratini S. Gene flow and demographic history of the mangrove crab Neosarmatium meinerti: A case study from the western Indian Ocean. Estuar Coastal Shelf Sci. 2010;86:179-188. doi:10.1016/j.ecss.2009.11.002 [ Links ]
84. Silva IC, Mesquita N, Paula J. Lack of population structure in the fiddler crab Uca annulipes along an East African latitudinal gradient: Genetic and morphometric evidence. Mar Biol. 2010;157:1113-1126. doi:10.1007/ s00227-010-1393-9 [ Links ]
85. Ridgway T, Sampayo EM. Population genetic status of the Western Indian Ocean: What do we know? Western Indian Ocean J Mar Sci. 2005;4:1-9. [ Links ]
86. Grant WS, Leslie RW. Inter-ocean dispersal is an important mechanism in the zoogeography of hakes (Pisces: Merluccius spp.). J Biogeogr. 2001;28:699-721. doi:10.1046/j.1365-2699.2001.00585.x [ Links ]
87. Grant WS, Bowen BW. Living in a tilted world: Climate change and geography limit speciation in Old World anchovies (Engraulis: Engraulidae). Biol J Linn Soc. 2006;88:673-689. doi:10.1111/j.1095- 8312.2006.00651.x [ Links ]
88. D'Amato ME, Harkins GW, De Oliveira T, Teske PR, Gibbons MJ. Molecular dating and biogeography of the neritic krill Nyctiphanes. Mar Biol. 2008;155:243-247. doi:10.1007/s00227-008-1005-0 [ Links ]
89. Teske PR, Oosthuizen A, Papadopoulos I, Barker NP. Phylogeographic structure of South African Octopus vulgaris revisited: Identification of a second lineage near Durban harbour. Mar Biol. 2007;151:2119-2122. doi:10.1007/s00227-007-0644-x [ Links ]
90. Palero F, Lopes J, Abello P, Macpherson E, Pascual M, Beaumont MA. Rapid radiation in spiny lobsters (Palinurus spp.) as revealed by classic and ABC methods using mtDNA and microsatellite data. BMC Evol Biol. 2009;9:263. doi:10.1186/1471-2148-9-263, PMid:19900277, PMCid:2777881 [ Links ]
91. Rius M, Teske PR. A revision of the Pyura stolonifera species complex (Tunicata, Ascidiacea), with a description of a new species from Australia. Zootaxa. 2011;2754:27-40. [ Links ]
92. Felsenstein J. Accuracy of coalescent likelihood estimates: Do we need more sites, more sequences, or more loci? Mol Biol Evol. 2006;23:691-700. doi:10.1093/molbev/msj079, PMid:16364968 [ Links ]
93. Irwin DE. Phylogeographic breaks without geographic barriers to gene flow. Evolution. 2002;56:2383-2394. doi:10.1111/j.0014-3820.2002. tb00164.x, PMid:12583579 [ Links ]
94. Stephens M, Smith N, Donnelly P. A new statistical method for haplotype reconstruction from population data. Am J Hum Genet. 2001;68:978-989. doi:10.1086/319501, PMid:11254454 [ Links ]
95. Flot J-F. CHAMPURU 1.0: A computer software for unraveling mixtures of two DNA sequences of unequal lengths. Mol Ecol Notes. 2007;7:974-977. doi:10.1111/j.1471-8286.2007.01857.x [ Links ]
96. Jarman SN, Ward RD, Elliott NG. Oligonucleotide primers for PCR amplification of coelomate introns. Mar Biotech. 2002;4:347-355. doi:10.1007/s10126-002-0029-6, PMid:14961246 [ Links ]
97. Teske PR, Beheregaray LB. Intron-spanning primers for the amplification of the nuclear ANT gene in decapod crustaceans. Mol Ecol Res. 2009;9:774-776. doi:10.1111/j.1755-0998.2009.02534.x [ Links ]
98. Chow S, Hazama K. Universal PCR primers for S7 ribosomal gene introns in fish. Mol Ecol. 1998;7:1255-1256. PMid:9734083 [ Links ]
99. Rosenblum EB, Belfiore NM, Moritz C. Anonymous nuclear markers for the eastern fence lizard, Sceloporus undulatus. Mol Ecol Notes. 2007;7:113-116. doi:10.1111/j.1471-8286.2006.01547.x [ Links ]
100. Teske PR, Forget FRG, Cowley PD, Beheregaray LB. Microsatellite markers for the roman, Chrysoblephus laticeps (Teleostei: Sparidae), an overexploited seabream from South Africa. Mol Ecol Res. 2009;9:1162-1164. doi:10.1111/j.1755-0998.2009.02595.x [ Links ]
101. Galbusera PHA, Gillemot S, Jouk P, Teske PR, Hellemans B, Volckaert FAMJ. Microsatellite markers developed for the endangered Knysna seahorse, Hippocampus capensis, and their use in the detection of a genetic bottleneck. Mol Ecol Notes. 2007;7:638-640. doi:10.1111/j.1471- 8286.2007.01688.x [ Links ]
102. Dos Santos SMR, Klopper AW, Oosthuizen CJ, Bloomer P. Isolation and characterization of polymorphic tetranucleotide microsatellite loci in the pelagic perciform fish Pomatomus saltatrix (Linnaeus, 1766) from South Africa. Mol Ecol Res. 2008;8:1065-1067. doi:10.1111/j.1755- 0998.2008.02156.x [ Links ]
103. Chopelet J, Helyar S, Mann B, Mariani S. Novel polymorphic microsatellite loci for the protogynous hermaphrodite slinger sea bream (Chrysoblephus puniceus, Sparidae). Mol Ecol Res. 2009;9:1223-1226. doi:10.1111/j.1755- 0998.2009.02615.x [ Links ]
104. Bester-van der Merwe AE, Roodt-Wilding R, Volckaert FAM, D'Amato ME. Historical isolation and hydrodynamically constrained gene flow in declining populations of the South-African abalone, Haliotis midae. Conserv Genet. 2011;12(2):543-555. doi:10.1007/s10592-010-0162-0 [ Links ]
105. Bensch S, Akesson M. Ten years of AFLP in ecology and evolution: Why so few animals? Mol Ecol. 2005;14:2899-2914. doi:10.1111/j.1365- 294X.2005.02655.x, PMid:16101761 [ Links ]
106. Badenhorst D, Roodt-Wilding R. Application of various DNA extraction methodologies on abalone, Haliotis midae, larvae and juveniles for fluorescent AFLP analysis. Aquaculture Res. 2007;38:546-549. doi:10.1111/j.1365-2109.2007.01698.x [ Links ]
107. Griffiths CL, Robinson TB, Lange L, Mean A. Marine biodiversity in South Africa: An evaluation of current states of knowledge. PLoS ONE. 2010;5:e12008. doi:10.1371/journal.pone.0012008, PMid:20689849, PMCid:2914023 [ Links ]
108. Rouault M, Penven P, Pohl B. Warming in the Agulhas Current system since the 1980s. Geophys Res Lett. 2009;36:L12602. doi:10.1029/2009GL037987 [ Links ]
109. Lin H-C, Sanchez-Ortiz C, Hastings PA. Colour variation is incongruent with mitochondrial lineages: Cryptic speciation and subsequent diversification in a Gulf of California reef fish (Teleostei: Blennioidei). Mol Ecol. 2009;18:2476-2488. doi:10.1111/j.1365-294X.2009.04188.x, PMid:19389167 [ Links ]
110. Von der Heyden S. 'Carry on sampling' - assessing marine fish biodiversity and discovery rates in southern Africa. Divers Distrib. 2011;17:81-92. doi:10.1111/j.1472-4642.2010.00712.x [ Links ]
111. Beerli P. How to use MIGRATE or why are Markov chain Monte Carlo programs difficult to use? In: Bertorelle G, Bruford MW, Hauïe HC, Rizzoli A, Vernesi C, editors. Population genetics for animal conservation, volume 17 of conservation biology. Cambridge: Cambridge University Press, 2009; p. 42-79. [ Links ]
112. Hey J, Nielsen R. Integration within the Felsenstein equation for improved Markov chain Monte Carlo methods in population genetics. Proc Natl Acad Sci USA. 2007;104:2785-2790. doi:10.1073/pnas.0611164104, PMid:17301231, PMCid:1815259 [ Links ]
113. Hickerson MJ, Stahl E, Takebayashi N. msBayes: Pipeline for testing comparative phylogeographic histories using hierarchical approximate Bayesian computation. BMC Bioinf. 2007;8:268. doi:10.1186/1471-2105-8- 268, PMid:17655753, PMCid:1949838 [ Links ]
114. Beerli P, Palczewski M. Unified framework to evaluate panmixia and migration direction among multiple sampling locations. Genetics. 2010;185:313-326. doi:10.1534/genetics.109.112532, PMid:20176979, PMCid:2870966 [ Links ]
115. Blastoch A, Boning CW, Schwarzkopf FU, Lutjeharms JRE. Increase in Agulhas leakage due to poleward shift of southern hemisphere westerlies. Nature. 2009;462:495-498. doi:10.1038/nature08519, PMid:19940923 [ Links ]
116. Lutjeharms JRE, Durgadoo JV, Schapira M, McQuaid CD. First oceanographic survey of the entire continental shelf adjacent to the northern Agulhas Current. S Afr J Sci. 2010;106(9/10):7-9. [ Links ]
 Correspondence to:
Correspondence to:
Peter Teske
Postal address: Molecular Ecology and Systematics Group, Botany Department, Rhodes University
Grahamstown 6140, South Africa
Email: pteske101@gmail.com
Received: 10 Nov. 2010
Accepted: 16 Mar. 2011
Published: 26 May 2011
© 2011. The Authors. Licensee: OpenJournals Publishing. This work is licensed under the Creative Commons Attribution License.














