Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Science
On-line version ISSN 1996-7489
Print version ISSN 0038-2353
S. Afr. j. sci. vol.106 n.11-12 Pretoria Nov./Dec. 2010
http://dx.doi.org/10.4102/sajs.v106i11/12.211
RESEARCH LETTER
Characterisation of Shigella species isolated from river catchments in the North West province of South Africa
Constance Wose KingeI; Moses MbeweII
ISchool of Environmental & Health Sciences, North-West University, Mafikeng campus, South Africa. email: conskinge@yahoo.ca
IISchool of Agricultural Sciences, North-West University, Mafikeng campus, South Africa
ABSTRACT
The occurrence and distribution of Shigella species in water from the five river catchments in the North West province of South Africa were investigated. Shigella is a Gram-negative, non-motile, facultative anaerobic bacillus that causes shigellosis, an important cause of morbidity and mortality in high-risk populations (such as children, the elderly and immuno-compromised individuals) that depend on river water. A total of 54 water samples collected in winter (April 2007 to July 2007) and summer (December 2007 to March 2008) were cultured on Salmonella-Shigella agar by the spread-plate method. Suspected Shigella isolates obtained were characterised by primary biochemical (Triple Sugar Iron agar and agglutination) and molecular (polymerase chain reactions, PCR) tests. Amplification of the invasion plasmid gene (ipaH) by PCR was done to confirm the presence of Shigella spp. in water. In total, 214 Shigella boydii, 15 Shigella dysenteriae, 11 Shigella flexneri and 2 Shigella sonnei were confirmed by serotyping in both winter and summer samples. The ipaH gene (606 bp) was present in 176 and 49 of the winter and summer isolates, respectively. The presence of Shigella spp. in water was confirmed with over 90% specificity. The need for more effective management of these river catchments and the provision of potable water and sanitation facilities is needed to minimise the occurrence and transmission of water-borne diseases caused by these and other pathogenic bacteria.
Keywords: polymerase chain reaction; ipaH gene; river catchments; serotyping; Shigella species
INTRODUCTION
Shigella species are Gram-negative, non-motile, non-encapsulated, non-lactose fermenting, facultative anaerobes that are pathogenic to humans.1 Their mode of transmission is usually through the ingestion of contaminated food and water, as well as person-to-person contact.2 The genus Shigella consists of four species and at least 47 serotypes, which include Shigella dysenteriae, Shigella flexneri, Shigella boydii and Shigella sonnei.2,3 They are the major cause of shigellosis or bacillary dysentery affecting, in particular, immuno-compromised individuals, children and the elderly.1 Worldwide, the disease burden is estimated to be 150 million cases, with 1 million deaths per year in the developing world.4 In 2009, the disease burden in South Africa was 1812 cases for both invasive and non-invasive shigellosis, predominantly in children below the age of 5 years.4
Shigellosis is characterised by destruction of the colonic epithelium as a result of an inflammatory response induced upon bacterial invasion of the mucosa.5 Differentiation of Shigella isolates is classically based on serotyping and biochemical assays.1 Antigenic characterisation relies exclusively on their somatic (O) antigens because they produce neither flagellins nor capsular antigens.5 Biosynthesis of the O antigen is carried out by genes located in the galF and gnd clusters. These genes are classified into three different groups, (1) genes for the synthesis of nucleotide sugars specially used as O antigen residues, (2) genes encoding sugar transferases and (3) O unit processing genes normally consisting of O unit flippase gene (wzx) and polymerase gene (wzy).6 Based on their antigenic properties, 46 Shigella serotypes have been recognised but, with the exception of S. boydii, all Shigella serotypes show more than 73% DNA relatedness to Escherichia coli K-12.6,7
Molecular techniques, such as restriction of amplified O-antigen gene cluster loci,7 pulsed field gel electrophoresis, PCR, DNA hybridisation, ribotyping, and multilocus enzyme electrophoresis-mdh gene sequencing and their combinations, have been used to characterise Shigella species.8,9,10,11,12 Several PCR protocols using different molecular markers have been used for the detection of Shigella species and other related organisms in environmental, faecal and food samples.1,2,13,14 The invasion plasmid antigen H (ipaH) gene, a virulence gene whose protein product is necessary for invasion of colonic epithelial cells and also for the detection of Shigella in the environment, was used because it is carried by all four Shigella species as well as enteroinvasive E. coli (EIEC).15
Incidences of shigellosis outbreaks as a result of poor water quality have been reported throughout the world.16 In Sierra Leone, a dysentery outbreak caused by S. dysenteriae type 1 and S. flexneri was reported in the Moyamba and Koinadugu districts. Reports provided by the Ministry of Health gave a total of 3094 cases of shigellosis with 132 deaths from 6 December 1999 to 16 January 2000.16 In 2003, 379 cases of shigellosis and 23 deaths were reported in Paoua and Bazoum, north-west of Bangui in the Central African Republic.17 In that same year, a cholera outbreak was also reported in Monrovia, Liberia, affecting 1857 people.17 A year later, North Darfur (Sudan) reported a shigellosis outbreak caused by S. dysenteriae type 1, with 1340 cases of bloody diarrhoea and 11 deaths.18 South Africa is no exception, with incidences of diarrhoea outbreaks caused by S. flexneri and S. sonnei reported in the Mpumalanga, Limpopo and Northern Cape provinces.19 While shigellosis outbreaks have not been reported in the North West province, the possibility of such an occurrence cannot be underestimated as other water-borne diseases such as cholera and typhoid have been previously reported.16,20,21 Therefore with no available information of shigellosis outbreaks in the North West province, this study aims to create awareness of the presence of Shigella spp. in river water and preparedness for any potential full-blown disease outbreak.
MATERIALS AND METHODS
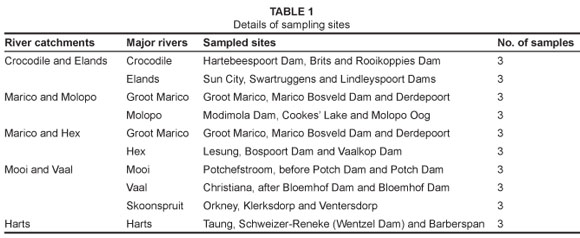
A total of 54 water samples were collected in 500-mL sterile Schott Duran bottles from the major rivers in the Crocodile and Elands, Marico and Molopo, Marico and Hex, Mooi and Vaal, and Harts river catchments in the North West province (Table 1). Water samples were collected from three sites from each river during the months of April 2007 to July 2007 (winter) and December 2007 to March 2008 (summer). Samples were transported to the Microbiology Research Laboratory of the North-West University.
Isolation of Shigella
Ten-fold serial dilutions of water samples were performed using 2% Buffered Peptone water (Biolab, Merck Diagnostics, South Africa). Aliquots of 1 mL of each dilution were cultured on Salmonella-Shigella agar by the spread-plate method.22 Plates were incubated at 37 ºC for 24 h.13 Suspected isolates were picked from culture plates and subjected to further analysis for the identification and isolation of possible Shigella isolates.
Bacterial strains
Shigella boydii strain ATCC® 9207, Shigella sonnei strain ATCC® 25931, Shigella flexneri strain ATCC® 12022 and Shigella dysenteriae strain ATCC® 49345 were used as positive controls.23 Because the ipaH gene is also present in EIEC, E. coli strain ATCC® 25922 was used as a negative control.24
Biochemical characterisation
Triple Sugar Iron agar test
All isolates were characterised using Triple Sugar Iron (TSI) agar to test for glucose, sucrose and lactose fermentation. Bergey's manual of systematic bacteriology25 was used as an identification aid.
Agglutination test
A total of 235 and 287 isolates, for winter and summer respectively, were presumptively identified as Shigella spp. by the TSI agar test and then subjected to serotyping by slide agglutination. Serotyping was done using S. boydii Poly C, C1, C2 and C3; S. dysenteriae Poly A Types 1, 2, 3, 4, 5, 6 and 7; S. sonnei Poly D Phase I and II and S. flexneri Poly B Types I, II, III, IV, V and VI antisera according to the manufacturer's instructions (Davies Diagnostics, Johannesburg, South Africa).
Molecular characterisation
Genomic DNA extraction
Bacterial genomic DNA was extracted as described previously.26 The concentration of the extracted DNA in solution was determined spectrophotometrically (UV Visible spectrophotometer model S-22, Boeco, Germany) at a wavelength of 260 nm and the purity was measured at 280 nm as previously described.27 The integrity of the purified template DNA was assessed by conventional 0.8% (w/v) agarose gel.
Polymerase chain reactions (PCR)
PCRs were performed in final volumes of 25µL, with a reaction mixture containing 0.2µg/µL of the prepared template DNA, 50 pmol of each oligonucleotide primer, 1X PCR master mix and Dnase-Rnase free distilled water (Inqaba Biotechnological Industries Pty Ltd, Pretoria, South Africa). The primers IpaH-F 5-'CCTTGACCGCCTTTCCGATA-3' and IpaH-R 5'-CAGCCACCCTCTGAGGTACT-3' (Inqaba Biotechnological Industries Pty Ltd, Pretoria, South Africa) were used. The amplifications were performed using a Peltier Thermal Cycler (model PTC-220 DYADTM DNA ENGINE; MJ Research Inc., Waltham, MA, USA) under the PCR conditions described previously.28 PCR-amplified DNA fragments were separated by electrophoresis on 1% agarose. A wide-range molecular weight DNA marker (100-bp ladder, Inqaba Biotechnological Industries Pty Ltd) was used on each gel as the standard. The gels were stained in ethidium bromide (0.1µg/mL) for 2 h. PCR products were visualised using a GeneSnap Bio-Imaging System (SYNGENE model GBOX CHEMI HR) and the images captured using image acquisition software version 6. 08. 04.
RESULTS
Based on the TSI agar test, 235 and 287 presumptively positive Shigella isolates were identified for the winter and summer samples, respectively. Among the 235 positive isolates obtained for winter, serological analysis confirmed a total of 168 S. boydii, 11 S. dysenteriae, 9 S. flexneri and 1 S. sonnei. The results indicated the presence of S. boydii and S. dysenteriae in all five catchments, whereas S. flexneri was present in only three catchments and S. sonnei in only one catchment (Table 2). Similarly, of the 287 Shigella isolates identified by the TSI test in the summer samples, serological analysis confirmed the presence of 44 S. boydii, 4 S. dysenteriae, 2 S. flexneri and 1 S. sonnei. The results indicated the presence of S. boydii in all five catchments and S. dysenteriae in two catchments, S. flexneri and S. sonnei were both present in the Mooi and Vaal catchment only (Table 3).
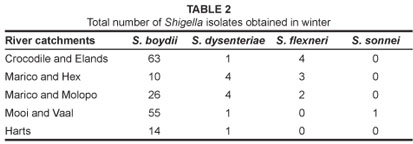

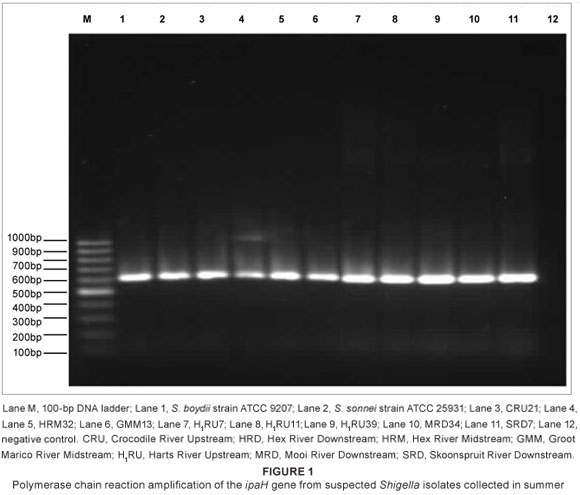
The high S. boydii contamination levels in the winter samples in all catchments are of concern. Levels were as high as 88% and 79% in the Harts and Crocodile and Elands catchments, respectively, whilst levels in the other three catchments were between 50% and 74%. For the summer samples, S. boydii contamination levels were highest (34%) in the Mooi and Vaal catchment followed by the Crocodile and Elands catchment (21%). The level in the Harts catchment was 13% while the Mooi and Vaal catchment and the Marico and Hex catchment recorded less than 10% agglutination each. All 189 and 51 isolates confirmed by serotyping to be Shigella spp. for winter and summer seasons, respectively, were subjected to PCR for the detection of the ipaH gene. The ipaH gene (606 bp) was present in 176 of the winter and 49 of the summer isolates. Figure 1 presents the positive results of PCR detection of the ipaH gene for a few of the isolates collected in summer. The PCR confirmed the presence of Shigella in river water and showed 93% and 96% specificity for winter and summer Shigella isolates, respectively.
DISCUSSION
Culture methods have traditionally been used to identify Shigella species; however these methods have proved less effective because they only detect a small fraction of the actual number of shigellosis cases, probably because of a low number of causative organisms, competition from other commensal organisms and changes in the ambient temperature during sample collection.15,29 To optimise the detection rates, we focused on the use of conventional serotyping and PCR typing methods to detect Shigella in river waters. In similar studies, the ipaH gene was used as an indicator to detect the presence of Shigella in environmental waters.23,28,30,31 Although the PCR assay is rapid and highly specific, limitations such as false positives or negatives also exist because of the presence of impurities.31 However, DNA extracted from all suspected isolates and control strains used in the study was checked for the presence of any impurity by measuring the optical density at 280 nm and 260 nm. The ratio of the two optical densities was calculated and compared with that of the standard values, the presence of any impurity was thereby eliminated. The PCR assay used was highly sensitive and the presence of Shigella spp. using 0.2µg/µL of DNA was confirmed with over 90% specificity.
Generally, river water harbours a vast majority of enteropathogens derived from municipal sewage discharges, rainfall runoffs from agricultural farms and faecal waste from humans, pets, farm animals and wildlife, because river water is untreated.32 Shigella spp. are sensitive to chlorination at normal levels and can survive for only up to 4 days in river water. In addition, it takes 1 to 2 days to establish a laboratory diagnosis by culture of a patient specimen, making the recovery of Shigella from environmental samples almost impossible.33 However, the findings of this and previous studies2,13 detected Shigella in surface waters and sewage samples, which is an indication that surface waters can possibly transport Shigella strains and that there is a possibility of a continuous source of contamination into the rivers. The detection of Shigella bacteria in the water samples suggests the likelihood of other pathogenic bacteria and confirms the presence of faecal contamination in the river catchments sampled.
Although Shigella bacteria predominantly cause food-borne diseases, shigellosis outbreaks as a result of consumption of contaminated water have also been reported, especially in developing countries with inadequate sanitation facilities.20,21,23 Shigella epidemics spread through contaminated food and water. It is not known what dose of Shigella is necessary to cause disease, but the transmission of the pathogen is believed to be facilitated by a very low infectious dose.34 In the North West province of South Africa, the percentage of households with sustainable access to an improved water source has steadily improved from 78% in 2001 to 97% in 2005.35 However, an approximate 1 million people are still without access to basic water supply – mostly those in rural villages, farmlands and informal settlements – and still rely on surface water resources, which exposes them to a high risk of contracting water-borne diseases.36
CONCLUSIONS AND RECOMMENDATION
The presence of Shigella spp. in river water, as demonstrated by this and other studies, is significant and may be of public health concern. The chromosomally located multicopy virulence gene, ipaH, which is also known for its role in producing invasive characteristics, was found in all of the Shigella strains identified. Therefore, PCR screening of environmental samples for the ipaH gene should provide a better indicator of the possible presence of Shigella. We have previously shown37 that river water in the North West province is a possible route of human contamination with pathogenic bacteria such as Shigella, by direct or indirect consumption of river water through its use for recreation, domestic and agricultural purposes. The need for more effective management of these river catchments and the provision of potable water and sanitation facilities is needed to minimise the occurrence and transmission of water-borne diseases.
REFERENCES
1. Penatti MPA, Hollanda LM, Nakazato G, et al. Epidemiological characterization of resistance and PCR typing of Shigella flexneri and Shigella sonnei strains isolated from bacillary dysentery cases in Southeast Brazil. Braz J Med Biol Res. 2007;40(2):249–258. [ Links ]
2. Theron J, Morar D, Preez M, Brözel VS, Venter SN. A sensitive seminested PCR for the detection of Shigella in spiked environmental samples. Water Res. 2001;35(4):869–874. [ Links ]
3. Kingombe CI, Cerqueira-Campos ML, Farber JM. Molecular strategies for the detection, identification, and differentiation between enteroinvasive Escherichia coli and Shigella species. J Food Prot. 2005;68:7549-7553. [ Links ]
4. Keddy K. The group for enteric, respiratory and meningeal disease surveillance in South Africa. Communicable Disease Surveillance Bulletin [serial online]. May 2010;8(2):25–27. [ Links ]
5. Parsot C. Shigella spp. and enteroinvasive Escherichia coli pathogenicity factors. FEMS Microbiol Lett. 2005;252(1):11-18. [ Links ]
6. Liu B, Senchenkova SN, Feng L, et al. Structural and molecular characterisation of Shigella boydii type 16 O antigen. Gene. 2006;380:46-53. [ Links ]
7. Coimbra RS, Grimont F, Grimont PAD. Identification of Shigella serotypes by restriction of amplified O-antigen gene cluster. Res Microbiol. 1999;150:543–553. [ Links ]
8. Rompré A, Servais P, Baudart J, de-Roubin M-R, Laurent P. Detection and enumeration of coliforms in drinking water: Current methods and emerging approaches. J Microbiol Methods. 2002;49:31–34. [ Links ]
9. Rolland K, Lambert-Zechovsky N, Picard B, Denamur E. Shigella and enteroinvasive Escherichia coli strains are derived from distinct ancestral strains of E. coli. Microbiol. 1998;144:2667–2672. [ Links ]
10. Pupo GM, Karaolis DK, Lan R, Reeves PR. Evolutionary relationships among pathogenic and non-pathogenic Escherichia coli strains inferred from multilocus enzyme electrophoresis and mdh sequence studies. Infect Immun. 1997;65:2685–2692. [ Links ]
11. Pupo GM, Lan R, Reeves PR. Multiple independent origins of Shigella clones of Escherichia coli and convergent evolution of many of their characteristics. Proc Natl Acad Sci USA. 2000;97:10567–10572. [ Links ]
12. Lee TM, Chang CY, Chang LL, Chen WM, Wang TK, Chang SF. One predominant type of genetically closely related Shigella sonnei prevalent in four sequential outbreaks in school children. Diagn Microbiol Infect Dis. 2003;45:173–181. [ Links ]
13. Moganedi KLM, Goyvaerts EMA, Venter SN, Sibara MM. Optimisation of the PCR-invA primers for the detection of Salmonella in drinking and surface waters following a pre-cultivative step. Water SA. 2007;33(2):195–201. [ Links ]
14. Faruque SM, Khan R, Kammuzzaman M, et al. Isolation of Shigella dysenteriae type 1 and S. flexneri strains from surface water in Bangladesh; comparative molecular analysis of environmental Shigella versus clinical strains. Appl Environ Microbiol. 2002;68(8):3908–3913. [ Links ]
15. Thiem VD, Sethabutr O, Seidlein L, et al. Detection of Shigella by a PCR assay targeting the ipaH gene suggests increased prevalence of shigellosis in Nha Trang,Vietnam. J Clin Microbiol. 2004;42(5):2031–2035. [ Links ]
16. World Health Organization (WHO). Considerations in evaluating the cost-effectiveness of environmental health interventions. WHO/SDE/WSH/00.10. Geneva: WHO; 2000. [ Links ]
17. World Health Organization (WHO). Quantifying selected major risks to health. The world health report 2002. Geneva: WHO; 2003. [ Links ]
18. World Health Organization (WHO). Evaluation of the costs and benefits of water and sanitation improvements at the global level. WHO/SDE/WSH/04.04. Geneva: WHO; 2004. [ Links ]
19. National Institute for Communicable Diseases. Diarrhoeal disease outbreak-school trip. Communicable Diseases Communiqué. 2008;7(10):1–2. [ Links ]
20. National Department of Health. Cholera outbreak in Mpulanga, Issues: Cholera. Archive of cholera updates. C2003 [cited 2003 Dec 31]. Available from: http://www.doh.gov.za/docs/reports/2003/cholera/mpumalanga.html [ Links ]
21. Outbreak Report, Department of Health. Diarrhoea and typhoid outbreak in Delmas under control. Department of Health, September 2005. Available from: http://www.doh.gov.za/docs/pr/2005/pr0926.html [ Links ]
22. Standing Committee of Analysts. The microbiology of drinking water 2002. Part 7: The enumeration of heterotrophic bacteria by pour and spread plate techniques. Methods for the examination of waters and associated materials. London: Environment Agency; 2002. [ Links ]
23. Wang R-F, Cao W-W, Cerniglia CE. Phylogenetic analysis and identification of Shigella species by molecular probes. Mol Cell Probes. 1997;11:427–432. [ Links ]
24. Lu J-J, Perng C-L, Lee S-Y, Wan C-C. Use of PCR with universal primers and restriction endonuclease digestions for detection and identification of common bacterial pathogens in cerebrospinal fluid. J Clin Microbiol. 2000;38(6):2076–2080. [ Links ]
25. Garrity GM, Boone DR, Castenholz RW, Bergey DH. Bergey's manual of systematic bacteriology. 2nd ed. New York: Springer, 2001. [ Links ]
26. Doyle JJ, Doyle JL. A rapid total DNA preparation procedure for fresh plant tissue. Focus. 1990;12:13–15. [ Links ]
27. Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: A laboratory manual. 2nd ed. New York: Cold Spring Harbor Laboratory Press; 1989. [ Links ]
28. Kong RYC, Lee SKY, Law TWF, Law SHW, Wu RSS. Rapid detection of six types of bacteria pathogens in marine waters by multiplex PCR. Water Res. 2002;36(11):2802–2812. [ Links ]
29. Taylor WI, Schelhart D. Effect of temperature on transport and plating media for enteric pathogens. J Clin Microbiol. 1975;2:281–286. [ Links ]
30. Obi CL, Bessong PO, Momba MNB, Potgieter N, Samie A, Igumbor EO. Profiles of antibiotic susceptibilities of bacterial isolates and physico-chemical quality of water supply in rural Venda communities, South Africa. Water SA. 2004;30(4):515–520. [ Links ]
31. Rompré A, Servais P, Baudart J, de-Roubin M-R, Laurent P. Detection and enumeration of coliforms in drinking water: Current methods and emerging approaches. J Microbiol Methods. 2002;49:31–34. [ Links ]
32. Arvanitidou M, Kanellou K, Vagiona DG. Diversity of Salmonella species and fungi in northern Greek rivers and their correlation to faecal pollution indicators. Env Res. 2005;99:278–284. [ Links ]
33. Eaton AD, Clesceri LS, Rice EW, Greenberg AE, editors. Standard methods for the examination of water and wastewater. 21st ed. American Public Health Association, Washington, DC. 2005; p. 118–119. [ Links ]
34. Shears P. Shigella infections. Ann Trop Med Parasitol. 1996;90:105–114. [ Links ]
35. Northwest Province Environment Outlook. A report on the state of the environment, 2008 [homepage on the Internet]. No date [cited 2010 Aug 24]. Available from: http://www.nwpg.gov.za [ Links ]
36. Northwest Department of Agriculture, Conservation and Environment. Development of an ambient environmental monitoring programme for the Northwest Province, 2007. Report to the Northwest Province Department of Agriculture, Conservation and Environment. Northwest Department of Agriculture, Conservation and Environment (NWDACE), 2007. [ Links ]
37. Wose Kinge CN, Ateba CN, Kawadza DT. Antibiotic resistance profiles of Escherichia coli isolated from different water sources in the Mmabatho locality, Northwest Province, South Africa. S Afr J Sci. 2010;106(1/2):1-6. [ Links ]
 Postal address:
Postal address:
Department of Biological Sciences, School of Environmental & Health Sciences
North-West University, Mafikeng campus
Private Bag X2046
Mmabatho 2735, South Africa
Received: 08 Apr. 2010
Accepted 28 Aug. 2010
Published: 11 Nov. 2010
Correspondence to: Constance Wose Kinge
This article is available at: http://www.sajs.co.za
© 2010. The Authors. Licensee: OpenJournals Publishing. This work is licensed under the Creative Commons Attribution License.














