Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Science
On-line version ISSN 1996-7489
Print version ISSN 0038-2353
S. Afr. j. sci. vol.106 n.7-8 Pretoria Jul./Aug. 2010
RESEARCH LETTER
Optimisation of automated ribosomal intergenic spacer analysis for the estimation of microbial diversity in fynbos soil
Etienne SlabbertI; Carel J. van HeerdenII; Karin JacobsI
IDepartment of Microbiology, Stellenbosch University, South Africa
IICentral Analytical Facility, Stellenbosch University, South Africa
ABSTRACT
Automated ribosomal intergenic spacer analysis (ARISA) has become a commonly used molecular technique for the study of microbial populations in environmental samples. The reproducibility and accuracy of ARISA, with and without the polymerase chain reaction (PCR) are important aspects that influence the results and effectiveness of these techniques. We used the primer set ITS4/ITS5 for ARISA to assess the fungal community composition of two sites situated in the Sand Fynbos. The primer set proved to deliver reproducible ARISA profiles of the fungal community composition with little variation observed between ARISA-PCRs. Variation that occurred in a sample due to repeated DNA extraction is expected for ecological studies. This reproducibility made ARISA a useful tool for the assessment and comparison of diversity in ecological samples. In this paper, we also offered particular suggestions concerning the binning strategy for the analysis of ARISA profiles.
Keywords: automated ribosomal intergenic spacer analysis; binning; fungal ecology; fynbos fungi; molecular community fingerprinting; soil fungi
INTRODUCTION
Numerous studies on soil diversity have been conducted using small subunit recombinant DNA (rDNA).1,2,3,4 Although molecular profiling techniques provide little direct evidence to the function of organisms in the soil, they have become invaluable for the understanding of soil microbial diversity and community composition.5,6,7,8 Molecular techniques, such as rDNA methods, provide a good indication of the community structure, including the diversity composition and the evenness of communities.
The method of rRNA intergenic spacer analysis (RISA) provides for an estimation of community diversity and community composition. RISA allows one to estimate microbial diversity without the need to culture organisms. Culturing methods were shown to reveal only about 1% of the total microbial diversity.9,10,11 In addition, the bias in favour of fast-growing organisms and against slow-growing organisms is, to a large extent, eliminated with the RISA technique. The RISA technique also includes the diversity from non-culturable organisms in the soil, which was lacking in studies utilising traditional culturing media.12 RISA requires genomic DNA of the total microbial population under study to be extracted from the environmental sample. This method further involves the amplification of the selected DNA fragments with universal primers and subsequent electrophoresis on a polyacrylamide gel. The RISA technique has been enhanced by the addition of an automated component to the technique by using an automated genetic analyser.5
The automated ribosomal intergenic spacer analysis (ARISA) method is an effective, rapid and fairly inexpensive process that can be used to estimate the diversity and composition of microbial communities without demonstrating a bias towards fast-growing or dominant species.6 This is especially useful in ecological studies, where a large number of samples need to be processed and diversity needs to be determined at a spatial and temporal level. Both fungal and bacterial communities can be determined via this method, depending on the primers used. The method, when denoted as fungal ARISA (F-ARISA), targets the DNA of the intergenic spacer region 1, the 5.8S small subunit and the intergenic spacer region 2 of the fungal communities. This region, especially within the intergenic spacer regions 1 and 2, displays significant heterogeneity in length and nucleotide sequence between species. Bacterial ARISA (B-ARISA) targets the intergenic region between the 16S and the 23S subunits of the rDNA genes in the rRNA operon, which displays size and sequence heterogeneity between species.13
ARISA-PCR involves the use of fluorescently labelled oligo-nucleotide primers commonly labelled with the fluorescent markers ROX and FAM.5,13 Electrophoresis of the total amplified DNA is performed on an automated system, which detects the fluorescently labelled DNA fragments with the aid of a laser and a charge coupled device camera.5 ARISA-PCR, and subsequently the ARISA profiles, are highly reproducible, as demonstrated by Cardinale et al.14 The reproducibility of ARISA is determined in terms of peak occurrence, peak height and peak size, which allows direct comparisons to be drawn between different ARISA profiles as well as between different studies.15 The standard deviation for fragments smaller than 1 kilo-base pair (kbp) was shown to be between 1 base pair (bp) and 2 bp.5 The reproducibility of the abundance of the ARISA fragments is also high; the peaks with the largest contribution to the total fluorescence are the most reproducible. The variability in the number of PCR cycles does not have an influence on the ARISA pattern but, rather, influences the sensitivity of the ARISA.5
The ARISA technique does leave a small margin for error resulting from background fluorescence. For this reason, cut-off levels are implemented for the fluorescent intensity of the ARISA profile, which is 0.5% of the total fluorescence. The heterogeneity of the sampling environment and the randomness of the sampling procedure play a role in the standardisation of the method. The PCR amplification of the targeted DNA may introduce errors in the length of the internal transcribed spacer (ITS) region.1 In addition, the ABI310xl genetic analyser is recognised to be less accurate when the size of the fragments are increased.1 The possibility also exists that two fragments of the same size may be represented as one peak on the ARISA profile. These problems with the ARISA method should be recognised and incorporated into a binning strategy, which places peaks into specific groups and allows for a more accurate comparison of different samples.16
The literature has proposed various empirical binning strategies. Fisher and Triplett5 observed size variations across replicate ARISA profiles of 1-2 bp for fragment sizes below 1 kbp, 3-5 bp for fragments up to 1150 bp and up to 13 bp for the largest size fragments. Hewson and Fuhrman13 used a bin size of 3 bp for fragment lengths up to 500 bp and a bin size of 7 bp for fragments larger than 500 bp. Brown et al.1 suggested that windows 3 bp in width be used for fragments ranging from 400 bp to 700 bp, windows 5 bp in width for those from 700 bp to 1000 bp and windows 10 bp in width for fragments from 1000 bp to 1200 bp. Larger fragments do not, however, necessarily result in higher similarities between similar samples due to the splitting of peaks between adjacent bin windows.
The aim of this study was to optimise and evaluate the ARISA protocol to determine microbial populations in fynbos soil. This niche is expected to contain a large number of species due to its sandy oligotrophic nature. The sandy acidic fynbos soil also contains a high concentration of humic substances, which makes the obtaining of pure DNA challenging.17
MATERIAL AND METHODS
Sensitivity of DNA extraction and ARISA-PCR
The sensitivity of the DNA extraction method for ARISA was tested on a random soil sample collected at Kalbas Kraal in the Western Cape Province of South Africa (33º34'14.20" S, 18º37'43.00" E), using the ZR soil microbial DNA kit (Zymo Research, Orange, United States of America). Penicillium spores are more robust than vegetative cells and were used to provide a stringent test for the extraction method.18 Penicillium was also found to be the dominant species in these soils. Spores of Penicillium spp. were suspended in a phosphate buffered saline (pH 7) solution. The concentration of the cells in the suspension was determined by direct microscope count using a haemocytometer (Superior Marienfeld Laboratory Glassware, Marienfeld, Germany). Spores were used to prepare a dilution series of 104 spores, 103 spores, 102 spores, 101 spores and 1 spore per millilitre. An amount of 1 mL of each diluted spore suspension was added to 1 g of both autoclaved (121 ºC, 100 kPa, 20 min) soil samples and unsterilised soil samples. Total DNA was extracted from the soil using the ZR soil microbial DNA kit (Zymo Research) and quantified spectrophotometrically with a Nanodrop 2000c (Thermo Scientific, Waltham, USA). PCR was performed using fungal specific primers ITS4 and FAM-labelled ITS5. The reaction mixture contained 1 µL of the purified genomic DNA, 500 nM of each primer and 23 µL of KapaTaq readymix (KapaBiosystems, Cambridge, USA) in a total volume of 25 µL. The PCR conditions consisted of an initial denaturing step at 95 ºC for 3 min, followed by 40 cycles of 95 ºC for 30 s, 51 ºC for 30 s and 72 ºC for 30 s. The reaction was completed with a final extension at 72 ºC for 5 min and then cooled and held at 4 ºC. Samples were run on a 1% agarose gel stained with ethidium bromide and visualised under a UV light for the presence of amplified products.
The sensitivity and effectiveness of the ARISA-PCR was considered by determining its ability to detect DNA within a mixed sample. Total genomic DNA was extracted for a soil sample collected at Kalbas Kraal as described earlier. Spore suspensions from Penicillium spp. were quantified using a haemocytometer as described earlier. The spore suspension was vortexed for 1 min at full speed after the addition of 2 mm steel beads. The number of Penicillium spores that were lysed was determined by counting the spores after lyses. The number of spores lysed were diluted to 100, 80, 40, 20 and 10 spores per soil DNA extraction, and added to the genomic DNA from the soil. The number of spores corresponded to the number of genome copies in the PCR.
Conditions for ARISA
The PCR products of every sample were run on an ABI310xl genetic analyser (Applied biosystems, Foster City, USA) to obtain an electropherogram of the different fragment lengths and fluorescent intensities. ARISA-PCR samples were run with the size standard LIZZ 600, containing sizes from 60 bp to 600 bp in length. GeneMapper 4.1 software19 converted the fluorescence electropherogram data representing operational taxonomic units (OTUs) into peaks indicating the fluorescent intensity. Peak heights were favoured over peak area for further analysis due to concerns that peak areas for larger peaks may be subjected to inaccuracies. The variance of peak area values is higher than for peak height. This may be due to peak smoothing and height algorithms that are simpler and a more consistent measure of peak height. Peaks which represented 1% or more of the total fluorescence were considered for analysis. Similarities between samples were represented by calculating the pairwise Whittaker similarity index.13
Evaluating binning sizes for ARISA
Different PCR reactions from a single DNA extraction were compared to determine the effect of different bin sizes on the similarity of the profiles. Bin sizes ranging from 1 bp to 7 bp were considered. Similarities were compared by calculating the Whittaker similarity index. The number of unique OTUs was determined for each binning protocol, along with the number of corresponding and conflicting OTUs. Evaluating binning sizes requires lenient peak selection criteria with no peaks under 0.5% of total fluorescence considered for analysis. This was done to include the effect lower intensity peaks had on the binning strategy. Profile similarities increase due to a high stringency of peak selection across all binning sizes and this phenomenon should be limited in order to observe the real effect bin size has on the similarities of the profiles. Four different profiles were considered and data of all profiles were normalised to the mean values.
Evaluating the use of ARISA to study environmental samples
The significance of the variability of ARISA was evaluated when environmental samples with the primer set ITS4/ITS5 were compared. Soil samples were collected from Sand Fynbos, approximately 10 km outside the town of Malmesbury in the Western Cape. Samples were collected from a range of plots on different sites. Profile variability from two different sites at Camphill Village and Kalbas Kraal, located approximately 6.5 km apart, was evaluated by calculating the Whittaker similarity index. Two different profiles of two unique samples within the plot at Kalbas Kraal were also compared to evaluate variability of different samples within the same plot. The variability within these samples was compared by performing ARISA on different DNA extractions of the same sample. Lastly, the variability of PCR within the same DNA extraction was evaluated. The distance relationship of the similarities at the different levels of sampling was illustrated by means of cluster analysis. Cluster analysis was performed by complete linkage of the Pearson-r correlation values of the Whittaker similarity indices between the sites using Statistica 9 software20.
Construction of a size standard
A size standard was created to enable the use of ARISA for species with peaks larger than 500 bp. A ROX-labelled M13 forward primer was used in conjunction with six other non-labelled primers to generate six ROX-labelled DNA fragments using the plasmid pGEM-3zf as the template. The fragment sizes were calculated by counting the number of bases from the 5' end of the labelled primer to the 5' end of the unlabelled primer. These PCR fragments were mixed with a standard ROX 500 size standard to create a size standard (called ROX 1.1) ranging from 75 bp to 1121 bp.
RESULTS AND DISCUSSION
Sensitivity of the DNA extraction on ARISA
The results of the sensitivity test indicated that Penicillium spores could be detected in the DNA extractions from sterile soil at 40 spores per gram of soil, according to the peaks observed in Figure 1. The detection of the spores in the untreated soil samples with higher peak diversity, however, required a higher concentration of spores (100 spores per gram of soil) to distinguish between the spores and the background noise and to compensate for competitive binding to the environmental DNA (Figure 1). This compared favourably to other studies, where, Van Elsas et al.21 were able to detect Trichoderma harzianum as low as 103 spores per gram of soil on an agarose gel, while Doaré-Lebrun et al.18 found Aspergillus carbonarius to be detectable at levels of 103 cells in a mixed culture and Penicillium expansum in the range of 105 cells. The detectable limit of fungal spores in a sample is generally higher than that of vegetative cells.
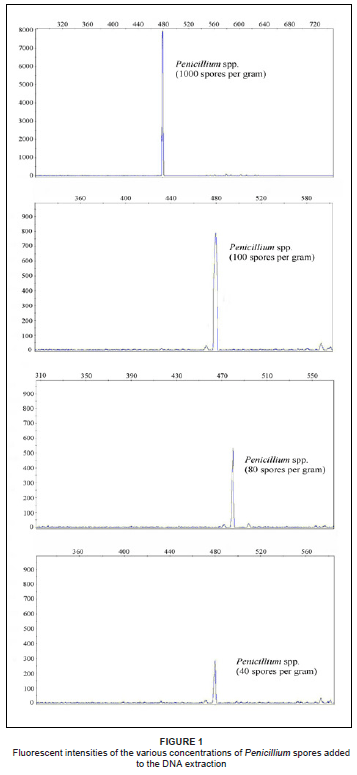
Sensitivity of the ARISA-PCR
The higher dilutions of 20 spores per gram of soil and 10 spores per gram of soil were impossible to detect, probably due to competitive binding by high concentrations of DNA templates (Table 1). Peaks that were detected at solutions of 10 spores to 20 spores per gram of sterile soil were extremely variable, with less than one in four PCR's detecting the Penicillium spp. at very low levels. This result indicates that DNA concentration should be at least the equivalent of 40 spores per gram of soil extracted to produce a consistent peak on the electropherogram. The sizing of the fragment varied between 479.32 bp and 480.02 bp; the peaks were thus sized with an error in precision of less than 1 bp.
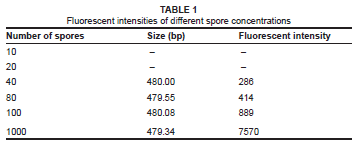
Evaluating binning sizes for ARISA
The different binning sizes examined for the various samples indicated that the maximum similarity between identical samples was reached at a bin size of 4 bp. The Whittaker index value, however, did not show a significant difference between bin sizes 2 bp, 3 bp and 4 bp. The bin size of 3 bp resulted in the highest number of peaks shared, namely 23, with 2 peaks not shared. The binning size of 5 bp resulted in an average of nine OTUs that were not shared. The Whittaker index increased at a bin size of 5 bp due to unwanted overlaps that occurred at this bin size, with nine OTUs not shared. Larger bin sizes did not necessarily result in a more analogous profile because a peak may have been incorrectly placed in a bin with other peaks, even if it was clearly distinct. Furthermore, an increase in bin size also resulted in a decrease in the total number of OTUs (Table 2). The ideal binning protocol thus required maximum similarity between parallel samples, with the maximum number of total and shared OTUs.
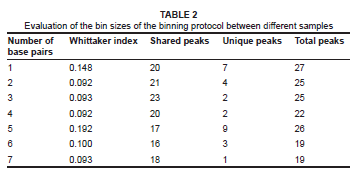
Reproducibility of ARISA profiles
The Whittaker index revealed that the ARISA profiles were highly reproducible when considering duplicate samples of ARISA from the same DNA extraction (Figure 2). The duplicate ARISA-PCR showed 98% similarity with a linkage distance of 0.01 after cluster analysis with normalised data and analysed using a bin size of 3 bp. All peaks were reproduced in the ARISA profile and only differed in terms of their relative proportions of fluorescence intensity. The Whittaker index revealed high similarity between repeated DNA extractions from the same soil sample. ARISA profiles showed a percentage similarity of 93% with a 0.1 linkage distance. The Whittaker index decreased when DNA extractions of different samples on the same plot were used for ARISA. The linkage distance value increased to 0.7 with a percentage similarity of 78%. The Whittaker similarity index again decreased when a sample for a different site (i.e. Camphill Village) was compared in the cluster analysis. The similarity of the ARISA profile after repeated PCRs with the same sample and high similarity with repeated DNA extractions, demonstrated that analysis was reproducible. The findings were as expected and similar to another study that found greater variation between sites than within the site and greater variation between PCR reactions of DNA extractions from the same plot than from different plots.6 The ARISA method was found to be robust regardless of the primer set used and the specific PCR conditions.
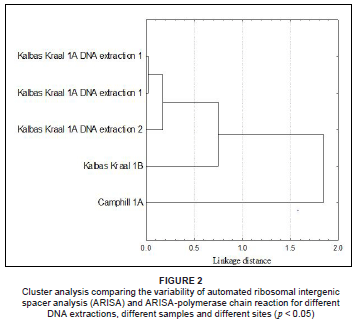
ARISA was also evaluated for bacterial-specific primers. However, detected peaks with sizes up to 1000 bp in length were observed. This required a sized standard which incorporated larger fragments. ROX 1.1 was constructed and included the following size fragments: 75 bp, 100 bp, 139 bp, 150 bp, 160 bp, 200 bp, 300 bp, 340 bp, 350 bp, 400 bp, 450 bp, 490 bp, 500 bp, 583 bp, 683 bp, 782 bp, 932 bp, 991 bp and 1121 bp. The same binning strategy proved to be sufficient for bacterial ARISA.
CONCLUSION
The primer set ITS4/ITS5 has not been commonly used for the purpose of ARISA. This primer set performs well when testing the sensitivity and 40 templates per gram of soil would be sufficient to detect both dominant and minor species in environmental samples. The DNA extraction method tested detected cells at 100 spores per gram of soil and, as is the case with most studies looking at environmental samples, the DNA extraction method is the defining step for the sensitivity of the method. The reproducibility of ARISA using the primer set ITS4/ITS5 is very high, with little variation, even under lenient filtering conditions.
ACKNOWLEDGEMENTS
We would like to acknowledge Stellenbosch University and the South African National Research Foundation for funding and the Western Cape Nature Conservation Board for allowing access to the Riverlands Nature Reserve.
REFERENCES
1. Brown MV, Schwalbach MS, Hewson I, Fuhrman JA. Coupling 16S-ITS rDNA clone libraries and automated ribosomal intergenic spacer analysis to show marine microbial diversity: Development and application to a time series. Environ Microbiol. 2005;14:1466-1479. [ Links ]
2. Borneman J, Triplett EW. Molecular microbial diversity in soils from eastern Amazonia: Evidence for unusual microorganisms and microbial population shifts associated with deforestation. Appl Environ Microbiol. 1997;63:2647-2653. [ Links ]
3. Kuske CR, Barns SM, Busch JD. Diverse uncultivated bacterial groups from soils of the arid southwestern United States that are present in many geographic regions. Appl Environ Microbiol. 1997;63:3614-3621. [ Links ]
4. Fierer N, Breitbart M, Nulton J, et al. Metagenomic and small-subunit rRNA analyses reveal the genetic diversity of bacteria, archaea, fungi, and viruses in soil. Appl Environ Microbiol. 2007;73:7059-7066. [ Links ]
5. Fisher MM, Triplett EW. Automated approach for ribosomal intergenic spacer analysis of microbial diversity and its application to freshwater bacterial communities. Appl Environ Microbiol. 1999;65:4630-4636. [ Links ]
6. Ranjard L, Poly F, Lata JC, Mougel C, Thioulouse J, Nazaret S. Characterization of bacterial and fungal soil communities by automated ribosomal Intergenic spacer analysis fingerprints: Biological and methodological variability. Appl Environ Microbiol. 2001;67:4479-4487. [ Links ]
7. Lerner A, Herschkovitz Y, Baudoin E, et al. Effect of Azospirillum brasilense inoculation on rhizobacterial communities analyzed by denaturing gradient gel electrophoresis and automated ribosomal intergenic spacer analysis. Soil Biol Biochem. 2006;38:1212-1218. [ Links ]
8. Mills DK, Entry JA, Gillevet PM, Mathee K. Molecular-based approaches to soil microbiology assessing microbial community diversity using amplicon length heterogeneity polymerase chain reaction. Soil Sci Soc Am. 2007;71:572-578. [ Links ]
9. Richaume A, Steinberg C, Jocteur-Monrozier L, Faurie G. Differences between direct and indirect enumeration of soil bacteria: The influence of soil structure and cell location. Soil Biol Biochem. 1993;25:641-643. [ Links ]
10. Torsvik V, Goksoyr J, Daae FL. High diversity in DNA of soil bacteria. Appl Environ Microbiol. 1990;56:782-787. [ Links ]
11. Tsuji T, Kawasaki Y, Takeshima S, Sekiya T, Tanaka SA. New fluorescence staining assay for visualizing living microorganisms in soil. Appl Environ Microbiol. 1995;61:3415-3421. [ Links ]
12. Kennedy N, Clipson N. Fingerprinting the fungal community. Mycologist. 2003;7:158. [ Links ]
13. Hewson I, Fuhrman JA. Richness and diversity of bacterioplankton species along an estuarine gradient in Moreton Bay, Australia. Appl Environ Microbiol. 2004;70:3425-3433. [ Links ]
14. Cardinale M, Brusetti L, Quatrini P, et al. Comparison of different primer sets for use in automated ribosomal intergenic spacer analysis of complex bacterial communities. Appl Environ Microbiol. 2004;70:6147-6156. [ Links ]
15. Hansgate AM, Schloss PD, Hay AG, Walker LP. Molecular characterization of fungal community dynamics in the initial stages of composting. FEMS Microbiol Ecol. 2005;51:209-214. [ Links ]
16. Ruan Q, Steele JA, Schwalbach MS, Fuhrman JA, Sun F. A dynamic programming algorithm for binning microbial community profiles. Bioinformatics. 2006;22:1508-1514. [ Links ]
17. Lakay FM, Botha A, Prior BA. Comparative analysis of environmental DNA extraction and purification methods from different humic acid rich soils. J Appl Microbiol. 2007;102:265-273. [ Links ]
18. Doaré-Lebrun E, El Arbi A, Charlet M, et al. Analysis of fungal diversity of grapes by application of temporal temperature gradient gel electrophoresis - Potentialities and limits of the method. J Appl Microbiol. 2006;101:1340-1350. [ Links ]
19. Genemapper [computer program]. Version 4.1. Foster City: Applied biosystems; 2009. [ Links ]
20. Statistica [computer program]. Version 9. Tulsa: Statsoft; 2009. [ Links ]
21. Van Elsas JD, Duarte GF, Keijzer-Wolters A, Smit E. Analysis of the dynamics of fungal communities in soil via fungal-specific PCR of soil DNA followed by denaturing gradient gel electrophoresis. J Microbiol Methods. 2000;43:133-151. [ Links ]
 Correspondence to:
Correspondence to:
Etienne Slabbert
Postal address: Department of Microbiology, Stellenbosch University
Private Bag X1
Matieland 7602, South Africa
email: 13265695@sun.ac.za
Received: 04 Sept. 2008
Accepted: 11 June 2010
Published: 04 Aug. 2010
This article is available at: http://www.sajs.co.za




![Dichloro(bis[diphenylthiourea])cadmium complex as a precursor for HDA-capped CdS nanoparticles and their solubility in water](/img/en/prev.gif)









