Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Science
On-line version ISSN 1996-7489
Print version ISSN 0038-2353
S. Afr. j. sci. vol.105 n.7-8 Pretoria Jul./Aug. 2009
RESEARCH LETTERS
Circular dichroism as a means to follow DNA gymnastics: on the shoulders of giants
M. Mills; C. Lin; M. Chauhan; H.H. Klump*
Department of Molecular and Cell Biology, University of Cape Town, Rondebosch 7701, South Africa
ABSTRACT
This is the first report of DNA stem-loops self-assembled by 'foot-loop' interactions into either two-dimensional strings or three-dimensional spirals, distinguished by circular dichroism spectroscopy. All subunits are linked by cooperative Watson-Crick hydrogen bonds.
Key words: DNA, circular dichroism, nanostructures, stem-loop structures, self-assembly
Introduction
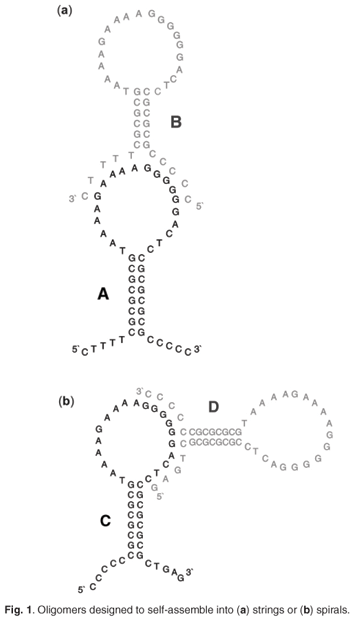
Many of the key goals of DNA-based nanotechnology entail the use of periodic arrays with tunable features such as binding patterns and cavities,1 with important applications in medicine as drug-delivery systems.2 Branched networks that result from ligating three-way junctions of DNA together have been reported.3,4 This study introduces a new method to build either linear or three-dimensional self-assembling structures. Pairs of DNA stem-loops (46 bases) were designed to self-assemble by foot-loop interactions to form networks of strings (A+B) or spirals (C+D) (Fig. 1). The inter-subunit interactions are restricted to cooperative Watson-Crick hydrogen bonds. No ligation is used—making the assembly completely reversible.
Methods
Oligonucleotide design and synthesis
The sequences were designed to form a double-stranded stem of alternating guanine/cytosine base pairs5 holding a single-stranded, asymmetrical loop sequence from the polypurine tract of HIV-1 (A4GA4G6A).6 The 'feet' (5'- and 3' extensions of the stem) were added to make complementary base pairs to the loop, thus forming either strings or spirals. Heterodimers were required to make the foot-loop interaction constitute an anti-parallel double helix. Oligonucleotides were synthesised by standard phosphoramidate chemistry using a Beckman 1000 M DNA synthesiser, and purified in dimethoxytrityl-on mode by reverse-phase high performance liquid chromatography (HPLC) using an acetonitrile gradient. Concentrations were expressed in strand molarity, using nearest-neighbour approximation for the extinction coefficients of the unfolded species.7
Ultraviolet (UV) melting
Thermal melting curves were recorded using a heating rate of 1°C min–1 in an Uvikon spectropolarimeter, with a custom-made heating block equipped with an Oasis analog-to-digital converter. The DNA strand concentration was 1.5 µM ml–1 and oligonucleotides were dissolved in 100 mMl–1 sodium chloride, 5 mMl–1 magnesium chloride, and 10 mMl–1 sodium cacodylate, at pH 7.0.8
Circular dichroism spectroscopy
Circular dichroism (CD) spectroscopy of 1.5 µM ml–1 DNA strand concentration in 100 mMl–1 sodium chloride, 5 mMl–1 magnesium chloride, and 10 mMl–1 sodium cacodylate, at pH 7.0 in a 1 ml jacketed quartz cuvette, was performed in a Jasco J-810 spectropolarimeter with ten accumulations per scan. Circular dichroism, with units of millidegrees, reflects detailed helical geometry and is defined as the difference between the absorption of left- and right-handed, circularly-polarised light measured as a function of wavelength (nm). The sample was kept at a constant temperature by a Haake D8 programmable water bath during each set of scans.
Polyacrylamide gel electrophoresis (PAGE)
Native PAGE was performed with samples at a DNA strand concentration of 1 µM ml–1, pre-heated to 80°C, and incubated at room temperature in 100 mMl–1 sodium chloride, 5 mMl–1 magnesium chloride, and 10 mMl–1 sodium cacodylate, pH 7.0, before loading on a 10% gel run at 70 volts with tris-borate running buffer (pH 8.3), and visualised under UV light by ethidium bromide intercalation.
Results
Melting behaviour of the networks
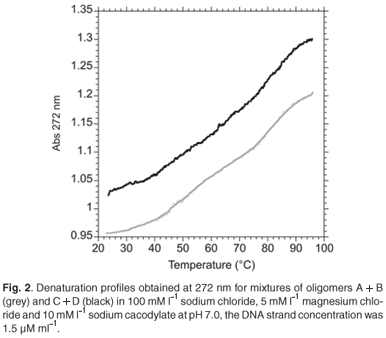
The melting of both assemblies (Fig. 1.) was biphasic at a neutral pH (Fig. 2.) corresponding to the denaturation of the intermolecular 'foot-loop' double helix (melting temperature (Tm) = 52°C), followed by the unfolding of the intramolecular stem double helix (Tm = 82°C) in 1 ml of 100 mMl–1 sodium chloride, 5 mMl–1 magnesium chloride, and 10 mMl–1 sodium cacodylate.
Circular dichroism
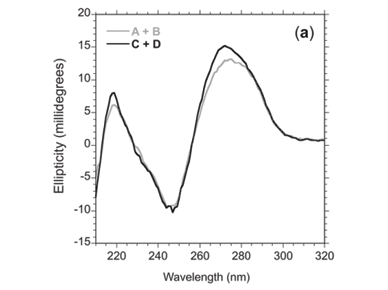
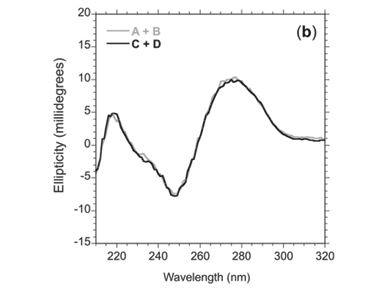
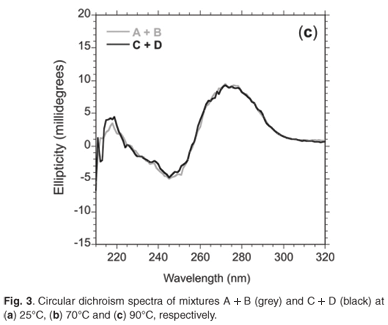
Circular dichroism spectra revealed that the network made up of oligomers C+D showed more helicity (15 m°) than oligomers A+B (13 m°) at 272 nm (Fig. 3a), as expected. Upon raising the temperature above the melting temperature (Tm, the temperature at which half of the observed absorbance change has been recorded) of the 'foot-loop' interactions, the spectra of the individual stem-loops were similar at 70°C (Fig. 3b). Further heating to 90°C resulted in similar spectra for the random coil species (Fig. 3c).
Polyacrylamide gel electrophoresis
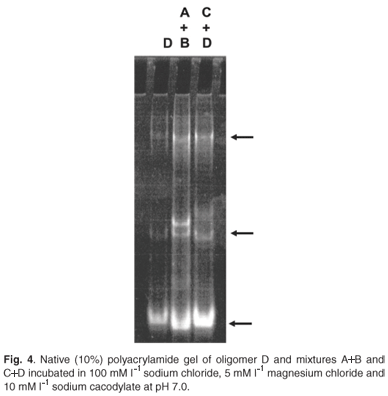
Lanes 1, 2 and 3 of Fig. 4 show the monomer stem-loop D and the dimers and higher networks formed by A+B and C+D, respectively. The migration patterns of lanes 2 and 3 were different, as expected, supporting the CD evidence that networks A+B and C+D had different secondary structures.
Conclusion
This is the first time that 'foot-loop' interactions have been shown to create either linear strings or spirals that were distinguished by CD. Future work will include electron and atomic-force microscopy of the networks, as well as incorporate a transcription factor-binding site into the stem sequence. Hetero-multimers, whether they are formed by single strands or represent i-motif, parallel or anti-parallel guanine-duplexes, Hoogsteen or anti-Hoogsteen triplexes, can self-assemble in this way under the controlling influence of ionic strength and/or pH.
We thank P. Ma (UCT) for DNA synthesis and the National Research Foundation, South Africa for financial support.
1. Seeman N.C. (2005). Structural DNA nanotechnology: an overview. Methods Mol. Biol. 303, 143–166. [ Links ]
2. Klump H.H., Chauhan M., Mills M. and Lin C. (2006). Progress in designing nucleic acid structures and fine-tuning their interactions. Arch. Biochem. Biophys. 453, 91–98. [ Links ]
3. Lin C. (2006). DNA based networks, nanostructures and meta-stable states. Ph.D. thesis, University of Cape Town, South Africa. [ Links ]
4. Husler P.L. and Klump H.H. (1995). Thermodynamic characterization of a triple-helical three-way junction containing a Hoogsteen branch point. Arch. Biochem. Biophys. 10, 149–166. [ Links ]
5. Völker J., Makube N., Plum G.E., Klump H.H. and Breslauer K.J. (2002). Conformational energetics of stable and metastable states formed by DNA triplet repeat oligonucleotides: implications for triplet expansion diseases. Proc. Natl. Acad. Sci. USA 99, 14700–14705. [ Links ]
6. Wain-Hobson S., Sonigo P., Danos O., Cole S. and Alizon M. (1985). Nucleotide sequence of the AIDS virus, LAV. Cell 40, 9–17. [ Links ]
7. Warshaw M.M. and Cantor C.R. (1970). Oligonucleotide interactions. IV. Conformational differences between deoxy- and ribodinucleoside phosphates. Biopolymers 9, 1079–1103. [ Links ]
8. Mills M., Völker J. and Klump H. (1996). Triple helical structures involving inosine: there is a penalty for promiscuity. Biochemistry 35, 13338–13344. [ Links ]
Received 18 May. Accepted 26 May 2009.
* Author for correspondence E-mail: horst.klump@uct.ac.za