Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Science
On-line version ISSN 1996-7489
Print version ISSN 0038-2353
S. Afr. j. sci. vol.104 n.7-8 Pretoria Sep./Aug. 2008
RESEARCH ARTICLES
Ectomycorrhizas in association with Pinus patula in Sabie, South Africa
G.L. HawleyI; A.F.S. TaylorII; J.F. DamesI, *
IDepartment of Biochemistry, Microbiology and Biotechnology, Rhodes University, P.O. Box 94, Grahamstown 6140, South Africa
IIDepartment of Mycology and Forest Pathology, Swedish University of Agricultural Science, Box 7026, S-750 07 Uppsala, Sweden
ABSTRACT
Forestry is an economically important industry in South Africa, involving extensive exotic plantations of Eucalyptus, Pinus and Acacia species. These tree species have fungal associations, such as ectomycorrhizas, that have become locally naturalized. The forestry industry is increasingly faced with problems of long-term sustainability, increasing soil acidity and depletion of soil nutrients. It is, therefore, essential that the fundamental importance of the ectomycorrhizal (ECM) symbioses in the nutrient cycling, growth, health and survival of these tree species should not be ignored. Research on the species diversity of ECM fungi associated with forestry plant species has been hampered by the difficulty of identifying the fungi involved in the symbiosis. This investigation focused on the ECM fungi associated with Pinus patula (Schlecht. et Cham.) grown in managed plantations in the Sabie region, Mpumalanga province, South Africa. ECM roots were morphotyped and DNA was extracted. The internal transcribed spacer (ITS) region was amplified using the ITS 1F and ITS 4 primers. The sequences were BLASTed using the GenBank and UNITE databases. Twenty-seven extractions were successfully amplified representing 17 different morphotypes. Of the 27 sequences, 21 were identified as ECM fungi and, from the BLAST results, eleven different ECM species could be identified. Selected ECM root types were morphologically and anatomically described according to root morphology, mantle structure, specialized hyphae and rhizomorph arrangement. Seven dominant field types were described and identified as two Amanita species, Scleroderma citrinum, a suilloid species, Thelephora terrestris, a tometelloid species and one resembled an Albatrellus species.
Introduction
Forestry in South Africa is extensive, covering approximately 1.3 million hectares,1 and is planted mainly with exotic species such as Pinus, Acacia and Eucalyptus. Naturalized ectomycorrhizal (ECM) fungi have been found associating with these tree species, and are necessary for successful tree establishment in managed plantations.2 The diversity and associated mechanisms of the symbiosis in a South African context have received only limited attention. Doidge3 published the last comprehensive list of fungi in South Africa in 1950, which included ECM fungal species from the genera Amanita, Boletus, Cantharellus, Lactarius, Scleroderma, Rhizopogon and Russula.
Previous attempts to document the ECM fungal species diversity occurring in Pinus plantations were conducted using sporocarp surveys. Lundquist4 produced a good listing of all the fungi associated with the genus Pinus, but it is limited in its information as there is no reference to the mycorrhizal status of the fungi. The list is based on the sporocarp deposits in the National Collection of Fungi, which is biased against undetectable ECM fungi below-ground, or non-sporulating fungi. Van der Westhuizen and Eicker5 recorded 21 Basidiomycota ECM fungi from sporocarp data, basing the mycorrhizal status on current understandings of genera and species known to form ectomycorrhizas. Although above-ground diversity confirms the presence of species, it is understood that above- and below-ground species diversity and frequency may differ considerably, given the micro-niches and dynamics within the soil.6,7 It is also possible for fungal species that do not form sporocarps and hypogeous or resupinate sporocarps to go undetected.
In South Africa, research on the below-ground diversity at ECM root tip level is limited. Dames et al.8 characterized two ECM morphotypes from field samples growing in association with Pinus patula (Schlecht. et Cham.), identifying them as Scleroderma citrinum Pers. and Boletus pinicola Rea. Carlson9,10 observed a response of ECM fungal species associated with P. patula to acidic precipitation. Carlson9,10 detected a change in morphotype composition and concluded that there was no decrease in root colonization.
Traditional morphological characterization of ECM root tips is regarded as an important identification technique. Currently, six confirmed ECM fungal species, including the descriptions by Dames et al.8 associated with P. patula, have been characterized. Amanita muscaria (L) Hook., Laccaria laccata (Scop.) Fr., Suillus brevipes (Peck) Kuntze and Thelephora terrestris Ehrh. were characterized by Mohan et al.11,12 after inoculating nursery seedlings with sporocarp material and harvesting the roots after five months.
Molecular and DNA amplification techniques have been increasingly used to resolve or aid a number of systematic and ecological challenges associated with the identification of ECM fungi,13 such as species phylogeny and delineation from field samples as well as allowing for fast and accurate identification, and therefore allowing processing of large community/population sampling. Molecular techniques have proved useful, especially when morphological data are non-informative or not sufficiently refined. Molecular techniques that have been employed included restriction fragment length polymorphism,14–16 DNA probing,15 DNA sequencing of the large ribosomal subunit and internal transcribed spacer regions. The molecular data derived from field samples can then be compared to known data (restriction patterns or sequences) from identified sporocarp material or by comparison with sequences submitted to large online databases.
The forestry industry in South Africa is facing increasing problems of soil nutrient depletion, soil acidity due to pollution, and legislation of land used for forestry. Licensing is required, limiting the expansion of land for forestry plantation (National Water Act, No. 36 of 199817). The industry is therefore faced with having to strategically manage the current plantations and to render the soils re-useable as a sustained resource. No long-term strategy has been devised to develop and maintain soil microbial health or to maximize nutrient availability. Khalil et al.2 reported that, in South Africa, P. patula survival and establishment after transplanting was dependent on ECM establishment.
Continued ECM research in South Africa will depend on comprehensive records of fungal species diversity recorded from both above- and below-ground collections to determine their population dynamics. These records should rely on both morphological and molecular techniques to determine above- and below-ground species identification and description. The motivation was to create an initial database of ECM fungal species associating with P. patula species grown in managed plantations in South Africa. Characterizing root tip material in terms of their DNA sequences and pairing these with classical morphological descriptions was carried out to achieve this.
Methods
Study site description
Two Pinus patula plantations, Brooklands and Driekop, were selected around Sabie in Mpumalanga province, which supports an intensive forestry industry. The sites are approximately 50 km apart. The climate of this area is subtropical, experiencing mild winters (6°C minimum to 27°C maximum as the daily average) and hot, humid summers (18°C minimum to 30°C maximum as the daily average) with a mean annual rainfall of 736 mm.18 One compartment was selected from each plantation.
The Brooklands plantation site, compartment E23A (25°11.2'S, 30°43.5'E), was planted in 1983 as a second rotation P. patula stand, at an altitude of 1410 m. The Driekop plantation site, compartment C25 (25°02.9'S, 30°53.4'E), was planted in 2000, with plots of P. patula, P. elliotii and P. taeda. This site is situated at an altitude of 1600 m. Only the P. patula plots were sampled for this study. The mature (Brooklands) and young (Driekop) compartments were chosen such that successive ECM fungal species could be represented.
Sampling of root material
Soil cores (diameter 5 cm) were taken from selected plots in each of the two compartments. These were taken approximately 1 m from the base of the tree. The corer extracted soil to a depth of 20 cm, and the root-containing soil was placed in sealed plastic bags and kept at 4°C until processed.
Approximately 40–45 cores were collected during each event from Brooklands and Driekop, respectively. Five sampling events were conducted in June 2002, December 2002, April 2003, December 2003 and March 2005. All the roots were washed clean of any soil and organic matter using a 2-mm sieve and were inspected under a dissecting microscope (Leica S4E). Where possible, the material was observed fresh or preserved in 50% ethanol for future examination.19 ECM root tips exhibiting similar morphological characteristics of colour, branching type and mantle surface texture were morphotyped.20 These were coded according to the characteristics and stored in 50% ethanol to maintain the integrity of the fungal DNA.
Molecular analysis
Extraction of DNA from ECM root tips. DNA sequencing was conducted at the end of 2002 and in mid-2003 at the Swedish University of Agricultural Science, Uppsala. DNA was extracted from individual tips with 2–6 representatives of each morphotype using a modified CTAB method.21 Single individual root tips were homogenized in 600 µm 2% CTAB solution and incubated at 65°C for 30 minutes. Proteinase K (20 µm, 20 mM) was added to the root material before incubation, to degrade any proteins, which could inhibit the polymerase chain reaction (PCR). The DNA was precipitated with isopropanol and the pellet was resuspended in 50 µm autoclaved milli-Q water.
The primer pair used in the PCR reaction were ITS1F22 and ITS4,23 such that amplification was not limited to the Basidiomycota, as members of Ascomycota and Zygomycota are also known to form ECM.19,21,24 High Fidelity Taq polymerase (ExpandTM High Fidelity PCR System, Boehringer Mannheim, Germany) was used for the PCR reaction. The PCR amplification reactions contained both positive and negative controls, of which the DNA template contained a positively amplified fungal ITS region and autoclaved milli-Q water, respectively. The PCR reaction medium contained, in percentage volumes, 41.8% sterile milli-Q water, 10% 10× reaction buffer (MgCl2), 10% dNTPs (2 mM of each base), 6.5% 25 mM MgCl2, 3% 10 µM of each primer and 0.75% 3.5 U/µl Taq polymerase, and was conducted with the following temperature cycles: 94°C for 2 min; 94°C for 60 s, 50°C for 30 s, 72°C for 60 s for 35 cycles; final extension at 72°C for 7 min. Products of the PCR reactions were fractionated on a 2% agarose gel. Double bands that were observed on the gel were considered as contamination or co-amplification of more than one species, and were discarded.
Successful amplifications were cleaned with a PCR Purification Kit (Qiagen Cat. No. 28004) and re-amplified using a Terminator Ready Reaction Mix (BigDYE V3.1 ABI, U.S.A.) with both primers separately and sequenced using the ABI 310 DNA sequencer (Perkin Elmer). The forward and reverse sequences were aligned using the Vector NTI suite package (InforMax) and edited. The sequences obtained in this study for the different ECM fungal species were deposited in the GenBank25 database and accession numbers obtained. Non-mycorrhizal fungi were submitted to GenBank.25
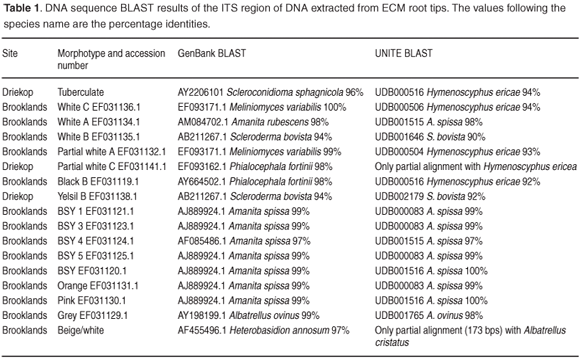
Sequence BLAST analysis. The consensus sequences were submitted to the GenBank25 and UNITE26 databases and compared with similar sequence references, for identification. Identity percentages were used to compare sequences within genotypic groups. In this study, the preferential cut-off for species inclusion was 98%, below which the sequences were considered to represent different species.
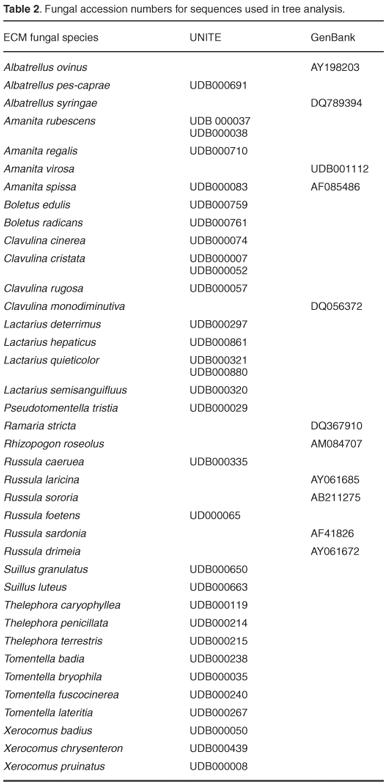
From Table 1 and Figs A–D (in supplementary material online at www.sajs.co.za), it is evident that a number of different morphotypes resulted in the same species alignment result. As a result of differing sequence lengths, BLAST alignment results can be misleading by misaligning with closely related species. To confirm the taxonomic position of the ECM root tip sequences generated, similarity trees were derived. This was accomplished by downloading several known sequences from GenBank25 and UNITE26 (Table 1), and sequences generated from sporocarp collections,27 together with the ECM root tip sequences, which were aligned in ClustalW28 and visualized in Treeview (Win32, 1.6.6, 2001). The ITS region, although useful for species distinction, should not be used for higher-order resolution as the variation between different genera, particularly in the ITS 1 and ITS 2 regions, would not allow for homologous base-pair alignment. Species within the same genus were therefore aligned and analysed with appropriate outgroup species from sister genera.
Morphological and anatomical description
Selected representatives of each morphotype were used for further morphological and anatomical description. The key morphological features that were examined included root tip branching, shape of branches, mantle colour and surface texture, presence of rhizomorphs and emanating hyphae, as described by Agerer.20 Anatomical characteristics that were observed included detailed records of mantle morphology, characteristics of emanating hyphae and rhizomorphs and structure of the Hartig net in terms of hyphal arrangement, hyphal dimensions, presence of specialized cells and mantle layering.11,12, 29–31 Mantle sections were peeled from whole roots, mounted in lactoglycerol32 or water, and examined for surface hyphal arrangement and key distinguishing anatomical features, including pigmented and specialized hyphae. ECM root descriptions were based on the examination of 7–10 root tips for each morphotype.
Sectioning, staining and photography
Cross- and longitudinal sections were examined to characterize various layers of hyphal arrangement and determine the presence of specialized cells.20 The sectioning of root material was conducted by mounting individual root tips of each morphotype in 2% glycerol.32 Cross- and longitudinal sections, approximately 50 µm thick, were prepared using a freeze microtome (Mectron model 150, Frigistor). The resulting sections were placed in a drop of water in 1.5 ml Eppendorf tubes. The lactoglycerol trypan blue staining procedure32,33 for whole root staining was followed and modified by extending the hydrogen peroxide soak until darker sections were bleached. During the course of staining, the samples were centrifuged at 5000 rpm for 30 s if root sections became suspended. Sections were slide mounted in lactoglycerol and anatomical characteristics were examined using a compound microscope (Olympus BX50).
ECM morphological characteristics were examined after immersion of samples in water, using a dissecting microscope (Wild Photomakroskop M400) and photographed on 100 ASA daylight slide film. Anatomical characteristics were examined using bright field microscopy (Olympus BX50) and photographed on 100 ASA black and white film.
Results
Molecular analysis
The ECM morphotype sequences were BLAST analysed in relation to the GenBank25 and UNITE26 databases. The results given in Table 1 indicate incongruence within morphotypes. Different species are evident within a single morphotype and a number of different morphotypes constitute the same genotypic species.
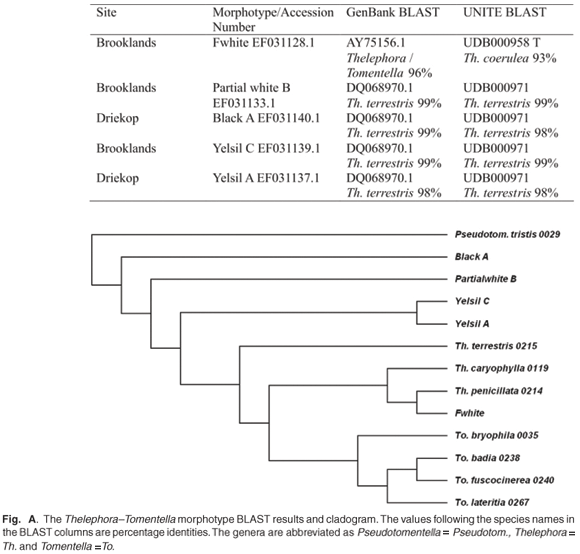
For example, results showed that Thelephora terrestris was represented by several different morphotypes: Partial white B, Black A, Fwhite, Yelsil C and Yelsil A (Fig. A). Fwhite was aligned with Thelephora coerulea. The morphotype sequences that aligned with Thelephora species were aligned with Tomentella and Thelephora species downloaded from UNITE.26 In the derived tree the morphotype Fwhite was grouped together with Thelephora penicillata, indicating that the alignment with T. caryophylla was potentially inaccurate. All the other morphotypes—Yelsil A, Yelsil C, Partial white B and Black A—group with Thelephora terrestris at basal nodes.
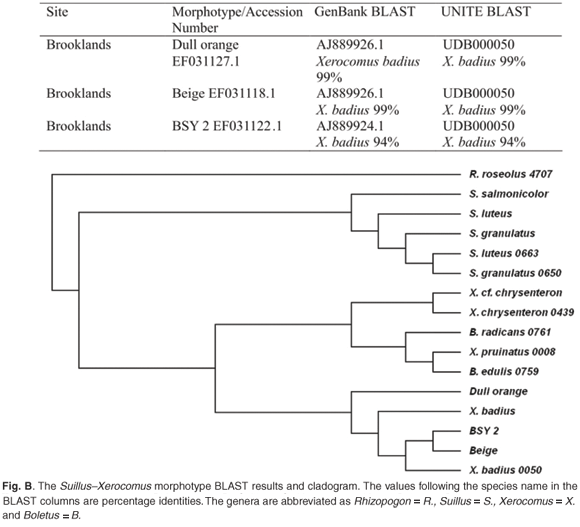
Xerocomus badius was represented by morphotypes BSY 2, Beige and Dull orange. The taxonomic position of the Xerocomus morphotypes was investigated by aligning the sequences together with species of Boletus and Suillus (Fig. B). The morphotypes Dull orange, BSY2 and Beige formed a monophyletic group with X. badius (UNITE). All other BSY morphotypes—Grey-pink, Orange and White A—aligned with Amanita spissa (Table 1).
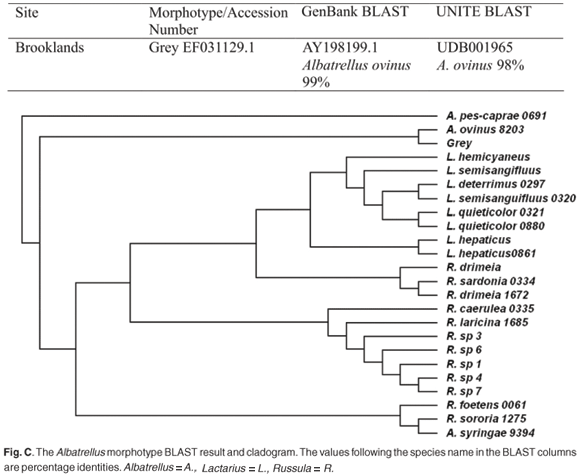
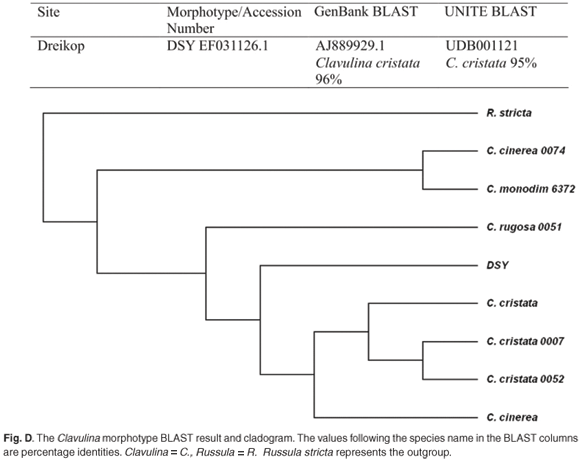
The genus Scleroderma aligned poorly to both morphotypes White B and Yelsil B, with expectation values of e-164 and e-128, respectively (Table 1). Albatrellus ovinus was represented by morphotype Grey. The similarity tree of the three genera, Russula, Lactarius and Albatrellus (Fig. C), confirmed the position of the morphotype Grey within the genus Albatrellus. DSY aligned well with Clavulina cristata (Fig. D).
White C and Partial white A were aligned with Hymenoscyphus sp. The morphotype Beige/white resulted in a strong alignment with Heterobasidion annosum on the GenBank database. Discrepancies between the database BLAST results, e.g. for Tuberculate and White C, are the result of missing species in either database and the limitation of the UNITE database containing only ECM fungal species.
Morphological description
Many of the ECM morphotypes were rare or infrequent and were therefore not selected for detailed description. From the Brooklands plots the dominant morphotypes White, BSY, Orange and Grey were selected. Three dominant morphotypes from Driekop, Black, Yelsil and Tuberculate, were characterized. Of the above types, White, Orange, Yelsil, Black and Tuberculate were common between the two sites, but varied in frequency of occurrence.27 The appearance and descriptions of each morphotype are presented in Figs 1–7.
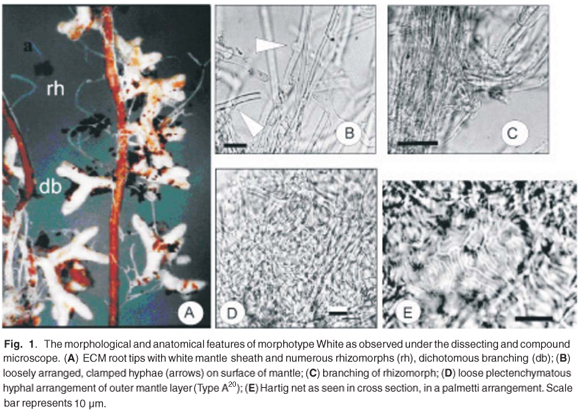
Morphotype White
Sequence matches to this morphotype were Hymenoscyphus sp., Amanita spissa and Scleroderma bovista (Table 1). Morphotype White was derived from the Brooklands study site and was recorded on 33% of all the ECM root tips that were examined.27
Root tip morphology
Root tip branching was dichotomous, long and straight (Fig. 1A) and the dimensions of the last order axis tips ranged from 0.5–1.2 mm in length and from 0.25–0.3 mm in diameter. The mantle surface consisted of loosely arranged stringy hyphae that trap air, making the mantle hydrophobic, and giving the root tips a silvery sheen. The mantle was white, bruising dull yellow with no soil attachments on the mantle. Frequent rhizomorphs emanating from the mantle were white to light yellow/ opaque, and had smooth margins with few hyphal branches. Emanating hyphae were found along the length of the colonized root.
Anatomical morphology
The mantle was composed of two plectenchymatous layers (Type A20). The outer layer was loosely arranged (Fig. 1D) with hyphal dimensions of 2.9–3.9 µm in width and 14.6–34.0 µm in length. The inner layer was more compact, 2.9–3.9 µm in width and 9.7–21 µm in length. Rhizomorphs were observed to be thicker in diameter at branching points, forming nodia and internodia (Fig. 1C). The diameter of the rhizomorph ranged from 34.0–38.8 µm. Emanating hyphae were clamped (Fig. 1B) and were 2.5–4.9 µm in width and 43.6–58.2 µm in length from septum to septum. The Hartig net was in a palmetti arrangement20 that developed from branching hyphae aligned alongside each other to form a single layer between the cells (Fig. 1E). The Hartig net hyphae penetrated 1 or 2 layers of the root cortical cells.
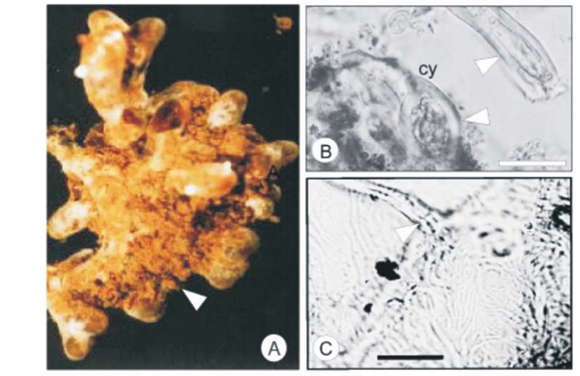
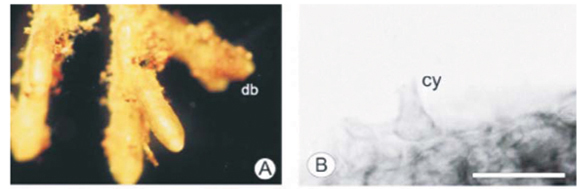
Morphotype BSY
The morphotype BSY was represented by two species, Xerocomus badius (see Fig. B in supplementary material online) and Amanita spissa (Table 1), based on sequence data. BSY was described from the Brooklands study site and was noted on 21% of all the ECM root tips that were recorded from there.27
Root tip morphology
Branching of the root tips was coralloid to irregularly pinnate (Fig. 2A), with the last order axis ranging from 2–2.5 mm in length and 0.5 mm in diameter. The mantle was a dirty yellow in colour with a slight sheen, indicating hydrophobic properties. The ramification was straight and the mantle surface texture smooth to reticulate. Soil particles covered the entire root, concentrating in the base of the root branches. Rhizomorphs were dark yellow to tan in colour, but infrequent.
Anatomical morphology
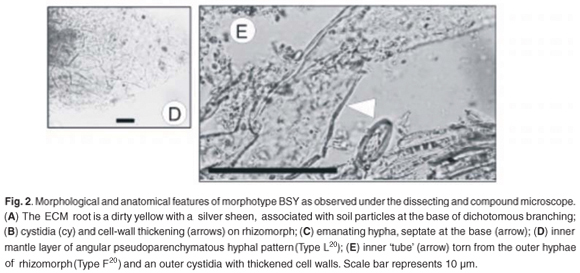
Emanating hyphae were infrequent, and were septate at the base of attachment (Fig. 2C). The outer mantle was arranged in a loose delicate plectenchymatous hyphal net, with hyphal dimensions of 1.8–2.9 µm in width and 10–11.6 µm in length. The inner mantle consisted of strongly angular pseudoparenchymatous cells (Fig. 2D) (Type L20), 6–10 µm in width and 9–11 µm in length. The rhizomorphs were highly differentiated (Type F20). There appeared to be an inner thin-walled tube surrounded by shorter thick-walled hyphae (Fig. 2E). Thick-walled cystidia were observed on the outer mantle and the outermost cells of the rhizomorph (Fig. 2B, E). The Hartig net was in a palmetti arrangement20 and penetrated 1 to 2 layers of the cortical cells.
Morphotype Black
Two fungal species, Thelephora terrestris (Fig. A in supplementary material online) and Phialocephala fortinii (Table 1), were identified from sequences of the Black morphotype. Morphotype Black was described from the Driekop study site and was recorded on 7.5% of all the ECM root tips examined.27
Root tip morphology
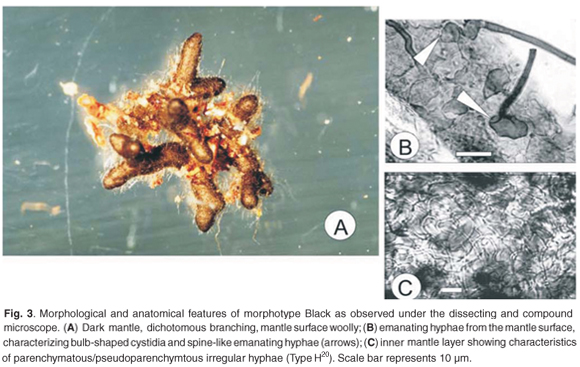
Branching of the root tips was unramified to short dichotomous and straight (Fig. 3A). The last order axis was 0.5–0.8 mm in length and 0.3–0.4 mm in diameter. The mantle was highly pigmented (Fig. 3A) and black in colour. The texture of the mantle surface was reticulate to woolly and was formed by emanating hyphae that then extended straight off the mantle to form spine-like structures. Soil particles were held between emanating hyphae. No rhizomorphs were observed, but emanating hyphae were found along the length of the ECM root tip, concentrating around younger, acropetal areas. Emanating hyphae formed fan-like morphologies.
Anatomical morphology
Emanating hyphae were frequent, dark and clamped. Cystidia were observed attached to the mantle by a globular base (Fig. 3B). Spine-like hyphae emanated from the globular structure, which was separated from the base by septa. The spine-like hyphae measured 75 µm in length from septum to septum and 3 µm in diameter. The mantle consisted of one layer of irregularly shaped transitional hyphae (Type H20) from plectenchymatous to pseudoparenchymatous arrangement (Fig. 3C). The hyphal dimensions in this layer were 4–8 µm in width and 7–10 µm in length. The Hartig net formed a typical palmetti arrangement20 and penetrated 3 or 4 layers of the root cortical cells.
Morphotype Orange
Morphotype Orange was identified from the ITS sequence as Amanita spissa (Table 1). Approximately 8.4% of all the ECM root tips examined from the Brooklands study sited were colonized by morphotype Orange.27
Root tip morphology
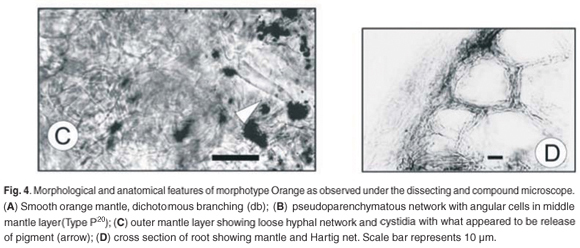
The branching associated with the Orange morphotype was short dichotomous to coralloid, and straight (Fig. 4A). The length and diameter of the last order axis were 0.5–0.7 mm and 0.2–0.5 mm, respectively. The mantle surface was smooth with a few hyaline emanating hyphae. The mantle was bright orange in colour, dulling to a darker orange with age. No rhizomorphs were observed.
Anatomical morphology
The outer mantle layer consisted of a delicate hyphal net, the hyphae ranging from 5–7 µm in width and 15–20 µm in length calculated from septum to septum. The net covered a middle mantle layer of pseudoparenchymatous cells with an angular configuration (Type P20) and hyphal dimensions of 8–15 µm in width and 14–18 µm in length (Fig. 4B). The inner layer was pseudoparenchymatous to plectenchymatous (Type H20), in another net-like arrangement with hyphal dimensions of 3–5 µm width and 10–25 µm length. Cystidia (Fig. 4C) with what appeared to be a pigment at the tip were observed. The Hartig net, in palmetti arrangement,20 penetrated 3 or 4 layers of cortical cells (Fig. 4D).
Morphotype Yelsil
Two species of ECM fungi were identified within this morphotype from the sequence identification, namely Thelephora terrestris (Fig. A) and a Scleroderma species (Table 1). The morphotype Yelsil was described from the Driekop study site and colonized 9.4% of all the ECM root tips examined at the site.27
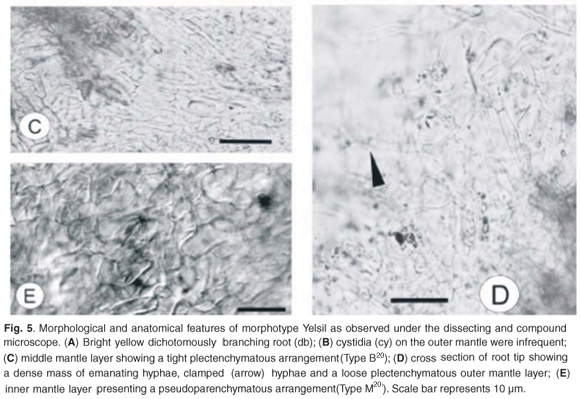
Root tip morphology
The root of the Yelsil morphotype was a luminescent lemon-yellow that branched dichotomously (Fig. 5A). The last order axis was 0.7–1.2 mm in length and 0.4–0.5 mm in width. The mantle was smooth with a few short, emanating hyphae.
Anatomical morphology
Three layers were observed in the mantle of the Yelsil morphotype. The outer layer is arranged in a loose plectenchymatous network with clamped emanating hyphae (Fig. 5D). The hyphal dimensions of the outer mantle layer were 2–3 µm in width and 23–40 µm in length measured between septa. Infrequent flask-shaped cystidia were observed on the outer mantle (Fig. 5B). The middle layer was tightly plectenchymatous (Type B20) and the hyphae were observed to be 3–4 µm in width and 10–15 µm in length (Fig. 5C). The inner layer was pseudoparenchymatous (Fig. 5E) (Type M20), with hyphal dimensions of 4–7 µm in width and 8–10 µm in length. Rhizomorphs were not observed. The Hartig net penetrated 2 layers of the root cortical cells.
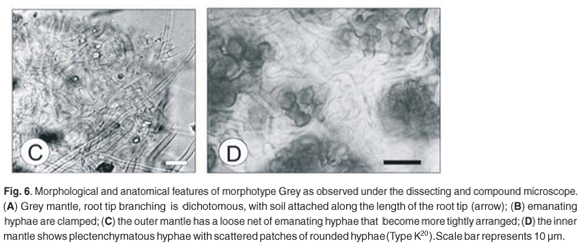
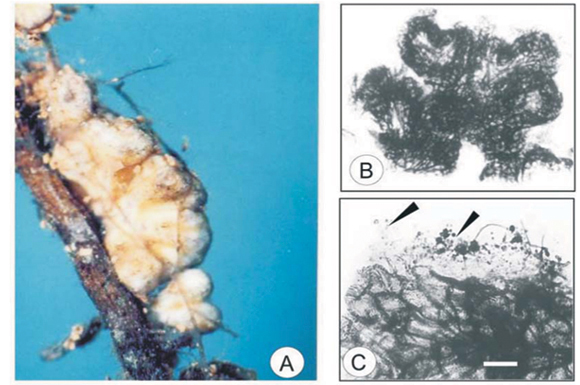
Morphotype Grey
This morphotype was identified as Albatrellus ovinus from the ITS sequence BLAST (Fig. C). The morphotype Grey was found only on approximately 2% of the ECM root tips examined in the Brooklands site.27
Root tip morphology
The Grey morphotype was dichotomously branched (Fig. 6A) with straight unramified ends. The length and width of the last order branches were 0.6–0.7 mm and 0.25–0.3 mm, respectively. The mantle surface was smooth to slightly woolly. Soil particles were attached to the length of the mantle.
Anatomical morphology
Two layers were observed in the mantle. The outer mantle layer has a plectenchymatous arrangement with squarrose branching (Fig. 6C) (Type E20), with hyphal length and width ranging from 15–20 µm and 3–4 µm, respectively. The inner mantle was plectenchymatous with scattered clusters of rounded hyphae (Type K20) (Fig. 6D) and had hyphal dimensions of 7–9 µm in length and 5–7 µm in width. No rhizomorphs were observed. Emanating hyphae were clamped and long (Fig. 6B), ranging between 60–75 µm in length from septum to septum and 3–4 µm in diameter. The Hartig net was in a palmetti arrangement,20 penetrating 2 layers of the root cortical cells.
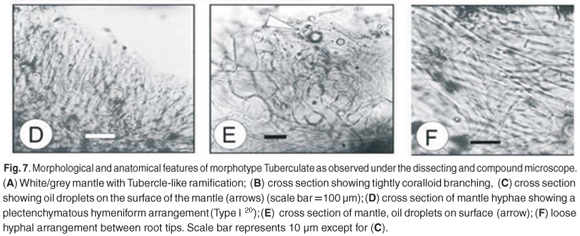
Morphotype Tuberculate
The Tuberculate morphotype did not amplify as an ECM fungus (Table 1) and was therefore not identified from the sequence data. This morphotype was encountered frequently at the Driekop study site and colonized approximately 9% of the ECM root tips examined there.27
Root tip morphology
The Tuberculate morphotype was greyish-white in colour (Fig. 7A) and was densely coralloid (Fig. 7B), becoming dark grey with age. The tubercle-like aggregations were roughly oval in shape and measured between 2–3.5 mm in width and 3–5 mm in length. The mantle was smooth to slightly reticulate.
Anatomical morphology
The mantle hyphae protruded perpendicularly to the root surface (Fig. 7D) in a plectenchymatous hymeniform arrangement (Type I20), 5.8–10.6 µm in width and 18.4–24.5 µm in length. The hyphal ends contained frequent amounts of oil droplets (Fig. 7C). From a surface view of the mantle, the hyphae appeared round (Fig. 7E). The hyphae of the mantle between the branches were tightly arranged and plectenchymatous (Fig. 7F). The rhizomorphs were observed as a mass of emanating hyphae with thickened hyphal walls. The Hartig net was in a typical palmetti arrangement,20 penetrating 4 to 5 layers of the root cortical cells.
Discussion
Molecular analysis
The BLAST alignments of ECM morphotypes (Table 1) resulted in interesting matches.
Discrepancies between the databases for two of the morphotype sequences, Tuberculate and White C, can be explained by the lack of particular species in either database. In the case of Tuberculate, the GenBank BLAST25 retrieved a non-ECM fungus, therefore the purely ectomycorrhizal UNITE26 database would not result in the same species. The Tuberculate sequence was aligned, on GenBank, with Scleroconidioma sphagnicola, which is a fungal pathogen that has been found in the moss Sphagnum fuscum.34 The alignment with S. sphagnicola indicated that the fungal DNA amplified was not from an ECM fungus but potentially that of a co-habiting fungus.
BLAST alignments of several morphotypes led to further investigation of Phialocephala fortinii and Hymenoscyphus ericae (Table 1). The latter represents a number of closely related taxa that form an aggregate; it was previously thought that these fungi were primarily ericoid mycorrhizas.35 Strains from this aggregate have recently been shown to produce ECM associations with Betula pubescens, Pi cea abies and Pinus sylvestris, and ericoid mycorrhizas with Vaccinium vitis-idaea.36 Villarreal-Ruiz et al.37 showed that the same strain of H. ericae can form both ECM and ericoid mycorrhizas with P. sylvestris and Vaccinium myrtillus, respectively. The results from the matches retrieved from GenBank of morphotype White C and Partial white A (see Table 2 in supplementary material online) match the morphotype sequences of Meliniomyces variabilis with 100% and 99% identities, respectively. M. variabilis forms part of the Rhizoscyphus ericae (= Hymenoscyphus ericae) aggregate and has recently been allocated to it own genus.38 The mycorrhizal status of this genus its unknown.38 The presence of two morphotypes for this species could be a result of different stages of colonization. The Partial white morphotype could represent only partial colonization of the root tip.
The Partial white C morphotype found at Driekop and the Black B morphotype found at Brooklands (Table 1) both have a BLAST match to Phialocephala fortinii on GenBank. The UNITE database produced a relatively poor match to H. ericae (92%). With a 98% identity to sequences from GenBank (Table 1), these morphotypes were identified as P. fortinii. This species is a dark septate endophyte (DSE), found globally, colonizing numerous plant species.The functional biology of DSEs, and whether they are pathogenic or symbiotic, is uncertain.39 The mechanism by which the hyphae colonize the root was examined by Yu et al.39 and found to be similar to that of arbuscular mycorrhizal fungi. Additionally, this genus is closely related to the ericoid ectomycorrhizal genus Hymenoscyphus.35
Five morphotypes, Black A, Fwhite, Yelsil C, Yelsil A and Partial white B, were matched to Thelephora species, a resupinate/ bracket ectomycorrhizal fungus (Fig. A). Fwhite was poorly aligned (93%) with a different species, T. coerulea, according to the UNITE database, which corresponds with the inconclusive BLAST result from GenBank. Comparisons between the alignments of all the morphotypes and their taxonomic position (Fig. A) indicate that Fwhite is a different species, grouping with Thelephora penicillata. The presence of T. terrestris can be confirmed by above-ground sporocarp collections27 and has been documented both in the field and in nurseries in South Africa.2,4
Xerocomus badius was represented by three morphotypes (Fig. B) according to the concurring results of both GenBank and UNITE databases. The morphotypes Dull Orange and Beige resulted in strong alignments with X. badius, with 99% identity. BSY2 aligned poorly with X. badius (94%), suggesting that this morphotype is closely related, but not the same species. The taxonomic placement of this morphotype did, however, match within the X. badius group in the derived tree (Fig. B), indicating that more-rigorous analysis is required to determine the taxonomic species.
The Grey morphotype was strongly aligned with Albatrellus ovinus on the GenBank and UNITE databases (Fig. C), with 99% and 98% identities, respectively. The taxonomic position of this morphotype as A. ovinus was confirmed in the derived tree (Fig. C online), where this morphotype grouped with a known sequence of this species. However, no sporocarp collections of this species have ever been recorded in South Africa. Morphotype DSY was tentatively identified as Clavulina cristata, as the ITS sequence aligned with this species in relation to GenBank and UNITE sequences, 96% and 95% identity, respectively. The species' position was verified in the derived tree (Fig. D online), where DSY grouped with known sequences of C. cristata.
Four morphotypes have been identified as Amanita spissa from the GenBank and UNITE databases: Pink, White A, BSY 1, 3, 4 and 5, BSY and Orange (Table 1). BLAST alignments of morphotypes BSY4 and White A resulted in lower percentage identities (97% and 98%, respectively, from UNITE) to A. spissa. GenBank gave a match of A. rubescens with White A. This mismatch between the two databases may be a result of misidentified sequence material being deposited in them. It is important to note that all three of these species are closely related and have been placed in the Sect Validae (Fr.) Quél.40 Within the genus, the ITS region displays high rates of divergence that do not allow proper species distinction. For this reason, the ITS regions may not be sufficiently informative for identification to species level. The LSU region, which is more conserved, is currently used in the literature for the phylogenetic analysis of this genus.40 Amanita spissa and A. rubescens have been recorded by Lundquist4 and Van der Westhuizen and Eicker5 in Pinus stands in South Africa.
The morphotypes White B and Yelsil B were poorly aligned with Scleroderma bovista on the GenBank and UNITE databases (94% and 90–92%, respectively; Table 1), which indicates that the identity of the ECM fungi falls within the genus, but could potentially be a different species. Sporocarps of S. citrinum were found at high frequency at the Brooklands study site,27 confirming the above statement.
The morphotype Beige/white displayed a fair alignment (97% identity) on GenBank, with Heterobasidion annosum (Table 1). This species is a wide-ranging, basidiomycetous plant pathogen that has been studied at an infected site of Pinus sylvestris and Betula pendula,41 and is considered to be highly infective over extended periods of time. The fungi colonize fresh plant tissue normally exposed as stumps during felling and infects surrounding plants through the rhizosphere. The consequences of infection that are experienced are reduced wood quality, growth losses and mortality of Pinus sylvestris.42 No record of this fungal infection has been made in South Africa.
It is clear that there is some genotypic variation within morphotypes (e.g. the BSY group), and yet some conformity between different morphotypes (e.g. Pink, BSY and Orange all resolve as Amanita spissa) (Table 2). Mah et al.43 observed similar results in a study where 22 genotypes were derived from eight morphotypes. A genotypic analysis of morphotypes found by Erland et al.44 recorded eight species within six morphotypic groups. Phenotypic variations may be a result of the morphological changes of the ECM root tip that occur during the ageing process. Mantle colour may change or may be altered in different environmental conditions. However, Sakakibara et al.45 found an average 93% genotypic conformity with the eight morphotypes examined and employed techniques of anatomical analysis which, although more accurate, may be impractical for large-scale population surveys. A possible 11 species may be identified from the BLAST results of the root tip DNA sequences examined in this study.
Morphological descriptions
The most informative features of the ECM root tip morphology observed were the cystidia and rhizomorphs (when present) and patterns discernible in the mantle layers. Dichotomous branching of the root was not a unique distinguishing feature, which may have been due to the natural occurrence of dichotomous branching of Pinus roots in the abscence of ECM fungi.46 This study described seven morphotypes from P. patula plantations in Sabie.
The morphotype White was identified as Scleroderma citrinum according to the colour atlas key,20 in that clamps were present (Fig. 1B) and contact-clamps and contact septa were lacking. There are, however, differences in the root ramification. The morphotype examined in this study branched dichotomously (Fig. 1A), whereas the branching presented in the colour atlas is monopodial-pyramidal. The White morphotype is similar to morphotype WITS 01, also described as S. citrinum, by Dames et al.,8 from P. patula, but differs in the colour of the rhizomorph. Dames et al.8 found the rhizomorphs to be yellow, but notes that the variation in colour could be attributed to age. The white ECM root identified by Mohan et al.,12 from P. patula as Amanita muscaria (L.) Hook, was compared with the morphotype in this study, but differed in that the specimen in this investigation contained clamp connections and lacked tannin cells in the mantle slide mounts.
The mantle morphology of morphotype BSY is remarkably similar to Orange in that the mantle has an angular pseudoparenchymatous hyphal arrangement in the middle to inner layer. This hyphal arrangement led to the comparison with Amanita citrina in the colour atlas.20 Features that were common between the BSY and A. citrina were an outer plectenchymatous hyphal arrangement and a strongly angular pseudoparenchymatous hyphal layer (Fig. 2D). The rhizomorphs also displayed a common differentiation consisting of a large inner tube (Fig. 2E). The morphotypes differed in the mantle layer, where an inner pseudoparenchymatous layer was described for A. citrina. Also, the hyphae in the outer mantle of the ECM morphotype described in this study consisted of frequent cystidia and hyphae with thickened cell walls (Fig. 2B), which are not observed with A. citrina. This indicates that BSY is possibly a different species of the same genus.
The Black morphotype was identified as a tomentelloid ECM fungus. Keying out to genus or species was not possible. Comparisons with Tomentella species were made, although occasional specimens of Cenococcum geophilum were observed, however, which were characterized by stellate pseudoparenchymatous mantles.47 Other, rarely found, black root tip types were comparable to Hymenoscyphus ericae and were characterized by plectenchymatous hyphal arrangement in the mantle.48 The similar morphologies of these black root tips may result in morphotyping inaccuracies. The morphotype Black in this study did compare with Tomentella albomarginata in that emanating hyphae extended from globular hyphae in the mantle (Fig. 3B). Although not all the mantle layers described from T. albomarginata49 correlate with the current morphotype, a similar pattern of irregular, transitional pseudoparenchymatous to plectenchymatous type hyphae were comparable.
The strong angularity of the hyphae in the pseudoparenchymatous mantle layer was comparable with the morphotype Orange and Amanita genera in the colour atlas.20 Although there is only one Amanita species illustrated, A. citrina,50 the key to mycobionts of Pinus, presents two Amanita species. A comparison of mantle structure between A. citrina and morphotype Orange can be made. Both species share loose, ring-like plectenchymatous surface hyphae (Fig. 4C) and a definite angular pseudoparenchymatous middle layer (Fig. 4B), as well as clamped emanating hyphae. The only differences that were noted between morphotype Orange and A. citrina were the root colour, the lack of emanating hyphal structures, and the presence of cystidia associated with Orange. The deduction therefore is that morphotype Orange is not A. citrina, but possibly a species within the same genus.
The morphotype Yelsil was keyed out in the colour atlas20 as Thelephora terrestris and was compared with the photographic illustration provided. The description of T. terrestris, in association with Picea abies (L.) Karst.,51 differed little from the root tip in this study in terms of mantle morphology. The root ramification was the only differing feature in that the morphotype described in this study branched dichotomously as opposed to monopodial-pinnate, but this may be a result of the dichotomous root branching exhibited in Pinus species, as discussed earlier. The Yelsil morphotype characterized in this study was also comparable to the T. terrestris described on P. patula by Mohan et al.11 T. terrestris is a common and widely distributed ECM fungus and known as an aggressive colonizer, especially in nurseries in South Africa, the presence of which is therefore not surprising.52
The morphotype Grey is not represented in the Pinus key in the colour atlas.20 It was, nevertheless, compared to the Albatrellus ovinus (Schaeff.: Fr.) Kotl. & Pouzar plate.53 Similarities and dissimilarities were noted. Both specimens were grey in colour, and were described as having a loose layer of emanating hyphae (Fig. 6C). In both cases the outer mantle was plectenchymatous, the arrangement becoming more tightly packed towards the inner mantle (Fig. 6D). The difference between them was the presence of cystidia, which were not reported from the Grey morphotype in our study. The branching reported in association with Picea abies53 was represented as monopodial, where the present description associated with Pinus patula was dichotomously branched. The species identification of this ECM fungal morphotype is therefore inconclusive. Albatrellus species have not been recorded in South Africa, and therefore there are no sporocarp data to confirm or provide insight into the presence of this species.
The morphotype Tuberculate was keyed out as Rhizopogon roseolus (Corda) TM Fries, in that the ECM branching was tightly coralloid (Fig. 7B), the emanating hyphae contained oil droplets (Fig. 7C) and were not embedded in gelatinous matrix, the mycorrhizal system was smooth and clamps were absent.20 The morphotype may also have been a tubercle-forming Suillus species, as ECM roots formed by this genus are commonly tuberculate, e.g. S. pictus.54,55
Concluding remarks
The characterization of seven morphotypes has led to the identification of Scleroderma citrinum, two Amanita species (morphotypes BSY and Orange), a tomentelloid species, Thelephora terrestris, and a possible suilloid and Albatrellus species. ECM root morphologies documented in this investigation can be used to direct future ECM root studies in South Africa, creating a database of recorded South African species with which to consult. With the exception of Albatrellus, all the genera discussed—Amanita, Scleroderma, Suillus and Thelephora—are well represented by several species in South Africa and have been recorded in association with Pinus species.4,5 It is necessary, though, to emphasize that root tip morphology can be highly variable, especially with regard to the age of ECM root tips, the stage of colonization, as well as geographic differences within the same species.
The identification and characterization of the ECM fungi occurring in South Africa is by no means complete, with only seven morphotypes having been described. The other morphotypes observed were infrequent or rare and are indicative of greater fungal species diversity than previously suspected.27 Morphological characterization of ECM roots is necessary for further understanding within this field and may be enhanced by ECM synthesis from sporocarp material or by tracing ECM via rhizomorphs and hyphae from the sporocarp to the root.
We thank Kata Ihrmark for her guidance and for sharing knowledge that was crucial for the molecular analysis of all the fungal samples. We would also like to acknowledge funding from SIDA and the NRF, whose support provided the means to achieve the research presented.
1. Smit W. and Pitcher M. (2003). A case study on ensuring sustainable management of planted forests: The economic, social and environmental role of commercial plantations in South Africa. Ensuring SRM session. UNFF Intersessional Experts Meeting on the Role of Planted Forests in Sustainable Forest Management, 24–30 March 2003, New Zealand, Paper 15. [ Links ]
2. Khalil S., Labuschagne N. and Wingfield M.J. (1999). Pinus patula establishment problem associated with poor ectomycorrhizal development in previously cultivated soils. S. Afr. For. J. 186, 59–65. [ Links ]
3. Doidge E. (1950). The South African Fungi and Lichens to the end of 1945. Bothalia 5, 1–1094. [ Links ]
4. Lundquist J.E. (1986). Fungi associated with Pinus in South Africa, Part 1. The Transvaal. Suid-Afrikaanse Bosboutydskrif 138, 1–13. [ Links ]
5. Van der Westhuizen G.C.A. and Eicker A. (1987). Some fungal symbionts of ectotrophic mycorrhizae of pines in South Africa. S. Afr. For. J. 143, 20–24. [ Links ]
6. Dahlberg A. (2001). Community ecology of ectomycorrhizal fungi: an advancing interdisciplinary field. New Phytol. 150, 555–562. [ Links ]
7. Jonsson L., Dahlberg A. Nilsson M. Zackrisson O. and Kårén O. (1999). Ectomycorrhizal fungal communities in late-successional Swedish boreal forests, and their composition following wildfire. Mol. Ecol. 8, 205–215. [ Links ]
8. Dames J.F., Straker C.J. and Scholes M. (1999). Ecological and anatomical characterisation of some Pinus patula ectomycorrhizas from Mpumalanga, South Africa. Mycorrhiza 9, 9–24. [ Links ]
9. Carlson C. (1992). Effects of acidic precipitation on calcium and magnesium uptake by Pinus patula. M.Sc. thesis, University of the Witwatersrand, Johannesburg. [ Links ]
10. Carlson C. (1994). The influence of fertilisation on ectomycorrhizal colonisation of Pinus patula roots. Suid-Afrikaanse Bosboutydskrif 171, 1–6. [ Links ]
11. Mohan V., Natarajan K. and Ingleby K. (1993). Anatomical studies on ectomycorrhizas. I, The ectomycorrhizas produced by Thelephora terrestris on Pinus patula. Mycorrhiza 3, 39–42. [ Links ]
12. Mohan V., Natarajan K. and Ingleby K. (1993). Anatomical studies on ectomycorrhizas. II, The ectomycorrhizas produced by Amanita muscaria, Laccaria laccata and Suillus brevipes on Pinus patula. Mycorrhiza 3, 43–49. [ Links ]
13. Bruns T.D. and Gardes M. (1993). Molecular tools for the identification of ectomycorrhizal fungi – taxon specific oligonucleotide probes for suilloid fungi. Mol. Ecol. 2, 233–242. [ Links ]
14. Kårén O., Högberg N., Dahlberg A., Jonsson L. and Nylund J. (1997). Inter- and intraspecific variation in the ITS region of rDNA of ectomycorrhizal fungi in Fennoscandia as detected by endonuclease analysis. New Phytol. 136, 313–325. [ Links ]
15. Martin F., Delaruelle C. and Ivory M. (1998). Genetic variability in intergenic spacers of ribosomal DNA in Pisolithus isolates associated with pine, eucalyptus and Afzelia in lowland Kenyan forests. New Phytol. 139, 341–352. [ Links ]
16. Timonen S., Tammi H. and Sen R. (1997). Characterization of the host genotype and fungal diversity in Scots pine ectomycorrhiza from natural humus microcosms using isozyme and PCR–RFLP analyses. New Phytol. 135, 313–323. [ Links ]
17. Chamberlain D., Essop H., Hougaard C., Malherbe S. and Walker R. (2005). G:nensis part III: Technical note and appendices. Online: www.forestry.co.za/ fsa/ViewFile?file_id=91 [ Links ]
18. Weather SA. Online: www.weathersa.co.za/Climat/Climatestats/Nelspruit Stats.jspweather SA [ Links ]
19. Brundrett M., Bougherm N. Dell B. Grovem T. and Malajczuk N. (1996). Working with Mycorrhizas in Forestry and Agriculture. Australian Centre for International Agricultural Research, Canberra. [ Links ]
20. Agerer R. (1987–2002). Colour Atlas of Ectomycorrhizae. Einhorn-Verlag, Schwäbisch Gmünd, Germany. [ Links ]
21. Zeze A., Ophel-Kellerm K. and Smith S. (1998). DNA isolation and development of molecular tools for identification and detection of VA mycorrhizal fungi. In Mycorrhizal Manual, ed. A. Varma, pp. 373–385. Springer, Berlin. [ Links ]
22. Gardes M. and Bruns T.D. (1993). ITS primers with enhanced specificity for basidiomycetes – application to the identification of mycorrhizae and rusts. Mol. Ecol. 2, 113–118. [ Links ]
23. White T.J., Bruns T., Lee S. and Taylor J. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications, eds. M.A. Innis, D.H. Gelfand J.J. Shinsky T.J. White, pp 315–322. Academic Press, San Diego. [ Links ]
24. Carlile M.J., Watkinson S. Gooday G.W. and Watkinson S.C. (2001). The Fungi, 2nd edn. Academic Press, London. [ Links ]
25. Altschul S.F., Madden T.L. Schaffer A.A. Zhang J. Zhang Z. Miller W. and Lipman D. (1997). Gapped BLAST and PSIBLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402. [ Links ]
26. Kõljalg U., Larsson K-H. Abarenkov K. Nilsson R.H. Alexander I.J. Eberhardt U. Erland S. Hoiland K. Kjoller R. Larsson E. Pennanen T. Sen R. Taylor A.F.S. Tedersoo L. Vrålstad T. and Ursing B.M. (2005). UNITE: a database providing web-based methods for the molecular identification of ectomycorrhizal fungi. New Phytol. 166, 1063–1068. [ Links ]
27. Hawley G.L. (2008). Ectomycorrhizal characterisation, species diversity and community dynamics of Pinus patula Schelcht. et Cham. plantations. Ph.D. thesis, Rhodes University, Grahamstown. [ Links ]
28. Chenna R., Sugawara H. Koike T. Lopez R. Gibson T.J. Higgins D.G. and Thompson J.D. (2003). Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 31, 3497–500. [ Links ]
29. Taylor A.F.S. and Alexander I.J. (1989). Ectomycorrhizal synthesis with an isolate of Russula aeruginea. Mycol. Res. 92, 103–107. [ Links ]
30. Massicotte H.B., Melville L.H. Peterson R.L. and Molina R. (1999). Biology of the ectomycorrhizal fungal genus, Rhizopogon. IV. Comparative morphology and anatomy of ectomycorrhizas synthesized between several Rhizopogon species on Ponderosa pine (Pinus ponderosa). New Phytol. 142, 355–370. [ Links ]
31. Kraigher H., Agerer R. and Javornik B. (1995). Ectomycorrhizae of Lactarius lignyotus on Norway spruce, characterisation by anatomical and molecular tools. Mycorrhiza 5, 175–180. [ Links ]
32. Smith S.E. and Dickson S. (1997). VA Mycorrhizas: Basic Research Techniques. Cooperative Research Centre for Soil and Land Management, Adelaide. [ Links ]
33. Koske R.E. and Gemma J.N. (1989). A modified procedure for staining roots to detect VA mycorrhizas. Mycol. Res. 92, 486–489. [ Links ]
34. Tsuneda A., Chen M.H. and Currah R.S. (2001). Characteristics of a disease of Sphagnum fuscum caused by Scleroconidioma sphagnicola. Can. J. Bot. 79, 1217– 1224. [ Links ]
35. Grünig C.R. (2003). Population biology of the tree-root endophyte Phialocephala fortinii. Ph.D. thesis (Natural Sciences), Swiss Federal Institute of Technology, Zürich. [ Links ]
36. Vrålstad T., Schumacher T. and Taylor A.F.S. (2002). Mycorrhizal synthesis between fungal strains of the Hymenoscyphus ericae aggregate and potential ectomycorrhizal and ericoid hosts. New. Phytol. 153, 143–152. [ Links ]
37. Villareal-Ruiz L., Anderson I.C. and Alexander I.J. (2004). Interaction between an isolate from the Hymenoscyphus ericae aggregate and roots of Pinus and Vaccinium. New Phytol. 164, 183–192. [ Links ]
38. Hambleton S. and Sigler L. (2005). Meliniomyces, a new anamorph genus for root-associated fungi with phylogenetic affinities to Rhizoscyphus ericae (= Hymenoscyphus ericae), Leotiomycetes. Stud. Mycol. 53, 1–27. [ Links ]
39. Yu T., Nassuth A. and Peterson R.L. (2001). Characterisation of the interaction between the dark septate fungus Phialocephala fortinii and Asparagus officinalis roots. Can. J. Microbiol. 47, 741–753. [ Links ]
40. Weiβ M., Yang Z.L. and Oberwinkler F. (1998). Molecular phylogenetic studies in the genus Amanita. Can. J. Bot. 76, 1170–1179. [ Links ]
41. Lygis V., Vasiliauskas R. and Stenlid J. (2004). Planting Betula pendula on pine sites infested by Heterobasidion annosum: disease transfer, silvicultural evaluation, and community of wood-inhabiting fungi. Can. J. For. Res. 34, 120–130. [ Links ]
42. Pukkula T., Möykkynen T., Thor M., Rönnberg J. and Stenlid J. (2005). Modelling infection and spread of Heterobasidion annosum in even-aged Fennoscandian conifer stands. Can. J. For. Res. 35, 74–84. [ Links ]
43. MahK., Tackaberry L.E., Egger K.N. and Massicotte H.B. (2001). The impacts of broadcast burning after clear-cutting on the diversity of ectomycorrhizal fungi associated with hybrid spruce seedlings in central British Columbia. Can. J. For. Res. 31, 224–235. [ Links ]
44. Erland S., Jonsson T., Mahmood S. and Finlay R.D. (1999). Below-ground ectomycorrhizal community structure in tow Picea abies forests in southern Sweden. Scand. J. For. Res. 14, 209–217. [ Links ]
45. Sakakibara S.M., Jones M.D., Gillespie M., Hagerman S.M., Forrest M.E., Simard S.E. and Durall D.M. (2002). A comparison of ectomycorrhiza identification based on morphotyping and PCR–RFLP analysis. Mycol. Res. 106, 868–878. [ Links ]
46. Kaska D.D., Myllylä R. and Cooper J.B. (1999). Auxin transport inhibitors act through ethylene to regulate dichotomous branching of lateral root meristems in pine. New Phytol. 142, 49–58. [ Links ]
47. Agerer R. and Gronbach E. (1988). Cenococcum geophilum. In Colour Atlas of Ectomycorrhizae, ed. R. Agerer, plate 11. Einhorn-Verlag, Schwäbisch Gmünd, Germany. [ Links ]
48. Brand F., Gronbach E. and Taylor A.F.S. (1992). Hymenoscyphus ericae. In Colour Atlas of Ectomycorrhizae, ed. R. Agerer, plate 73. Einhorn-Verlag, Schwäbisch Gmünd, Germany. [ Links ]
49. Agerer R. (1996). Tomentella albomarginata. In Colour Atlas of Ectomycorrhizae, ed. R. Agerer, plate 111. Einhorn-Verlag, Schwäbisch Gmünd, Germany. [ Links ]
50. Mleczko P. (2002). Amanita citrina. In Colour Atlas of Ectomycorrhizae, ed. R. Agerer, plate 159. Einhorn-Verlag, Schwäbisch Gmünd, Germany. [ Links ]
51. Agerer R. and Weiss M. (1990). Thelephora terrestris. In Colour Atlas of Ectomycorrhizae, ed. R. Agerer, plate 48. Einhorn-Verlag, Schwäbisch Gmünd, Germany. [ Links ]
52. Van Greuning J. and van der Westhuizen G. (1990). An ectomycorrhizal fungus of pine seedlings in an Eastern Transvaal Nursery. Suid-Afrikaanse Bosboutydskrif 155, 1–4. [ Links ]
53. Agerer R. (1996). Albatrellus ovinus. In Colour Atlas of Ectomycorrhizae, ed. R. Agerer, plate 91. Einhorn-Verlag, Schwäbisch Gmünd, Germany. [ Links ]
54. Olsson P.A., Münzenberger B. Mahmood S. and Erland S. (2000). Molecular and anatomical evidence for a three-way association between Pinus sylvestris and the ectomycorrhizal fungi Suillus bovinus and Gomphidius roseus. Mycol. Res. 104, 1372–1378. [ Links ]
55. Randall B.L. and Grand L.F. (1986). Morphology and possible mycobionts (Suillus pictus) of a tuberculate ectomycorrhiza on Pinus strobus. Can. J. Bot. 64, 2182–2191. [ Links ]
Received 13 April 2007. Accepted 5 July 2008.
This article is accompanied by a supplementary table and figures online at www.sajs.co.za
* Author for correspondence. E-mail: j.dames@ru.ac.za
Hawley G.L., Taylor A.F.S. and Dames J.F (2008). Ectomycorrhizas in association with Pinus patulain Sabie, South Africa. S. Afr. J. Sci.104, 273–283.