Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
South African Journal of Science
On-line version ISSN 1996-7489
Print version ISSN 0038-2353
S. Afr. j. sci. vol.104 n.7-8 Pretoria Sep./Aug. 2008
REVIEW ARTICLES
A role for barcoding in the study of African fish diversity and conservation
Ernst R. SwartzI, *; Monica MwaleI; Robert HannerII
ISouth African Institute for Aquatic Biodiversity, Private Bag 1018, Grahamstown 6140, South Africa
IICanadian Centre for DNA Barcoding, Biodiversity Institute of Ontario, University of Guelph, Guelph, Canada
ABSTRACT
Africa has a rich diversity of marine and freshwater fishes, but very little taxonomic expertise or funding to describe it. New approaches to using modern technology, such as DNA barcoding, can facilitate collaboration between field biologists, reference collections and sequencing facilities to speed up the process of species identification and diversity assessments, provided specimen vouchers, tissues, photographs of the specimen and DNA sequences (barcodes) are clearly linked. The FISH-BOL project in Africa aims to establish a collaborative Pan-African regional working group to facilitate barcoding of fish across the continent and the surrounding FAO marine regions. This is being established through existing African biodiversity networks and global biodiversity programmes that are already in place. Barcoding is expected to inform African fisheries management and conservation through more accurate identification of species and their different life-history stages, by speeding up biodiversity assessments. Barcoding is an important development, contributing towards an evolutionary history perspective on which to base Africa's conservation strategies.
Introduction
Biodiversity assessments and implementation of conservation actions worldwide are hampered by slow progress in taxonomic research, termed the Taxonomic Impediment.1 The existing workforce of taxonomists cannot cope with the overwhelming need for basic field surveys, species descriptions and systematic revisions to provide basic information for conservation planning. In addition, Costa and Carvalho2 suggested that few taxonomists are able to distinguish critically between more than 1000 taxa. The reality facing taxonomy is that there may be more species that remain to be discovered on Earth than those that have already been described.3 It is estimated that 1.4–1.8 million species have been described4–6 out of a possible total of approximately 7–15 million.6 Species and populations are going extinct at an alarming but poorly understood rate.7,8 Many species may be going extinct before they can be identified or described. This presents a problem for conservation planning and prioritization, obviously because species that have not been identified cannot be protected effectively.
This problem is especially evident in Africa, whose rich biodiversity is poorly known. Taxonomic expertise is lacking even for major and commercially important groups. The few taxonomists who are working in developing countries, home to more than 95% of globally described species, find it difficult to access basic taxonomic information such as species descriptions.9 Where taxonomic keys are available, they are rarely revised and often inadequate to identify specimens unambiguously to the species level.10 A further problem in Africa is that funding for taxonomic research and the museums or facilities that host biodiversity collections has been poor or non-existent, few new taxonomists are being trained, retirees are often not replaced and salaries are not competitive, such that the overall taxonomic expertise is declining.11,12
There have been strident calls on the taxonomy community to embrace new technologies and to form networks to speed up the description of biodiversity13,14 and to improve our ability to identify species.2,15–18 How this could be achieved without compromising rigorous taxonomic research principles is, however, questioned.19 There is legitimate concern that too little money is being spent on morphological taxonomy compared to molecular studies.20 A purely DNA taxonomy approach (e.g. Blaxter21), not to be confused with DNA barcoding,22 is too simplistic in our view. DNA taxonomy can, however, provide additional characters for discrimination, especially in cases where other characters vary among species and are thus difficult to interpret. Therefore we do not agree with de Carvalho et al.19 that a molecular approach will do little to address the real problems in taxonomy. In our experience, most traditional taxonomists welcome the opportunity to refer to molecular data, but DNA taxonomy should not replace taxonomic research that can be based on multiple character data sets.
Nucleotide sequence divergence in short, standardized gene regions (DNA barcodes) can be used to identify known species and facilitate the discovery of new ones.15,23 Mitochondrial DNA is a valuable marker in population genetic or phylogeographic studies because it is maternally inherited, evolves rapidly and recombination is rare or absent.24 Therefore, a part of the mitochondrial cytochrome oxidase subunit I (COI) has been chosen as a standard gene region for barcoding animals. Although there was no a priori reason for choosing COI among the 13 protein-coding mitochondrial genes,15 it has the advantage of having robust universal primers that can recover the 5' end of COI of most animal species. Barcoding of species has become cheaper through the technological advances made by other molecular programmes, especially the human genome project.25 Rapid barcoding and comparison with the growing database of COI sequences26 will increase the speed of identification of newly collected or unknown specimens. This may focus taxonomists' attention on unidentified lineages, and with the addition of morphological and other taxonomically relevant data, could lead to a faster rate of species description.
The Consortium for the Barcode of Life (CBOL) was established as a growing coalition of biodiversity organizations interested in developing barcoding as a global standard for DNA-based species identification. CBOL promotes FISH-BOL as an international campaign to barcode all marine and freshwater fish. FISH-BOL consists of ten regional working groups representing Africa, Australia, Oceania/Antarctica, the Americas (North, Central and South America), Europe and Asia (India, North East Asia, and South East Asia). Each of these regional working groups is charged with the task of organizing support and participation to barcode the ichthyofauna within their region, based on expert-identified voucher specimens.
The main purpose of barcoding is to identify species reliably, increase the rate of species discovery and raise the profile of taxonomic research.27 The vision for the African FISH-BOL programme is to co-ordinate a Pan-African coalition generating taxonomic reference data in a common format to fulfil the common goal of barcoding all Africa's fishes. The aim is eventually to establish a coherent information framework for revisionary fish taxonomy and diversity studies. This can only be achieved through the production of a dynamic catalogue of African fishes, complete with monitoring tools (such as barcodes) that can facilitate a greater understanding of natural history through molecular ecology studies.
In this review, we illustrate the role of FISH-BOL (www. fishbol.org) in taxonomic and diversity studies in Africa, with applications in conservation, fisheries management, identification of alien invasive fishes, law enforcement and resource management. We show that FISH-BOL will lead to greater access to African fish diversity information through a global alignment of resources and linkages to other global networks. We summarize the functioning and aims of the African FISH-BOL programme and draw conclusions about its potential role in African fish diversity assessments and conservation.
Why barcode fish in Africa?
Barcoding has an important role to play in phylogenetic and phylogeographic studies,28 especially in terms of generating hypotheses. With barcoding being applied broadly to African fishes, we anticipate it will focus attention on a plethora of species identification problems, which can then be resolved with specific morphological, phylogeographic and phylogenetic studies. Several fish genetic studies in Africa have already employed mitochondrial DNA sequence data.29–31 These studies are, however, not comparable for the purposes of species identification as different regions of the mitochondrial genome were used. The African projects proposed by FISH-BOL are also on a larger scale than previous molecular studies.
These projects are designed to identify species through a composite database based on the mitochondrial COI region of as many species as possible, to provide a hitherto unavailable 'horizontal genomics' perspective.28 This COI database can form the foundation for more detailed phylogeographic studies. In their planning phases, preliminary analyses of phylogenetic studies will be able to benefit from this species-rich COI database in important ways. One such benefit will be in selecting in- or outgroups suitable for molecular phylogenetic analyses in a particular study.
The identification of species remains, however, the main objective of barcoding. Owing to maternal inheritance, mtDNA reflects only a portion of the ancestry of a species. Additional information is necessary to test whether divergent mtDNA lineages constitute separate species.32–34 We therefore view provisional species assignments (that is, not formal descriptions) based only on mtDNA as working hypotheses. As new collections are made, individuals suspected of being unique (especially of equivocal taxonomic status) can be barcoded. If they are divergent or form part of a unique lineage, they can then be flagged for further taxonomic investigation.
The identification process using COI sequence data for fish is looking promising, as supported by recent examples of its application. Hubert et al.35 were able to distinguish 93% of 190 Canadian freshwater fishes using DNA barcodes. The inability of COI barcodes to distinguish the remaining 7% (13 species) was attributed to recent speciation, introgressive hybridization (only two species) or the possibility that some of these taxa may not be different species. Ward et al.36 found that all the 207 marine species that they sequenced from Australia could be discriminated using the mitochondrial DNA COI gene, with 98% of these being amplified using only one primer set. They found that only two sequences out of 754 tested appeared in the 'wrong' species identification because of hybridization or incorrect original identification. No transfers of mitochondrial COI sequences into the nuclear genome were observed. Pseudogenes or NUMTs may not be a serious problem in fish, as Venkatesh et al.37 did not find any mitochondrial pseudogenes in the Fugu nuclear genome, but Waters and Wallis38 did find pseudogenes in Galaxias from New Zealand. Pseudogenes did not significantly affect the barcoding of birds.39 The study of Ward et al.36 adds empirical evidence to expanding documentation, which shows that barcoding can discriminate between species, for example North Atlantic sharks,40 North American birds,41 neotropical butterflies,42 Arctic springtails,43 marine bryozoans,23 and Indo-Pacific coral reef stomatopods.44
The potential contributions of barcoding in African fisheries management should not be underestimated. Global capture fisheries yielded 95 million tonnes in 2004 with an estimated first-sale value of US$84.9 billion.45 Marine and freshwater ecosystems are under threat globally because of over-exploitation, alien invasion, habitat destruction, pollution and climate change.46,47 Accurate identification is essential in assessing the distribution, utilization and spread of fish species, therefore, prioritizing fish biodiversity assessments using barcoding is not only a global but also an African necessity.
In 2006, a CBOL meeting in Cape Town (South Africa) identified barcoding in fish stock management in African countries as a priority. Barcoding has an application in forensics for African fisheries departments to prosecute illegal fishing, even after processing has already occurred, because only a small part of the fish is needed for DNA analysis. An example is the illegal shark-fin trade, which is a major problem in most coastal African countries. Barcoding can identify different shark species,40 suggesting a potential role in law enforcement once a reference database has been established. An improved species identification system will contribute to better and more accurate statistics on stock trends, requested by the FAO since the 1960s.48
Barcoding may help link life-history stages, since little taxonomic information and few descriptions are available for fish larvae and eggs (e.g. refs 49–51). Incorrect identification of larvae will mislead understanding of speciation, diversity, niche partitioning and other ecosystem features.51 Barcoding can also be used in feeding biology to identify gut contents of fishes, especially organisms that are partially digested.52 It may also help prevent the spread of alien fish species, because even eggs that are being translocated can be barcoded, and alien parasites can be identified in a similar manner to the study of Siddall and Budinoff53 on alien leeches in North America. This is important for freshwater systems globally, since the continuous introduction of alien fish species has been identified as the most critical threat to freshwater biodiversity.46
Molecular studies have improved our understanding of aquatic biodiversity; for example, 15 historically isolated lineages were identified within the seven described Pseudobarbus species that occur in Lesotho and in the Cape Floristic Region in South Africa.54 Although this knowledge alone does not solve the taxonomic problems in the group, it makes a critical contribution to our understanding of the underlying taxonomic diversity. This genetic information has provided especially valuable insights into the formative processes that shaped the evolutionary history of these lineages.31 Ultimately, such studies may contribute to biodiversity conservation planning before formal taxonomic descriptions are completed or even before formal publication of results. Such a precautionary approach is justified, especially for freshwater aquatic organisms that seem to have higher levels of population differentiation than most marine and terrestrial species. In addition, taxonomic groups with the highest proportion of IUCN assessed threatened species (www.redlist.org), are those that rely on freshwater habitats.47 With more than half the accessible surface freshwater being used for humanity and about 60% of humanity living within 100 km of the productive coastlines,55 the conservation of freshwater and marine fishes is a priority.
One of the most important applications of a comprehensive, Pan-African barcoding study of fish will be more robust estimates of the phylogenetic diversity of different biogeographic regions. Phylogenetic diversity is a critical factor in conservation planning.56 A sufficient accumulation of barcodes will help conservation managers identify interim priority areas for conservation in the absence of good species data.
As PCR and sequencing technology advances—such as nanolitre-scale sequencing technology developments57—the evolution of a hand-held barcoder becomes more feasible. Fisheries managers, conservationists, legal authorities, ecologists, animal health professionals, agricultural practitioners, amateur naturalists and even the general angling public can collect molecular-based biodiversity data if such a 'barcoder' becomes widely available.58 This will markedly increase our capacity to identify species and understand the distribution of indigenous and alien fish species and their unique genetic lineages, especially in Africa where there are few trained taxonomists and sequencing facilities. It may also spark more interest in biodiversity, taxonomy and conservation issues if the general public have easier access to species information through rapid species identification.2,59
The FISH-BOL programme provides a platform for students trained in molecular systematics to collaborate with taxonomic experts to increase the transfer of knowledge and help train new taxonomists. Barcoding will not only provide a new avenue for researchers from different disciplines to collaborate more closely, but will also accelerate the process of species identification and discovery. These developments will be facilitated by a large online and publicly available reference database of DNA barcode information (Barcode of Life Data System, or BOLD26) for DNA sequence analyses and rapid species identification. BOLD may also help to link the user to taxonomic expertise and give the country of origin easier access to biodiversity information. The link between the barcode sequence, voucher specimen, image of the specimen and all the associated collecting and geographical information for future verification of species identity will add value to museum collections. Large-scale barcoding will also allow biomaterial banks to evaluate the effectiveness of their biomaterial curation techniques, as there is concern about the integrity of the nucleic acids of tissue samples stored in 98% ethanol or frozen at –20°C for more than five years.60 With so many fishes that remain to be investigated, FISH-BOL will not only increase the profile of museum collections, but will also expand existing scientific networks and collaborations throughout the continent.
A global network linked to the African FISH-BOL project
The FISH-BOL campaign will build on the success of sister projects that focus on other taxa, namely the All Birds Barcoding Initiative (ABBI) and the All Leps (Lepidoptera) Barcode of Life campaign. The FISH-BOL project will be more challenging than the All Birds project, not only because fish are far more diverse, but also because there is much less taxonomic information and expertise available. FISH-BOL, under the auspices of the Consortium for the Barcode of Life, has already assisted in driving a coherent approach within the taxonomic community. CBOL has built up an impressive network of collaborating organizations (for more information on CBOL and barcoding in general, see Savolainen et al.61 or visit www.barcoding.si.edu). One of these, the Census of Marine Life (CoML), has joined forces with CBOL and FISH-BOL to speed up the description of marine biodiversity.62 CoML will be a particularly valuable collaborator for FISH-BOL in Africa, because the programme has been productive in increasing exploration to assess diversity, distribution and abundance of marine life in African regions where little work had been done previously.
FISHBASE, the California Academy of Sciences' Catalog of Fishes and the Integrated Taxonomic Information System (ITIS) are other major depositories for updated taxonomic and biological information on fish species globally. FISH-BOL is currently using FISHBASE as its global taxonomic authority, but is also collaborating with Catalog of Fishes, ITIS and FISHBASE to incorporate their information into a resolved checklist for all fishes. FISH-BOL also supports the efforts of the International Commission on Zoological Nomenclature (ICZN) to establish a common registry of animal names. Some authors are already including barcodes of the holotype in their species descriptions,63,64 and this is expected to help reconcile and resolve ambiguities in the application of taxonomic names going forward.
The Global Biodiversity Information Facility (GBIF) has given scientists, conservationists and policy-makers access to geo-referenced, species databases across the world. Information relating to the South African National Fish Collection at the South African Institute for Aquatic Biodiversity (SAIAB), for example, has been linked through the South African Biodiversity Information Facility (SABIF) node to GBIF. FISH-BOL, through CBOL, is an associate participant in GBIF. Together, these initiatives aim to develop standards and protocols that will link voucher specimen data with barcode data. The U.S. National Center for Biotechnology Information (NCBI) is also working with GBIF and CBOL to make this happen and is the final depository (GenBank) for barcode sequences mainly administered through BOLD.26 This illustrates FISH-BOL and CBOL's commitment to work with established global programmes, networks and with the taxonomic community to increase the efficiency of describing biodiversity.
Several other global programmes and campaigns have attempted to inject new life into taxonomy over the years, and some have benefited African taxonomic research and inventories. For freshwater fish, the Cypriniformes Tree of Life (CToL) programme focused mainly on a phylogeny of cypriniform fishes, which includes the largest family of fishes, the Cyprinidae, with many species in Africa. In addition, the All Species Foundation, in partnership with the U.S. National Science Foundation (NSF), is attempting to describe all species on Earth in one human generation (25 years). They support the All Catfish Species Inventory programme that has been successful in increasing the number of species descriptions for fishes, in this case siluriform catfishes. These global initiatives, like many others, have been less successful in Africa, mainly because of a lack of taxonomic expertise in African countries to participate effectively. FISH-BOL will rely on its network of experts to identify fish and to train current African museum staff and field biologists in collecting data for barcoding.
The FISH-BOL programme will also explore other existing biodiversity networks in Africa to expand its operations. BioNET is attempting to promote taxonomy through the North African (NAFRINET), West African (WAFRINET), East African (EAFRINET) and the Southern African (SAFRINET) partnerships for taxonomy, the last becoming an official project of the Southern African Development Community (SADC). BioNET through EAFRINET has already held a regional meeting in Kenya and established the East African Barcoding Initiative (EABI). This group has recognized the need to barcode fish diversity and arranged a barcoding workshop to develop a proposal for a regional project on barcoding cyprinids. In South Africa, the South African Biosystematics Initiative (SABI) has stimulated research in taxonomy by providing dedicated government funding for taxonomic research. Nevertheless, there has been limited increase in the capacity of fish taxonomic research. The likely reasons for this deficiency include a lack of employment opportunities in museums or research institutions, and declining taxonomic expertise to train students. Only three fish taxonomic experts are formally employed in exclusively taxonomic positions in southern Africa, two at SAIAB in Grahamstown and one at the Iziko South African Museum in Cape Town. Although most new appointees in biodiversity-related posts have multidisciplinary skills (often including experience in molecular genetics and systematics), they lack the taxonomic expertise that is usually acquired only through years of research on a specific taxonomic group. This situation is a global problem, but especially exacerbates the Taxonomic Impediment in East, West and North Africa. It is therefore vital to increase collaboration by sharing the limited expertise and to encourage capacity building amongst African countries. If this is not addressed, Africa's fish diversity will not be managed in a sustainable manner.
SAIAB has signed a Memorandum of Understanding with the Canadian Centre for DNA Barcoding (CCDB) at the University of Guelph, and a Memorandum of Cooperation with CBOL to stimulate an African fish barcoding programme through the global FISH-BOL initiative. These developments act on the call by Klopper et al.65 to strengthen south–south and north–south collaboration to improve the capacity of taxonomic research in Africa. FISH-BOL will attempt to barcode all fish on Earth. This is indeed an ambitious task but it will be realistic to barcode all the available samples currently in collections within a relatively short period of time. Researchers from South Africa, Namibia, Zimbabwe, Zambia, Angola, Tanzania, Rwanda, Uganda, Kenya, Ethiopia, Cameroon, Benin, Ghana, Seychelles, Belgium and the United Kingdom have already expressed their interest in FISH-BOL through participation in African Regional Working Group activities.
Aims and challenges of the African FISH-BOL programme
The African FISH-BOL campaign will focus on all the fishes of the continent and the surrounding FAO marine regions 1, 34, 47 and 51 (see www.fishbol.org). The current FISH-BOL target for Africa of 8720 species that is based on FISHBASE is probably a major underestimate of the number of species that will eventually need to be investigated to make the campaign a success. The African Regional Working Group has been charged with editing the global checklist against African regional checklists. This review has already started with IUCN freshwater biodiversity projects in eastern and southern Africa, which suggest not only that many more fish species exist, but also that many of them remain undescribed.
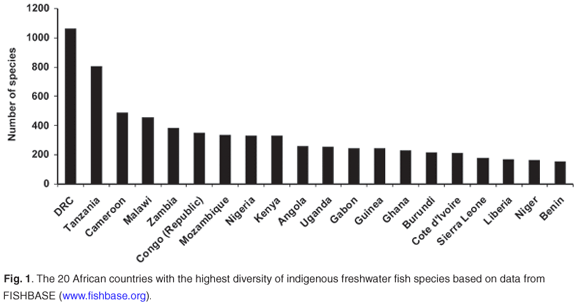
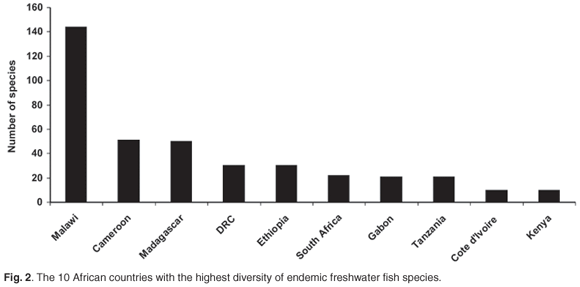
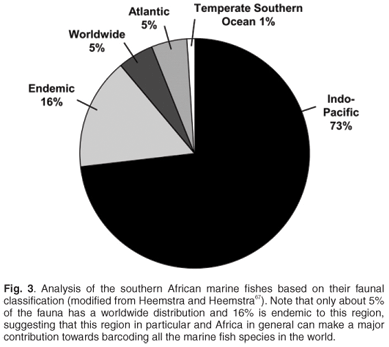
May66 suggested that the rate-limiting factor in describing biodiversity will remain the collection of new species in the field. This is especially true for Africa, where routine and geographically representative collection of species in the field remains problematic due to national conflicts, inaccessibility and a lack of capacity and logistical support. Of particular concern are those countries with the highest freshwater fish diversity and levels of endemism (Figs 1 and 2) based on data from FISHBASE (www.fishbase.org), without which the FISH-BOL programme will not achieve its goal of barcoding most of Africa's fish species. A successful programme in Africa will also make a major contribution to the barcoding of all the marine fish species in the world. For example, only about 5% of the southern African marine fish fauna has a worldwide distribution, whereas 16% is endemic to this region67 (Fig. 3).
The biggest challenge for the FISH-BOL project in Africa will be to collect, curate and accurately capture information in databases and deliver vouchers and samples of the species that are not already represented in natural history collections, to sequencing facilities. Field biologists interested in biodiversity, taxonomy and/or species identification can make an important contribution to this programme. Taxonomists will have a critical role in preparing and identifying vouchers for museum collections. BOLD has been designed specifically to link barcoding information to museum-registered specimens, so that taxonomic experts can go back to specimens to re-check identifications if necessary.26 This is crucial for new records or even conflicting results for known species. It will also be possible to submit sequences to GenBank (NCBI) directly from this database, where they will hold the status of 'Barcode' to signify their quality and direct linkage of DNA sequences to museum collection data. Barcoding therefore holds the promise to increase the usage of collections, and raise the profile of collections at a time when many museums suffer from budgetary constraints due to inappropriate governance and reduced government funding11,68,69 and will, we hope, also encourage better biomaterial banking practices.70,71 In the absence of other preferred facilities, SAIAB is currently the depository for tissues or extractions thereof (biomaterial bank), voucher specimens (main collection), images and associated locality and collection information (information system) that can be referred to once samples have been barcoded. All this information will be freely available to through SAIAB's web pages.
Conclusion
Many fish species remain to be discovered across the African continent and in its surrounding marine margins. We argue that barcoding holds a central role in the discovery and characterization of this diversity, and will also help democratize access to quality biodiversity information. Only time will tell how widely barcoding information will be used in fish species identification and aquatic biodiversity assessments in Africa. Preliminary results from the North American birds and Lepidoptera, North Atlantic shark and Australian marine fish projects have been positive. Although there are funding concerns, we envisage that FISH-BOL will provide previously inaccessible avenues of funding for taxonomic research in Africa. 'Barcoding' as a term immediately interests the public and therefore potential funders. The key challenge is for fish taxonomists, and the taxonomic community in general, to find ways in which this interest can be translated into greater support for taxonomic research, which will underpin the more efficient description of biodiversity.
The Consortium for the Barcode of Life (CBOL) is thanked for support to attend meetings where many of the aspects of barcoding referred to in this paper were discussed. Denis Tweddle (SAIAB) and three anonymous reviewers are thanked for comments on the manuscript. Our research is supported through the Ontario Genomics Institute within the Canadian Barcode of Life Network by Genome Canada, NSERC and other sponsors listed at www.bolnet.ca
1. Hoagland K.E. (1996). The Taxonomic Impediment and the Convention on Biodiversity. ASC News 24, 61–62, 66–67. [ Links ]
2. Costa F.O. and Carvalho G.R. (2007). The Barcode of Life Initiative: synopsis and prospective societal impacts of DNA barcoding of fish. Genomics Soc. Policy 3, 29–40. [ Links ]
3. May R.M. and Beverton R.J.H. (1990). How many species? Philos. Trans.: Biol. Sci. 330, 293–304. [ Links ]
4. Stork N.E. (1988). Insect diversity: facts, fiction and speculation. Biol. J. Linn. Soc. 35, 321–337. [ Links ]
5. Southwood T.R.E. (1978). The components of diversity. In Diversity of Insect Faunas, eds L.A. Mound and N. Waloff, pp. 19–40. Blackwell, Oxford. [ Links ]
6. Mace G.M. (2004). The role of taxonomy in species conservation. Phil. Trans. R. Soc. Lond. B 359, 711–719. [ Links ]
7. Thomas C.D., Cameron A., Green R.E., Bakkenes M., Beaumont L.J. et al. (2004). Extinction risk from climate change. Nature 427, 145–148. [ Links ]
8. Hughes J.B., Daily G.C. and Ehrlich P.R. (1997). Population diversity: its extent and extinction. Science 278, 689–692. [ Links ]
9. Agosti D. (2006). Biodiversity data are out of local taxonomists' reach. Nature 439, 392–392. [ Links ]
10. Balakrishnan R. (2005). Species concepts, species boundaries and species identification: a view from the Tropics. Syst. Biol. 54, 689–693. [ Links ]
11. Herbert D.G. (2001). Museum natural science and the NRF: crisis times for practitioners of fundamental biodiversity science. S. Afr. J. Sci. 97, 168–172. [ Links ]
12. Wynberg R. (2002). A decade of biodiversity conservation and use in South Africa: tracking progress from the Rio Earth Summit to the Johannesburg World Summit on Sustainable Development. S. Afr. J. Sci. 98, 233–243. [ Links ]
13. Wheeler Q.D., Raven P.H. and Wilson E.O. (2004). Taxonomy: impediment or expedient? Science 303, 285. [ Links ]
14. Polaszek A. (2005). A universal register for animal names. Nature 437, 477. [ Links ]
15. Hebert P.D.N., Cywinska A., Ball S.L. and De Waard J.R. (2003). Biological identifications through DNA barcodes. Proc. R. Soc. Lond. B 270, 313 – 321. [ Links ]
16. Schander C. and Willassen E. (2005). What can biological barcoding do for marine biology? Mar. Biol. Res. 1, 79 – 83. [ Links ]
17. Schindel D.E. and Miller S.E. (2005). DNA barcoding a useful tool for taxonomists. Nature 435, 17–17. [ Links ]
18. Miller S.E. (2007). DNA barcoding and the renaissance of taxonomy. Proc. Natl Acad. Sci. USA 104, 4775–4776. [ Links ]
19. de Carvalho M.R., Bockmann F.A., Amorim D.S., de Vivo M., de Toledo-Piza M. et al. (2005). Revisiting the taxonomic impediment. Science 307, 353. [ Links ]
20. Ebach M.C. and Holdrege C. (2005). DNA barcoding is no substitute for taxonomy. Nature 434, 697–697. [ Links ]
21. Blaxter M.L. (2004). The promise of a DNA taxonomy. Phil. Trans. R. Soc. B 359, 669–679. [ Links ]
22. DeSalle R. (2007). Phenetic and DNA taxonomy; a comment on Waugh. Bioessays 29, 1289–1290. [ Links ]
23. Gómez A., Wright P.J., Lunt D.H., Cancino J.M., Carvalho G.R. et al. (2007). Mating trials validate the use of DNA barcoding to reveal cryptic speciation of a marine bryozoan taxon. Proc. R. Soc. Lond. B 274, 199–207. [ Links ]
24. Moritz C., Dowling T.E. and Brown W.M. (1987). Evolution of animal mitochondrial DNA: relevance for population biology and systematics. Annu. Rev. Ecol. Syst. 18, 269–292. [ Links ]
25. Collins F.S., Morgan M. and Patrinos A. (2003). The Human Genome Project: lessons from large-scale biology. Science 300, 286–290. [ Links ]
26. Ratnasingham S. and Hebert P.D.N. (2007). BOLD: the Barcode of Life data system (http://www.barcodinglife.org). Mol. Ecol. Notes 7, 355–364. [ Links ]
27. Hebert P. and Gregory T. (2005). The promise of DNA barcoding for taxonomy. Syst. Biol. 54, 852–859. [ Links ]
28. Hajibabaei M., Singer G.A.C., Hebert P.D.N. and Hickey D.A. (2007). DNA barcoding: how it complements taxonomy, molecular phylogenetics and population genetics. Trends Genet. 23, 167–172. [ Links ]
29. Waters J.M. and Cambray J.A. (1997). Intraspecific phylogeography of the Cape galaxias from South Africa: evidence from mitochondrial DNA sequences. J. Fish Biol. 50, 1329–1338. [ Links ]
30. Wishart M., Hughes J., Stewart B. and Impson, D. 2006. Extreme levels of intra-specific divergence among Cape Peninsula populations of the Cape galaxias, Galaxias zebratus Castelnau 1861, reveals a possible species complex. Afr. J. Aquat. Sci. 31, 99–106. [ Links ]
31. Swartz E.R., Skelton P.H. and Bloomer, P. (2007). Sea-level changes, river capture and the evolution of populations of the Eastern Cape and fiery redfins (Pseudobarbus afer and Pseudobarbus phlegethon, Cyprinidae) across multiple river systems in South Africa. J. Biogeog. 34, 2086–2099. [ Links ]
32. Nielsen R. and Matz M. (2006). Statistical approaches for DNA Barcoding. Syst. Biol. 55, 162–169. [ Links ]
33. DeSalle R., Egan M. and Siddall M. (2005). The unholy trinity: taxonomy, species delimitation and DNA barcoding. Phil. Trans. R. Soc. B 360, 1905–1916. [ Links ]
34. Rubinoff D. (2006). Utility of mitochondrial DNA barcodes in species conservation. Conserv. Biol. 20, 1026–1033. [ Links ]
35. Hubert N., Hanner R., Holm E., Mandrak N.E., Taylor E., Burridge M., Watkinson D., Dumont P., Curry A., Bentzen P., Zhang J., April J. and Bernatchez L. (2008). Identifying Canadian freshwater fishes through DNA barcodes. Plos ONE 3, e2490–2490. [ Links ]
36. Ward R.D., Zemlak T.S., Innes B.H., Last P.R. and Hebert P.D.N. (2005). DNA barcoding Australia's fish species. Phil. Trans. R. Soc. B 360, 1847–1857. [ Links ]
37. Venkatesh B., Dandona N. and Brenner S. (2006). Fugu genome does not contain mitochondrial pseudogenes. Genomics 87, 307–310. [ Links ]
38. Waters J.M. and Wallis G.P. (2001). Cladogenesis and loss of the marine life-history phase in freshwater galaxiid fishes (Osmeriformes: Galaxiidae). Evolution 55, 587–597. [ Links ]
39. Kerr K.C.R., Stoeckle M.Y., Dove C.J., Weigt L.A., Francis C.M. et al. (2007). Comprehensive DNA barcode coverage of North American birds. Mol. Ecol. Notes 7, 535–543. [ Links ]
40. Greig T.W., Moore M.K., Woodley C.M. and Quattro J.M. (2005). Mitochondrial gene sequences useful for species identification of western North Atlantic Ocean sharks. Fish. Bull. 103, 516–523. [ Links ]
41. Hebert P.D.N., Stoeckle M.Y., Zemlak T.S. and Francis C.M. (2004). Identification of birds through DNA barcodes. PLoS Biology 2, e312. [ Links ]
42. Hebert P.D.N., Penton E.H., Burns J.M., Janzen D.H. and Hallwachs W. (2004). Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc. Natl Acad. Sci. USA 101, 14812–14817. [ Links ]
43. Hogg I.D. and Hebert P.D.N. (2004). Biological identification of springtails (Collembola: Hexapoda) from the Canadian Arctic, using mitochondrial DNA barcodes. Can. J. Zool. 82, 749–754. [ Links ]
44. Barber P. and Boyce S.L. (2006). Estimating diversity of Indo-Pacific coral reef stomatopods through DNA barcoding of stomatopod larvae. Proc. R. Soc. Lond. B 273, 2053–2061. [ Links ]
45. Anon. (2007). Fisheries and Aquaculture Department: the state of world fisheries and aquaculture 2006. FAO, Rome. [ Links ]
46. Cowx I.G. (2002). Analysis of threat to freshwater fish conservation: past and present challenges. In Conservation of Freshwater Fishes: Options for the future, eds M.J. Collares-Pereira, I.G. Cowx and M.M. Coelho, pp. 201–220. Fishing News Books, London. [ Links ]
47. Mace G., Masundire H., Baillie J., Ricketts T., Brooks T. et al. (2005). Biodiversity. Millennium Ecosystem Assessment, 2005 Current state and trends: findings of the Condition and Trends Working Group Ecosystems and human well-being, vol. 1. Island Press, Washington, D.C. [ Links ]
48. Lleonart J., Taconet M. and Lamboeuf M. (2006). Integrating information on marine species identification for fishery purposes. Mar. Ecol. Prog. Ser. 316, 231–238. [ Links ]
49. Leis J.M., Trnski T. and Beckley L.E. (2002). Larval development of Pagellus natalensis and what larval morphology indicates about relationships in the perciform fish family Sparidae (Teleostei). Mar. Freshwat. Res. 53, 367–376. [ Links ]
50. Pegg G.G., Sinclair B., Briskey L. and Aspden W.J. (2006). MtDNA barcode identification of fish larvae in the southern Great Barrier Reef, Australia. Sci. Mar. 70S2, 7–12. [ Links ]
51. Webb K.E., Barnes D.K.A., Clark M.S. and Bowden D.A. (2006). DNA barcoding: a molecular tool to identify Antarctic marine larvae. Deep-Sea Res. II 53, 1053– 1060. [ Links ]
52. Smith P.J., McVeagh S.M., Allain V. and Sanchez C. (2005). DNA identification of gut contents of large pelagic fishes. J. Fish Biol. 67, 1178–1183. [ Links ]
53. Siddall M.E. and Budinoff R.B. (2005). DNA-barcoding evidence for widespread introductions of a leech from the South American Helobdella triserialis complex. Conserv. Genet. 6, 467–472. [ Links ]
54. Swartz E.R. (2005). Phylogeography, phylogenetics and evolution of the redfins (Teleostei, Cyprinidae, Pseudobarbus) in southern Africa. PhD thesis. University of Pretoria, South Africa. [ Links ]
55. Vitousek P.M., Mooney H.A., Lubchenco J. and Melillo J.M. (1997). Human domination of Earth's ecosystems. Science 277, 494–499. [ Links ]
56. Faith D.P. (1992). Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10. [ Links ]
57. Blazej R.G., Kumaresan P. and Mathies R.A. (2006). Microfabricated bioprocessor for integrated nanoliter-scale Sanger DNA sequencing. Proc. Natl Acad. Sci. USA 103, 7240–7245. [ Links ]
58. Hollingsworth P.M. (2007). DNA barcoding: potential users. Genomics Soc. Policy 3, 44–47. [ Links ]
59. Holm P. (2007). The book of life goes online. Genomics Soc. Policy 3, 48–51. [ Links ]
60. Chakraborty A., Sakai M. and Iwatsuki Y. (2006). Museum fish specimens and molecular taxonomy: a comparative study on DNA extraction protocols and preservation techniques. J. Appl. Ichthyol. 22, 160–166. [ Links ]
61. Savolainen V., Cowan R., Vogler A., Roderick G. and Lane R. (2005). Towards writing the encyclopaedia of life: an introduction to DNA barcoding. Phil. Trans. R. Soc. B 360, 1805–1811. [ Links ]
62. Mason B. (2003). Marine survey sees net gain in number of fish species. Nature 425, 889–889. [ Links ]
63. Victor B.C. (2007). Coryphopterus kuna, a new goby (Perciformes: Gobiidae: Gobiinae) from the western Caribbean, with the identification of the late larval stage and an estimate of the pelagic larval duration. Zootaxa 1526, 51–61. [ Links ]
64. Pyle R.L., Earle J.L. and Greene B.D. (2008). Five new species of the damselfish genus Chromis (Perciformes: Labroidei: Pomacentridae) from deep coral reefs in the tropical western Pacific. Zootaxa 1671, 3–31. [ Links ]
65. Klopper R.R., Smith G.F. and Chikuni A.C. (2002). The Global Taxonomy Initiative in Africa. Taxon 51, 159–165. [ Links ]
66. May R. (2004). Tomorrow's taxonomy: collecting new species in the field will remain the rate-limiting step. Phil. Trans. R. Soc. B 359, 733–734. [ Links ]
67. Heemstra P. and Heemstra E. (2004). Coastal Fishes of Southern Africa. National Inquiry Service Centre and South African Institute for Aquatic Biodiversity, Grahamstown, South Africa. [ Links ]
68. Cotterill F.P.D. (2002). The future of natural science collections into the 21st century. Conferencia de clausura. Actas del I simposio sobre el patrimonio natural en las colecciones públicas en España. Departamento de cultura, diputación foral de Alava, Vitoria, pp. 237–282. [ Links ]
69. Suarez A.V. and Tsutsui N.D. (2004). The value of museum collections for research and society. BioScience 54, 66–74. [ Links ]
70. Prendini L., Hanner R. and DeSalle R. (2002). Obtaining, storing and archiving specimens and tissue samples for use in molecular studies. In Techniques in Molecular Systematics and Evolution, eds R. DeSalle, G. Giribet and W. Wheeler. Birkhäuser Verlag, Basel. [ Links ]
71. Walters C. and Hanner R. (2006). Platforms for DNA banking. In DNA Banks – Providing novel options for genebanks? Topical Reviews in Agricultural Biodiversity, eds M.C. de Vicente and M.S. Andersson, pp. 25–35. International Plant Genetic Resources Institute, Rome. [ Links ]
Received 10 April. Accepted 5 August 2008.
* Author for correspondence. E-mail: e.swartz@ru.ac.za














