Servicios Personalizados
Articulo
Indicadores
Links relacionados
-
 Citado por Google
Citado por Google -
 Similares en Google
Similares en Google
Compartir
Onderstepoort Journal of Veterinary Research
versión On-line ISSN 2219-0635
versión impresa ISSN 0030-2465
Onderstepoort j. vet. res. vol.80 no.1 Pretoria ene. 2013
ORIGINAL RESEARCH
The incursion, persistence and spread of peste des petits ruminants in Tanzania: Epidemiological patterns and predictions
Fredrick M. KivariaI; Olivier KwiatekII; Angolwisye M. KapagaIII; Emmanuel S. SwaiIV; Geneviève LibeauII; Winford MoshyIV; Albano O. MbyuziV; Joshua GladsonIII
INational Epidemiology Section, Ministry of Livestock and Fisheries Development, Tanzania
IICIRAD, Campus International de Baillarguet, France
IIICentral Veterinary Laboratory, Tanzania
IVVeterinary Investigation Centre, Arusha, Tanzania
VVeterinary Investigation Centre, Mtwara, Tanzania
ABSTRACT
Peste des petits ruminants virus, which causes a severe disease in sheep and goats, has only recently been officially declared to be present in Tanzania. An epidemiological study was carried out between September 2008 and October 2010 to investigate the incursion, persistence and spread of the virus in Tanzania. The investigation involved serosurveillance, outbreak investigation and computation of epidemiological indices such as the effective reproductive number, persistence and the threshold level for vaccination. Field and molecular epidemiological techniques were applied to isolate, characterise and trace the origin of the virus in Tanzania. A total of 2182 serum samples from goats and 1296 from sheep from 79 villages across 12 districts were investigated. Village-level prevalence of infection was variable (0.00% - 88.00%) and was higher in pastoral than in agro-pastoral villages. The overall antibody response to the virus was 22.10% (CI95% = 20.72% - 23.48%). About 68.00% and 73.00% of seropositive goats and sheep, respectively, did not show clinical signs. The proportion of seropositive animals differed significantly (p < 0.001) between age groups, sex and farming practices. Real-time polymerase chain reaction results showed that the isolated strains belong to lineage III, whose origin is in East Africa and the Middle East. This indicates that one of the northern neighbouring countries is most likely the source of infection. The computed overall effective reproductive number, the threshold level of vaccination necessary to eradicate the disease and persistence were 4.75% and 98.00%, respectively. These estimates indicate that achieving elimination of the peste des petits ruminants virus from pastoral flocks will require significant effort and development of highly effective intervention tools.
Introduction
In Tanzania, sheep and goats are important livestock species as they are able to survive unfavourable climatic conditions of arid and semiarid environments; hence, they are species of choice of pastoralists. These animals are widely distributed across different agro-ecological zones and production systems and form an integral part of the livelihood of rural households. However, the contribution of sheep and goats to food security in Tanzania has recently been threatened by outbreaks of peste des petits ruminants (PPR).
The first outbreak of PPR in sheep and goats in Tanzania was confirmed in December 2008. Since then, sporadic outbreaks have been reported in many parts of the country. The three fundamental steps of exotic pathogen emergence are incursion, persistence and spread in a geographical context. In this article, incursion refers to the first confirmed arrival of PPR virus (PPRV) in the country, persistence refers to the obstinate survival of PPRV in sheep and goat populations in spite of vaccination and other biosecurity measures undertaken to eliminate the virus, and spread refers to the movement of PPRV between different geographical localities. We use data on the first occurrence and the subsequent events of PPR incursion and spread in Tanzania to illustrate the epidemiological factors that drive these sequential events amongst the pastoral sheep and goat population. The epidemiological data, clinical and pathological features of this outbreak and recovery and identification of PPRV are also reported.
Materials and methods
The historical perspectives of peste des petits ruminants incursion into Tanzania
Incursion of PPR into Tanzania was suspected after a confirmed PPR outbreak in Kenya in August 2006. The possible presence of the disease in Tanzania was reported at the Tanzania Veterinary Association annual scientific conference by the Ngorongoro District Veterinary Officer in December 2007. In March 2008, a high mortality rate was observed amongst sheep and goats in the Ngorongoro district and a technical team was dispatched to investigate the cause of the increased deaths. A total of 112 sheep and goats were subjected to clinical, pathological and serological investigations. Both clinical and pathological investigations yielded inconclusive results, whilst serological investigation yielded negative results for PPR.
However, reports of increased deaths in sheep and goats continued in the Ngorongoro and Mara districts. In June 2008, a second team was dispatched to the Ngorongoro and Mara districts, where, respectively, 404 and 84 serum samples were collected. Of the 404 samples collected in the Ngorongoro district, 129 (31.93%) tested positive for PPR during serological screening. None of the samples from the Mara district tested positive. Based on these rapid epidemiological assessment (REA) results, the director of veterinary services designed a cross-sectional study that was aimed at gauging the magnitude and extent of PPR spread in the country.
Field investigation of the incursion of peste des petits ruminants
A cross-sectional study was carried out in 12 districts across the regions of Arusha, Kilimanjaro, Manyara and Tanga in the north of the country between September 2008 and July 2009. The aim of the study was to determine the magnitude of PPR in the study area and the extent of the southward spread. Approximately 4 362 838 goats and 2 079 151 sheep were estimated to be at risk in the study, representing about 57.75% and 32.08% of the national goat and sheep populations, respectively.
All sheep and goat flocks in the sampling area formed the population from which study flocks were randomly selected. Multistage sampling (Dohoo, Martin & Stryhn 2003), with four hierarchical stages, was used as sampling strategy. The first level of selection was the region; the Arusha, Kilimanjaro, Manyara and Tanga regions were purposely selected. Within each of the selected regions, specific districts (Karatu, Longido, Mbulu, Monduli, Ngorongoro, Siha, Simanjiro, Korogwe, Lushoto, Mkinga, Muheza and Tanga-municipal) were also purposely selected. Villages were randomly selected. Within each of the selected villages, 40 animals (either sheep or goats) were randomly selected. The study regions and districts were purposely selected based on perceived risk due to either their bordering Kenya or the presence of farming activities characterised by extensive livestock movement.
Owing to the characteristics of the pastoral and agro-pastoral livestock production systems, it is reasonable to consider all the sheep and goats in a village as one flock, with very little within-flock variation. A village was therefore considered to constitute a flock. The sample size was computed as suggested by Dohoo et al. (2003):

where z = 1.96, p = 0.32 (the prior prevalence from the REA study described earlier) and L = 0.05 (the desired level of precision or accuracy). The required sample size was accordingly calculated as n = 334.37.
However, PPR is a highly contagious disease. To obtain a similar accuracy to that with simple random sampling, the sample size had to be recalculated. The aim was to sample at least 40 animals from each village, and from the REA study the average rate of PPR homogeneity (p) was estimated to be 0.156. The sample size was subsequently recalculated as:

where n = 334.37, p = 0.156 and m = 40, representing the average number of animals to be sampled from each village (Dohoo et al. 2003). Therefore, the new sample size was equal to 2360.816 animals and 59 villages were to be included in this study. The design effect (D) was calculated as 7.084 according to:

The standard error (S) was calculated as 0.026 according to:

indicating that with this design the certainty associated with the true prevalence of sheep and goats with antibodies to PPR will be p ± 1.960 x 0.026 of the estimate. The prevalence was determined by dividing the total number of positive samples by the total number of samples (Dohoo et al. 2003). The sampling frame from which the 59 study villages were randomly selected was obtained from the district agriculture office.
Clinical and pathological investigation
In all villages that were visited, discussions were held with livestock owners and animals were inspected for evidence of classic PPR symptoms (severe mucopurulent nasal and ocular discharges, necrotic stomititis and respiratory distress). Dead animals were necropsied for evidence of PPR lesions and organ collection. Pieces of the lung, spleen, kidney, liver and mesenteric lymph nodes from one dead goat were collected for laboratory confirmation. In addition, two goats from the Ngorongoro district, which presented with clinical signs of PPR, were purchased and sacrificed for collection of microbiological samples and pathological lesions.
Laboratory techniques
Competitive enzyme-linked immunosorbent assay for peste des petits ruminants
A nucleotide monoclonal antibody-based competitive enzyme-linked immunosorbent assay (cELISA) was used for the detection of antibodies to PPRV (FAO/IAEA 1993; OIE 2000). This test was developed using a virus-neutralising monoclonal antibody directed against an epitope of the haemagglutinin protein specific to PPRV. All serum samples tested in the present study were processed in duplicate according to the standard protocol for the detection of rinderpest antibodies (FAO/IAEA 1993; OIE 2000). The results of the cELISA were interpreted using software from the Economic Development Institute developed by the Food and Agriculture Organization/International Atomic Energy Agency for the detection of rinderpest antibody (FAO/IAEA 1993). Results were interpreted as previously described by Libeau et al. (1995). Samples that showed colour inhibition (inhibition of the enzymatic colour reaction) > 50% when compared to wells containing the control (no serum), were considered positive. A confidence interval at the 95% level (CI95%) for the prevalence of antibodies to PPRV in the population studied was calculated using standard statistical methods (Snedecor & Cochran 1989).
Real-time polymerase chain reaction analysis and lineage definition
A total of six sheep samples from the Ngorongoro district in the Arusha region were analysed for viral identification and lineage definition. One sample consisted of ovine tissues (liver, spleen, heart and mediastinal lymph nodes) and five of heparinised whole blood. Total RNA was extracted and cDNA was amplified as previously described (Kwiatek et al. 2007). The polymerase chain reaction (PCR) used the pan-morbillivirus primers located in the middle of the gene (Nad1: CCAAGGATTGCAGAAATGATC, Nad2: GCTCCTGCACTAAACTTGTTC), which allow the amplification of the N-gene of all morbilliviruses to give a product of 230 bases. The PCR conditions were as follows: an activation of 10 min at 95 °C, 35 cycles of amplification (30 s at 94 °C, 30 s at 55 °C and 30 s at 72 °C) and a final extension step at 72 °C for 7 min. The PCR was performed in a 2720 thermal cycler (Applied Biosystems). The resulting PCR products were analysed by electrophoresis on an ethidium bromide-stained 1.5% agar gel.
The nucleic acid sequences obtained from PCR products amplified with the Nad1-Nad2 primer were aligned with sequences from PPRV strains that were maintained in our database. Phylogenetic analysis was performed on the 224 nucleotides located on the nucleoprotein gene of the virus. The known PPRV sequences were derived from strains isolated over a period of 20 years. They consisted of 'historical' strains, such as the Nigeria 75/1, from which the PPR vaccine was developed, and also the Dorcas strain isolated from a Dorcas gazelle in a zoological collection in the United Arab Emirates, (Furley, Taylor & Obi 1987). Strains isolated between 1988 and 2000 originated from Cameroon, India, Senegal, Turkey, Ghana, Guinea Bissau, Oman and the Ivory Coast. The alignment of nucleotide sequences was created by means of the Clustal W program (Vector NTI, Informax Inc., Rockville). Phylogenetic analysis was carried out as described by Kwiatek et al. (2007). Bootstrap confidence values were calculated on 1000 replicates. The RBOK vaccine strain of rinderpest virus was considered as an out-group.
Investigation on the persistence of peste de petits ruminants
A fundamental parameter determining the success (persistence and spread) of any infectious organism in a host population is the basic reproductive number, R0, which measures the potential for spread of infection. It is defined as the average number of new infections caused when an infection enters an entirely susceptible population (Anderson & May 1991). Two things are worth noting from this definition. First, it is an average value that can change from population to population or over time, depending on the patterns of contact or biological influences at play. Second, the subscripted identifier (zero) refers to the time at the start; that is, when invasion occurs. As the infection spreads, the population is no longer entirely susceptible. Contacts are therefore wasted on infected or immune individuals and the reproductive number is reduced to an effective reproductive number, Rt. The effective reproductive number has a threshold value of 1 to ensure persistence and can be used to estimate the level of immunisation or managerial change required to control an epidemic.
In this study, Rt was estimated from the average age at seroconversion by the method of Dietz (1993). The deterministic formula used was:

where Ls is the expected average farm gate lifespan of sheep and goats and As is the average age at seroconversion. An estimate of Rt was also determined as described by Heffernan and Wahl (2005), using reported morbidity and case fatality risk associated with PPR according to the formula:

where rt is the reported PPR morbidity and at is the average PPR case fatality risk.
The following assumptions were applied:
- From an epidemiological perspective, the small ruminant (sheep and goat) population is assumed to be a single, homogeneous, free-mixing host population (referred to as 'the flock') and contacts between individuals were random.
- The kidding and lambing rates were high enough to sustain an input of susceptible hosts into the population, thereby maintaining a constant chain of infection in the population.
- Screening of 250 sheep and 600 goat samples, randomly selected from the 2006-2008 sera that were collected from the study area during rinderpest elimination campaigns, yielded negative results for PPRV antibodies. It was therefore logical to assume that the study animals were seronegative during the 12 months before sampling, and therefore fully susceptible to the invading PPRV infection.
- The duration of the infectious period is not related to age.
- The deterministic model did not consider the probability of re-invasion of the recovered flock.
The probability of persistence or spread, ppers, over the next generation was computed as ppers = 1 - (1/Rt)n, where n is the number of infectious foci (the number of villages that reported PPR cases) and Rt equals R0 when there is no immunity or previous infection within the population. The threshold level of vaccination (Vt) necessary to eradicate the disease was calculated according to Vt = 1 - Rt-1, as described in Anderson and May (1991).
Investigation on the spread of peste des petits ruminants in Tanzania
In December 2009, an outbreak of a disease characterised by the sudden onset of yellowish diarrhoea, mucoid nasal and ocular discharges with matting of the eyelids and erosive lesions of the gums followed by death was reported from Likuna, a village in the Newala district of the Mtwara region in southern Tanzania. Participatory epidemiological investigations showed that a livestock trader, and a resident of the village, had brought 70 goats from the Pugu market in the Dar es Salaam region to be sold during the approaching seasonal festivals (Christmas and New Year). The trade goats started to die after three days, without noticeable clinical signs. To avoid losses, the trader sold the whole consignment cheaply to the nearby villages of Makote, Mnazi mmoja, Lidumbe and Mambika. The disease affected the resident stock with high morbidity and mortality rates, estimated at 90% and 73% respectively. Clinical examination was carried out in 30 randomly selected animals and necropsies were performed on 10 dead animals. Ocular and nasal swabs and sera were collected for laboratory confirmation from 217 randomly selected animals. The epidemic curve was plotted to see the pattern of the outbreak, using the daily records made by the Newala district veterinary officer.
Competitive ELISA showed that antibodies against PPRV were present in 59% of the 217 serum samples and PPRV protein was detected in 34% of the screened ocular and nasal swabs. By March 2010 the disease had spread to the nearby districts of Tandahimba and Masasi, where, respectively, 84% of the 180 and 3% of the 125 samples screened by "cELISA yielded positive results for antibodies against PPRV.
In April 2010 about 400 small ruminants were reported to have died because of an unknown disease in two divisions of Malinyi and Mtimbira in the Ulanga district of the Morogoro region. Epidemiological investigations suggested that the disease was PPR, which invaded the region through returning migratory pastoralists who had moved northwards from the Mtwara and Lindi regions. Serological investigations revealed the presence of antibodies against PPRV in 152 (76%) of the 200 serum samples tested by means of "cELISA. In 2011, antibodies to PPRV were demonstrated in trade cattle sharing common grazing lands with 225 small ruminants in the Mukuranga district of the Coast region. Although not susceptible to the disease, cattle are able to seroconvert in case of a high prevalence of PPR in small ruminants (Abubakar et al. 2011).
In October 2010 the Director of Veterinary Services conducted risk-based serosurveillance in all districts that border Mozambique, Malawi and Zambia. Samples were collected from animals at randomly selected high-risk areas, such as border crossings, auctions, slaughter slabs and grazing and watering points, and other areas where animals from a number of villages congregate. A total of 720 serum samples were collected under the Vaccine for Control of Neglected Animal Diseases in Africa (VACNADA) programme (funded by the European Union) and screened for antibodies against PPRV at the PANVAC laboratories in Addis Ababa, Ethiopia. Of the 720 samples, 266 (36.9%) tested positive for antibodies against PPRV. Clinical disease was detected in the Mahuta ward of the Tandahimba district, Mtwara and according to RT-PCR, PPRV protein was detected in 33.0% of the tissue samples taken from 28 goats here.
The chronology and probable movement of PPR in Tanzania is depicted in Figure 1.
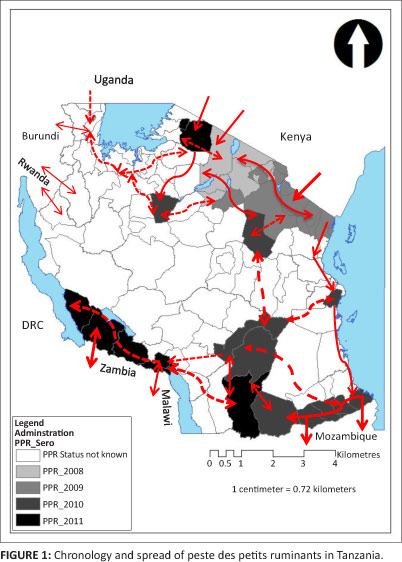
Results
Field investigations on the incursion of peste des petits ruminants
Clinical signs: Macro- and microscopical lesions
Clinical signs of PPR were observed in 90 goats and 60 sheep, independent of age or sex, in the Ngorongoro, Monduli, Longido and Siha districts. The clinical signs included diarrhoea, especially in young animals, elevated body temperatures (> 40 °C), serous rhinorrhoea and serous mucopurulent nasal discharges. Peri-nasal areas were often found encrusted with mucopurulent discharges. However, such clinical signs were not observed in the flocks in the other study districts.
Two necropsied goats showed inflammation of the apical lung lobes, with multiple abscesses evident in one of the goats. The mesenteric lymph nodes in both carcasses were swollen and the small intestines were hyperaemic. Microscopical lesions in the lungs were associated with bronchopneumonia and focal necrotic areas surrounded by inflammatory zones were also found.
Cross-sectional study results
A total of 3478 serum samples (2182 from goats; 1296 from sheep) were obtained from 79 villages1 between September 2008 and July 2009. The distribution of the study variables was such that 72.70% of the study animals were female, whilst 27.30% were male. Of the 3478 animals, 77.80% were classified as adults and 22.20% were classified as kids or lambs. The majority of the study animals (2162/3478; 62.16%) were from the agro-pastoral farming system, whilst 37.84% (1316/3478) came from the pastoral farming system.
Serological results and risk indicators for positive serological status against peste des petits ruminants
Of the tested animals, 22.10% (CI95%: 20.72% - 23.48%) were classified as seropositive for PPRV antibodies. Up to 88.00% of the study animals tested positive for PPRV, with a greater proportion of positive samples found in the Arusha, Manyara and Kilimanjaro regions (in descending order) than in the Tanga region. The percentage of PPRV-positive samples are categorised according to region, district, farming practice and animal species in Table 1. It was estimated that 68.00% of the 483 goats and 73.00% of the 286 sheep that tested positive for antibodies against PPRV had not displayed clinical signs. The proportion of seropositive animals differed significantly according to age, sex and farming practice, but there was no significant association between animal species and seropositivity. Results of the pairwise comparison are summarised in Table 2. Despite an overall frequency of 22.10% (CI95%: 20.72% - 23.48%), the seroprevalence of PPR appeared to be variable across the study districts; pastoral districts appeared generally to have suffered more from infection (Z = 15.38; p < 0.0001) than the agro-pastoral districts, and a seroprevalence of 0.00% was reported in two districts (Table 2).
Characterisation of the strain and determination of its geographical lineage
Two samples (one tissue sample and one from blood) were found positive for PPRV by means of RT-PCR. To obtain epidemiological information and analyse the genetic relation of the 2008 Tanzania strain (from the Ngorongoro district) with strains of different geographical origins, the nucleic acid sequences obtained from PCR products amplified with the Nad1-Nad2 primer were aligned with sequences from PPRV strains retrieved from GenBank or maintained in our database. Phylogenetic analysis was performed on the 224 nucleotides located on the nucleoprotein gene of the virus. The program MRBAYES (version 3.1.2) was used to perform Bayesian analysis. Tree manipulations were compiled using the FigTree program (version 1.3.1) (http://tree.bio.ed.ac.uk/software/figtree/). The RBOK wild strain of RPV and a measles virus strain were considered as out-groups. For support, phylogenetic analysis was also carried out as described in Kwiatek et al. (2007).
As expected, the 2008 Tanzania strain of PPRV fell into lineage III, as is the case for East African strains, and it was closely related to the historical strains Ibri and Dorcas isolated since 1987 in the Sultanate of Oman (Furley et al. 1987). The lineage is distinct from the one circulating in Asia from south India to Turkey (Özkul et al. 2002).
Figure 2 shows the Bayesian inference of PPRV strains. The phylogenetic reconstruction was made using MRBAYES software as mentioned earlier (Huelsenbeck & Ronquist 2001). The sequence evolution model proposed by Jobb (2011) was HKY. Consequently, the number of substitution types was set at six and the Bayesian reconstruction was run for 6 000 000 iterations. Results were analysed when the standard deviation between the Monte Carlo Markov chains fell below 0.01. Trees were sampled every 1000 generations; thus the consensus tree was built up with a burn-in phase of 25% and posterior probabilities were displayed as node labels. The tree was rooted by two out-groups, namely measles and rinderpest morbilliviruses collected from Genbank (FJ416068 and X98291 respectively), and the four PPRV lineages are highlighted in black brackets. Tanzania_2009 analysed in this study is underlined in lineage III.
Persistence and spread of peste des petits ruminants
Estimates of Rt derived through deterministic and morbidity or mortality techniques for each of the investigated districts, as well as those based on pooled data for the districts, were similar. These estimates, together with their respective vaccination threshold (Vt) estimates, are summarised in Table 3a and Table 3b. The overall persistence was estimated at 0.98.
As shown in Figure 3 the epidemic curve resembles a typical propagated outbreak, with peaks of primary, secondary and tertiary generations of cases spaced by an incubation period. The most probable period of exposure of the susceptible animals to PPRV was calculated as 3-5 days before the index case was reported.
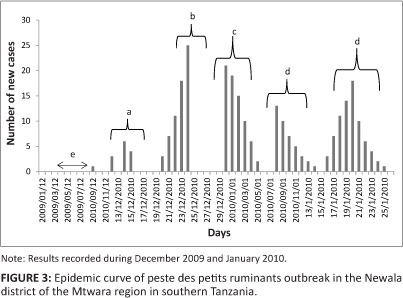
It was of epidemiological interest to observe that clinical signs were not present in 68% of the 483 goats and 73% of the 286 sheep that tested seropositive for antibodies against PPRV. The presence of these 'silent spreaders', coupled with uncontrolled extensive livestock movement within the country and across the international borders, may explain the detection of antibodies against PPRV without clinical disease in the south-western parts of the country (Tanzania-Zambia and Tanzania-Malawi borders), as shown in Figure 1.
Discussion
Our results are in agreement with observations made by Swai et al. (2009), who confirmed the presence of PPR in Tanzania. As PPR has not been previously reported in Tanzania, it can be considered an emerging disease in Tanzania. Absence of PPR in Tanzania was demonstrated serologically in 2000 (Wambura 2000). The identification of the PPRV strain, as well as the serological, virological and clinical findings of this study confirmed that the disease was PPR. Clinical and post-mortem examination may be sufficient for the diagnosis of PPR in endemic areas, but molecular methods of characterisation are compulsory in areas where the outbreaks occur for the first time (Scott 1990). Genotypic classification based on the N-protein gene appeared to be an effective method for discriminating between geographically distinct lineages (Kwiatek et al. 2007).
Pastoral districts appeared generally to have suffered more from infection (Z = 15.38; p < 0.0001) than the agro-pastoral districts (Table 2). Reasons for this may be related to extensive livestock movements and exchanges resulting in a high rate of contact between animals, which increases the potential for transmission of PPRV.
Pastoral animals will often travel long distances during the dry season in search of fodder and water. Consequently, a considerable proportion of the population becomes infected during this period, giving rise to the establishment of disease endemicity and continued, year-round circulation of the virus. Farmers are often not aware of the pitfalls of mixing sick and healthy animal for grazing. For example, the study team saw diseased small ruminants and in-contact animals being sold at the market despite the ongoing outbreak. The purchase of new stock from the market and the return of unsold animals will often precipitate in severe outbreaks. The movement of animals, therefore, plays an important role in the spread and maintenance of PPRV in flocks, as does the purchase of potentially infected animals and their subsequent introduction into naïve flocks.
Age appears to be a risk indicator (p < 0.001) for seropositive PPR status, as suggested by the observed age profile according to which about 43% of the flock consists of young animals. Although goats are considered to be more susceptible to PPRV than sheep (Abubakar et al. 2011), there was no association (p = 1.000) between animal species and seropositivity in the current study. This could be due to the fact that sheep and goats are equally exposed to the PPR risk factors such as movement and nutritional stresses, or it could be that the circulating field strain has overcome the natural resistance of sheep; different types of breed might affect the outcome of PPRV infection. The significant relationship (p = < 0.001) between sex and seropositive PPR status could be due to the uneven distribution of the two sexes in the study flocks.
The origin of PPRV in Tanzania is not well established, but it is believed that it might have been introduced to Tanzania through the movement of live, infected animals. This hypothesis is supported by the fact that the border districts such as Longido and Ngorongoro had the highest prevalence (Table 1) of PPRV antibodies; moreover, PPRV was isolated from tissue samples that originated from the Ngorongoro district. The results of molecular epidemiological techniques suggest cross-border transmission of actively circulating PPRV into Tanzania from neighbouring northern countries. The virus we isolated is PPRV lineage III, which includes viruses whose origins are in East Africa. An outbreak in Ethiopia in 1996 as well as one in the Middle East was of this type (Abubakar et al. 2011). Thus, one of the northern neighbouring countries in the East Africa region is most likely the source of infection.
The evolving East African situation regarding PPR for the period 2004-2009 can be traced in the World Organisation for Animal Health's WAHID database (http://www.oie.int/wahis/public.php?page=disease_timelines) (OIE 2009), where the different waves of incursion of PPR can be seen. The first is observed in Kenya in March 2006, a second in Uganda in 2007 and the last one in Tanzania in December 2008. The majority of north-eastern countries such as Ethiopia and Sudan have been reporting the disease for more than 10 years (Lefèvre et al. 1991). For PPRV transmission, infected and healthy animals need to be in contact (Lefèvre et al. 1991), which serves as an additional indication that the virus came into Tanzania through progressive introduction of live, infected animals from these countries. Trade, as well as the nomadic lifestyle of the people in this region, is to a great extent responsible for the phenomenon. Moreover, the general extension of PPR emergence in Africa is certainly linked to the expansion of trade in animals and subsequent movement of these animals into previously disease-free regions. This phenomenon is further advanced by the presence of a significant proportion of 'silent spreaders'.
The epidemic curve revealed that during the outbreak cases occurred over multiple incubation periods. The shape and other features of an epidemic curve (Figure 3) can suggest hypotheses about the time and source of exposure, the mode of transmission, the causative agent, incubation period and the efficacy of control measures (Dicker & Gathany 1992). The outbreak had a successive series of peaks, reflecting increasing and decreasing numbers of cases in each generation. The epidemic waned after five generations, probably because either the number of susceptible animals fell below some critical level or the zoosanitary measures taken to control the disease became effective.
The estimates of Rt for PPRV infection (Table 3a) indicate that, on average, four secondary PPR infectious cases resulted from each primary infection in pastoral flocks. The variation in Rt estimates observed (Table 3a) is probably due to farming system and environmental variation, but it may also reflect the stochastic nature of PPRV. At this estimated Rt value (4), it might be expected that minor outbreaks of PPR infection would occur in which far fewer individuals seroconvert. This expectation is consistent with the number of reported sporadic PPR episodes. However, the generally poor reporting level suggests that an Rt value of 4 may be an underestimate of PPRV infection in the pastoral population. Furthermore, the morbilliviruses are known to be highly contagious, with a basic R0 between 15 and 20 (Anderson & May 1991), which increases the chances of an infection spreading or persisting within the population. However, given that the serological survey was conducted when the epidemic was still evolving, the host-pathogen relationship may not have reached equilibrium. Furthermore, it is possible that the assumption that animals were seronegative during the previous 12 months might not be valid, as the epidemic had been evolving for some time before the animals were sampled. The value of Rt derived in this study should therefore be cautiously considered a minimum estimate of the true population values.
There are limitations to generalisation of this estimated Rt for PPRV infections in other sheep and goat populations. In the current study, the animals were assumed to constitute a free-mixing population, approximating a homogeneously mixed flock. Whilst representative of the small ruminant farming system found in Tanzania, this production environment is quite different from that found in highland areas, where grazing land is scarce. For example, the effective contact rate is likely to differ between tethered and free-ranging animals, which would affect estimates of Rt. Other factors, such as PPRV strain, genetics of the flock, pre-incursion flock immunity, climate, population density and frequency of contact between flocks, may also affect Rt. Thus, Rt for PPRV may vary from the present estimates in other sheep and goat populations and environments. Further refinement of the estimates, which should include variables such as seasonality, environmental contamination, and contribution of the wild life population, as well as estimation of Rt in other sheep and goat production settings will be required to determine the general applicability of these estimates.
An Rt value of 4 predicts that 63.8% - 84.6% of small ruminants in a population would have to be vaccinated with a 100.0% efficient vaccine to reduce Rt to less than 1 (Table 3a). If, as stated earlier, Rt is underestimated, the percentage vaccine coverage would also be underestimated and would need to be increased for effective control of PPRV. For an imperfect vaccine or other interventions that are unlikely to be completely efficient (e.g. quarantine, movement restriction and closure of primary and secondary livestock auctions), effective coverage would probably have to approach 100.0%. The situation is further complicated by the possibility of undetected disease in areas where stock populations are sparse, the possibility that the disease may establish in the wildlife population and the possibility of delayed detection and recognition, which would provide a potential source for re-infection of sheep and goats. Thus, whilst some interventions may reduce the number of infected sheep and goats, eliminating PPRV from pastoral flocks will require significant effort and development of highly effective intervention tools, as for rinderpest campaigns.
The control of transnational livestock movements across borders is difficult, especially in the northern districts where Maasai pastoralists are found on either side of the Tanzania-Kenya border. The resulting transborder movements of cattle, goats and sheep for breeding, trade and other socio-economic functions could be the main predictor for introduction, transmission and maintenance of PPRV in the country. Without proper control mechanisms, these animal movements could contribute to and accelerate the spread of the virus regionally. Indeed, the PPRV-positive samples from the Tanzania-Mozambique, Tanzania-Malawi and Tanzania-Zambia borders (Figure 1) indicate a southward movement of the disease. Therefore, the importance of PPR as a threat to small ruminants should be considered together with other economically important diseases and measures should be taken to prevent subsequent spread of such diseases into the region of the Southern African Development Community (SADC).
Conclusion
The objective of this work was to investigate the incursion, persistence and spread of PPRV in Tanzania. Our results contribute to the epidemiological understanding that PPRV is established in Tanzania. Extensive livestock movement, failure to observe basic zoosanitary measures, delayed reporting and lack of proper control strategies at the local and national levels are the main factors contributing to the persistence of PPRV in Tanzania. The establishment of PPRV in Tanzania serves as a reservoir for the rest of the SADC region through livestock movement and trade of infected animals.
Acknowledgements
We are grateful to the Maasai herdsmen who gave their time and allowed us to sample their animals. The help and participation of staff at the Department of Veterinary Services, Ministry of Livestock and Fisheries Development of Tanzania, is highly appreciated.
Competing interests
The authors declare that they have no financial or personal relationship(s) that may have inappropriately influenced them in writing this article.
Authors' contributions
F.M.K. (Ministry of Livestock and Fisheries Development) was the group leader and responsible for the study design and data analysis. O.K. (CIRAD) and G.L. (CIRAD) were responsible for the laboratory and molecular epidemiology work. A.M.K. (Central Veterinary Laboratory), J.G. (Central Veterinary Laboratory), E.S.S. (Veterinary Investigation Centre, Arusha), W.M. (Veterinary Investigation Centre, Arusha) and A.O.M (Veterinary Investigation Centre, Mtwara) were responsible for fieldwork.
References
Abubakar, M., Khan, H.A., Arshed, J.M., Hussain, M. & Qurban, A., 2011, 'Peste des petits ruminants (PPR): Disease appraisal with global and Pakistan perspective', Small Ruminant Research 96, 1-10. [ Links ]
Anderson, R.M. & May, R.M., 1991, Infectious Diseases of Humans: Dynamics and Control, Oxford University Press, London. [ Links ]
Dicker, R. & Gathany, N.C., 1992, Principles of Epidemiology, 2nd edn., U.S. Department of Health and Human Services, Public Health Service, Centre for Disease Control and Prevention (CDC), Atlanta. [ Links ]
Dietz, K., 1993, 'The estimation of the basic reproduction number for infectious diseases', Statistical Methods in Medical Research 2, 23-41. [ Links ]
Dohoo, I., Martin, W. & Stryhn, H., 2003, Veterinary Epidemiologic Research, Atlantic Veterinary College, Charlottetown. [ Links ]
Food and Agriculture Organization/International Atomic Energy Agency (FAO/IAEA), 1993, 'Peste des Petits Ruminants ELISA kit. Competitive enzyme immunoassay for detection of antibody to PPR virus. Bench protocol', version PPR 1.0, January 1993. Joint FAO/IEAE Programme, Animal Production and Health. Pirbright, United Kingdom: World Reference Laboratory for Rinderpest. [ Links ]
Furley, C.W., Taylor, W.P. & Obi, T.U., 1987, 'An outbreak of peste des petits ruminants in a zoological collection', Veterinary Record 121, 443-447. [ Links ]
Heffernan, J.M. & Wahl, L.M., 2005, 'Monte Carlo estimates of natural variation in HIV infection', Journal of Theoretical Biology 236, 137-153. [ Links ]
Huelsenbeck, J.P. & Ronquist, F., 2001, 'MRBAYES: Bayesian inference of phylogenetic trees (http://mrbayes.sourceforge.net/links.php)', Bioinformatics 17, 754-755. [ Links ]
Jobb, G., 2011, TREEFINDER, computer software, viewed 17 August 2011 from http://www..treefinder.de [ Links ]
Kwiatek, O., Minet, C., Grillet, C., Hurard, C., Carlsson, E. & Karimov, B., 2007, 'Peste des petits ruminants (PPR) outbreak in Tajikistan', Journal of Comparative Pathology 136(2-3),111-119. [ Links ]
Lefèvre, P.C., Diallo, A., Schenkel, S., Hussein, S. & Staak, G., 1991, 'Serological evidence of peste des petits ruminants in Jordan', Veterinary Record 128, 110. [ Links ]
Libeau, G., Diallo, A., Calvez, D. & Lefèvre, P.C., 1995, 'A competitive ELISA using anti-N monoclonal antibodies for specific detection of rinderpest in cattle and small ruminants', Veterinary Microbiology 31, 147-160. [ Links ]
Office International des Epizooties (OIE), 2000, OIE manual of standards for diagnostic tests and vaccines. List A and B diseases of mammals, birds and bees, OIE, Paris. [ Links ]
Özkul, A., Akca, Y., Alkan, F., Barrett, T., Karaoglu., T., Dagalp, S.B. et al., 2002, 'Prevalence, distribution, and host range of Peste des petits ruminants virus, Turkey', Emerging Infectious Diseases 8, 708-712. [ Links ]
Scott, G.R., 1990, 'Rinderpest and peste des petits ruminants', in E.P.J. Gibbs (ed.), Virus Diseases of Food Animals, vol. 2, Disease Monographs, Academic Press, New York. [ Links ]
Snedecor, G.W. & Cochran, W.G., 1989, Statistical Methods, 8th edn., Iowa State University Press, Ames, Iowa. [ Links ]
Swai, E.S., Kapaga, A., Kivaria, F., Tinuga, D., Joshua, G. & Sanka, P., 2009, 'Prevalence and distribution of Peste des petits ruminants virus antibodies in various districts of Tanzania', Veterinary Research Communications 33, 927-936. http://dx/doi.org/10.1007/s11259-009-9311-7 [ Links ]
Wambura, P., 2000, 'Serological evidence of the absence of peste des petits ruminants in Tanzania', Veterinary Record 146, 473-474. [ Links ]
 Correspondence:
Correspondence:
Fredrick Kivaria
PO Box 9152
Dar es Salaam, Tanzania
fredkiv@gmail.com
Received: 15 Mar. 2013
Accepted: 25 Aug. 2013
Published: 30 Oct. 2013
1. An additional 20 villages were sampled under the VACNADA project.














